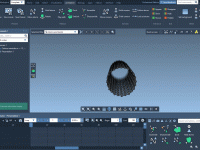
Mastering the Dolly Camera Animation in Molecular Modeling
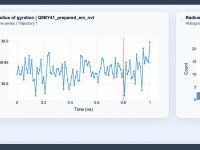
Understanding Compactness with Radius of Gyration Analysis
Streamlining System Preparation with GROMACS Wizard
Building Custom Functionalities with SAMSON Extensions
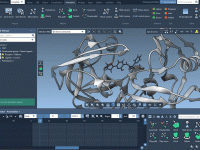
A Step-By-Step Guide to Aligning Protein Sequences and Structures with SAMSON
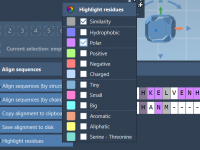
Molecular modelers often face the challenge of comparing proteins to identify conserved residues, understand structural similarities, or build homology models. Accurately aligning protein sequences and structures is crucial for gaining biological insights, but finding a fast, reliable, and streamlined way…
Mastering Transparency with the Appear Animation in SAMSON.
Achieve Effortless Reproducibility: Automating Molecular Workflows in SAMSON
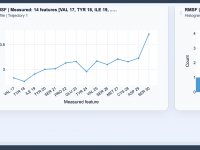
Visualizing Flexibility in Molecular Structures with RMSF Profiles
Streamlining Node Selection with SAMSON’s Path Attributes
As a molecular modeler, pinpointing specific nodes within complex molecular structures is a vital but often time-consuming task. Understanding how to effectively use selection parameters can significantly ease this process. Fortunately, SAMSON’s Node Specification Language (NSL) offers powerful tools to…