Molecular modelers often encounter the challenge of understanding and visualizing how proteins assemble within crystal structures and biological assemblies. This insight is crucial for protein-modeling workflows, particularly when designing molecular systems or analyzing quaternary structures. If you’ve ever struggled to unravel these symmetrical arrangements, SAMSON’s Symmetry Mate Editor provides an efficient and intuitive solution.
The Core Problem: Visualizing Protein Symmetries
Proteins often assemble into larger structures, such as symmetric complexes, as part of their crystallization process or biological function. These assemblies provide researchers with essential details about molecular interactions, binding sites, and structural organization. However, recreating such assemblies from the information embedded in PDB files can be complex when done manually.
SAMSON’s Symmetry Mate Editor simplifies this task by leveraging symmetry transformation data (available in CRYST1 or BIOMT records within PDB files) to generate replicas of proteins. This eliminates the need for time-consuming manual reconstruction and gives you a better understanding of protein arrangements in just a few clicks.
How to Use the Symmetry Mate Editor
To access and activate the editor, follow these simple steps:
- Using Find Everything: Press Shift + E and search for ‘Symmetry Mate Editor’.
- From the Editors Menu: Navigate through the left-side viewport menu: … > General > Symmetry Mate Editor.
Once activated, you’ll see control nodes appear in the viewport to represent symmetry transformations.
Interactive Exploration of Symmetry
SAMSON’s interactive tools make exploring symmetries both intuitive and efficient:
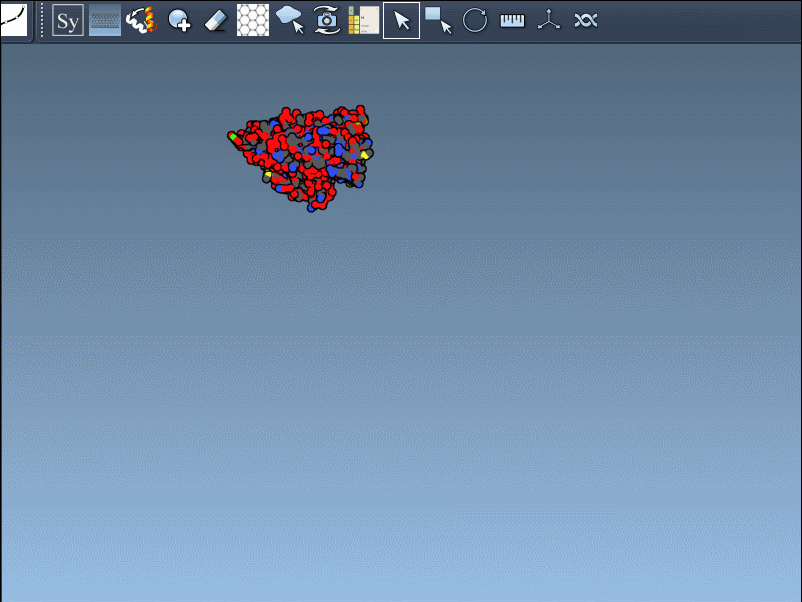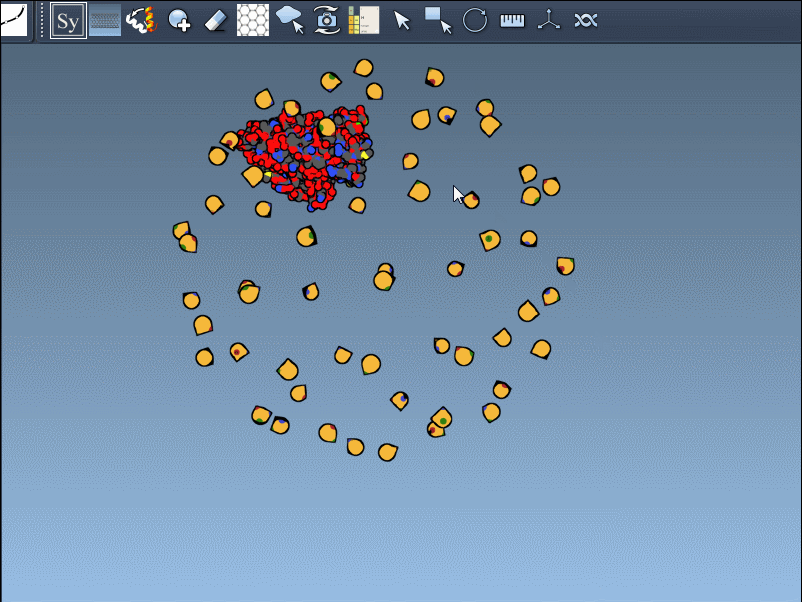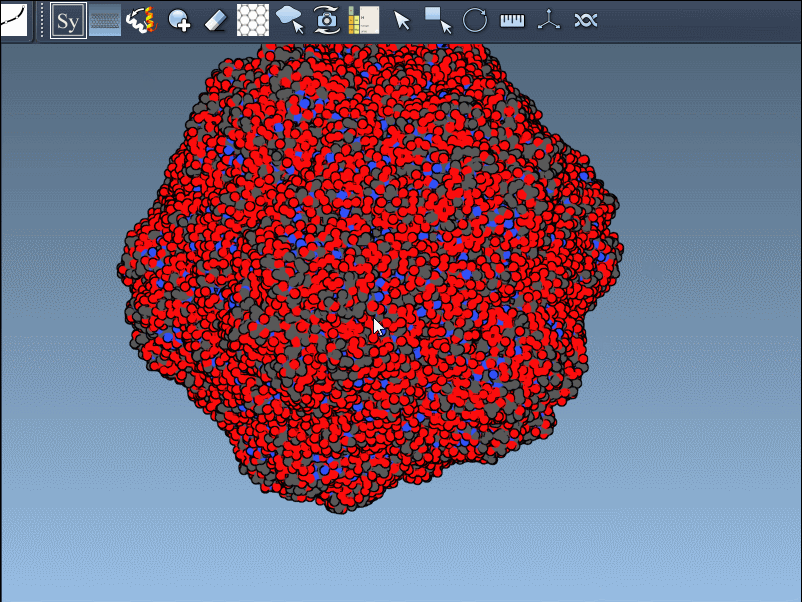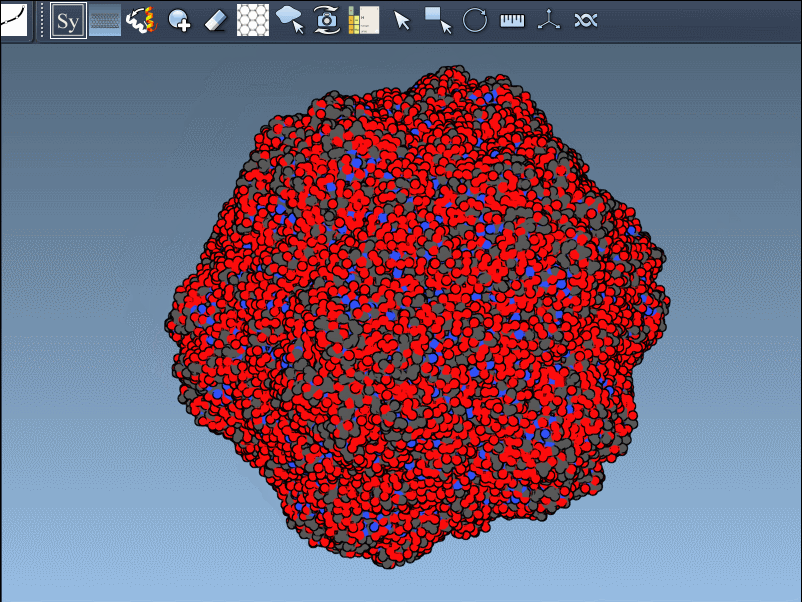
- Scroll to adjust the number of control nodes: Use Ctrl / Cmd + mouse wheel to scale visible control nodes.
