Category: Coding
Importing and exporting molecules in Python
Creating replicas of structures
Accessing the data graph in Python
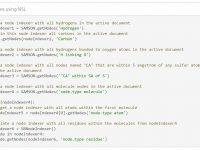
To efficiently manipulate nodes (atoms, bonds, side chains, molecules, etc.), use node indexers returned by the SAMSON.getNodes function applied to a Node Specification Language (NSL) expression. For example, SAMSON.getNodes(‘node.type atom’) returns an indexer containing all atoms in the active document.…
Create molecules using Python
Triggering GUI commands using Python
Creating Python Apps with GUIs
Importing and exporting molecules in Python
Creating replicas of structures
Accessing the data graph in Python

To efficiently manipulate nodes (atoms, bonds, side chains, molecules, etc.), use node indexers returned by the SAMSON.getNodes function applied to a Node Specification Language (NSL) expression. For example, SAMSON.getNodes(‘node.type atom’) returns an indexer containing all atoms in the active document.…