- Hover for real-time previews: Hovering over any node previews the corresponding replica instantly in your viewport. This allows quick identification of structures before committing to their generation.
- Left-click to finalize: Clicking a node permanently generates that replica – ideal for constructing complex molecular arrangements.
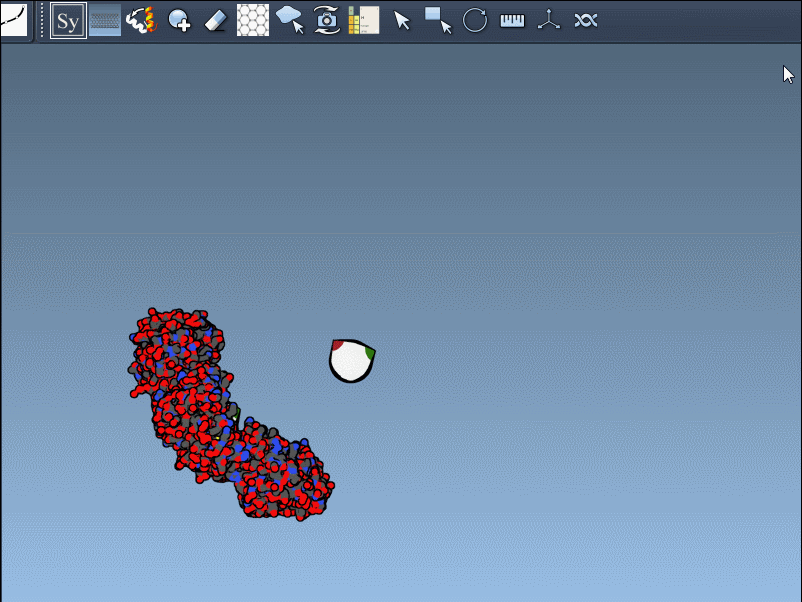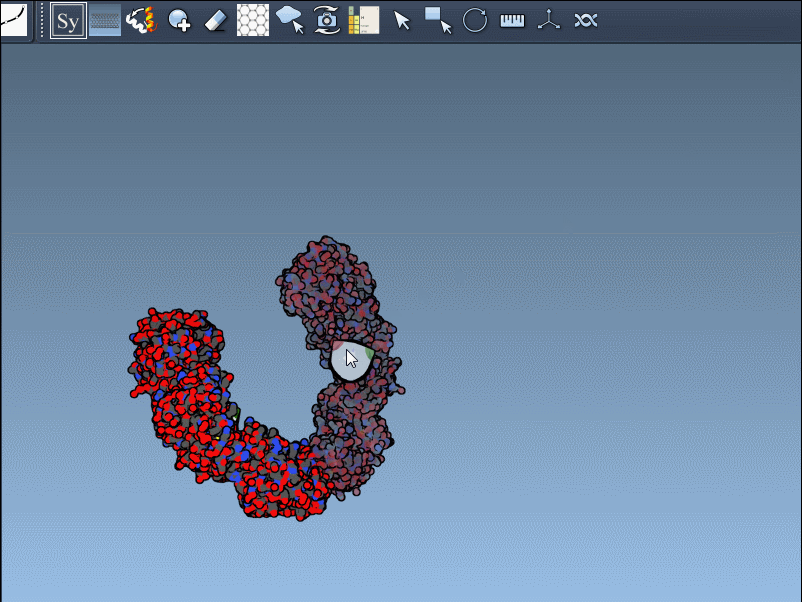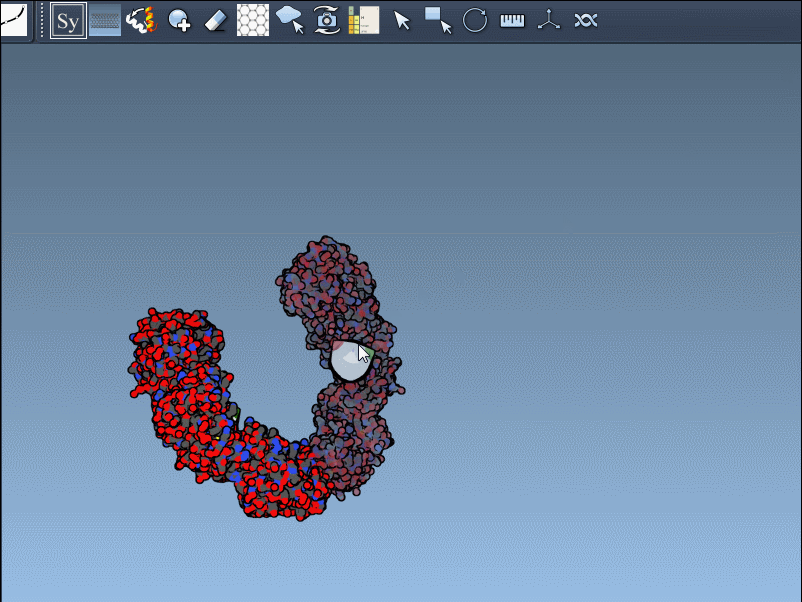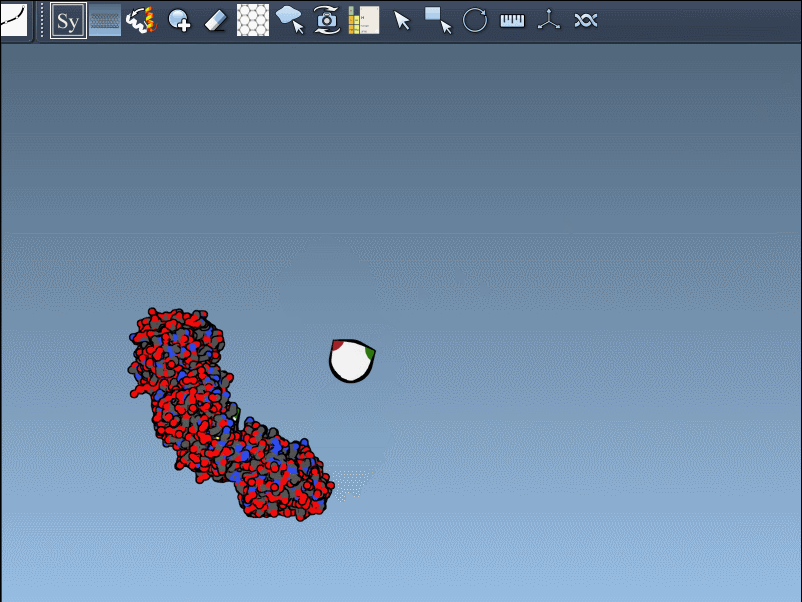
Want to create all mates simultaneously? Hold down Ctrl / Cmd while hovering and selecting a node to replicate the entire symmetric assembly in one go.
A Closer Look at CRYST1 vs. BIOMT Records
The editor supports both CRYST1 and BIOMT symmetry types, denoted in PDB files:
| Record Type | Description | Widget Color |
|---|---|---|
| CRYST1 | Symmetry from crystal lattice | White |
| BIOMT | Symmetry from biological assembly annotations | Yellow |
You can switch between these two modes via the interface, helping you identify the source of symmetry data and toggle between relevant configurations for your research. Here’s what it looks like in action:
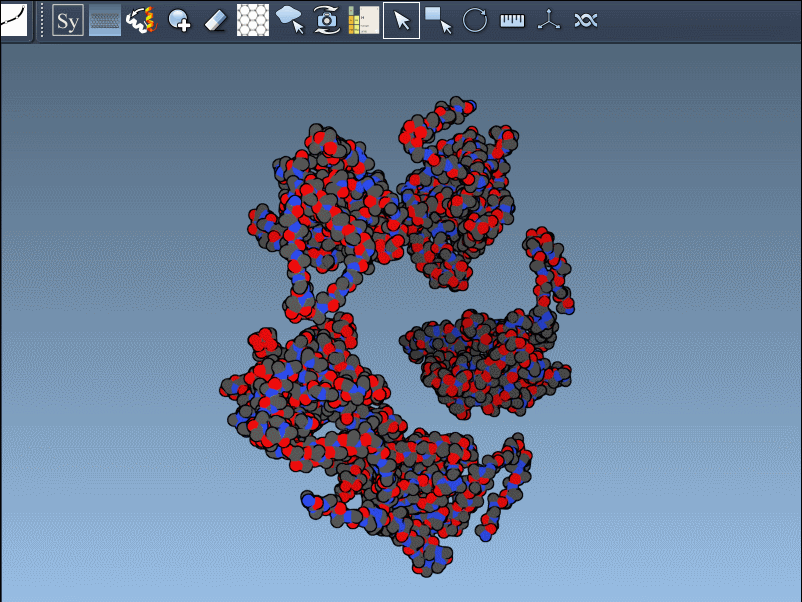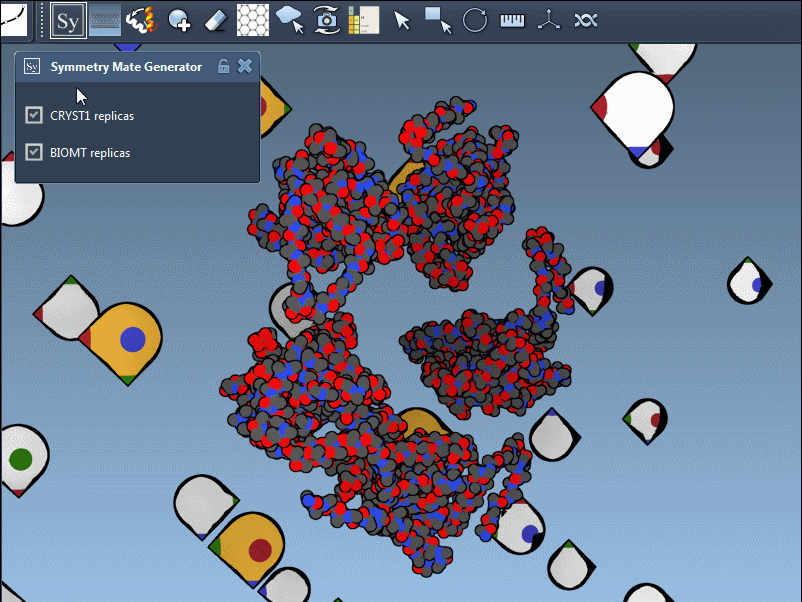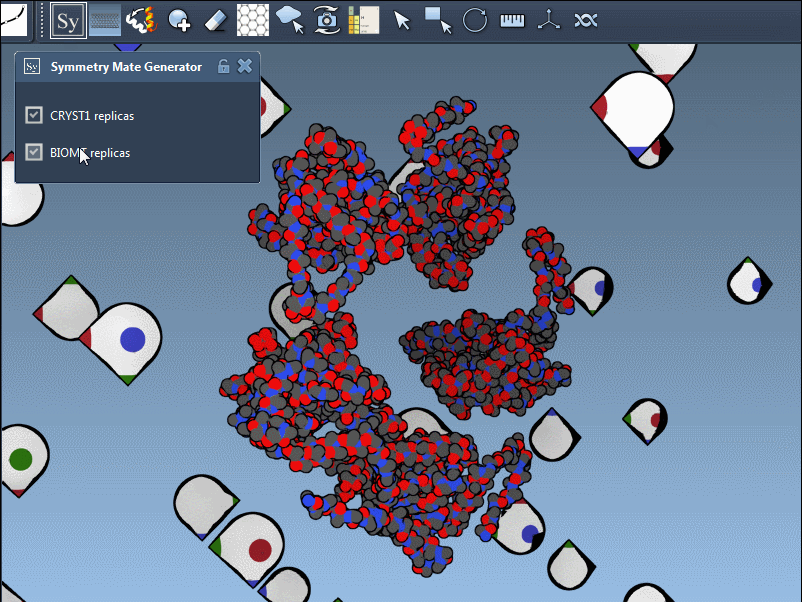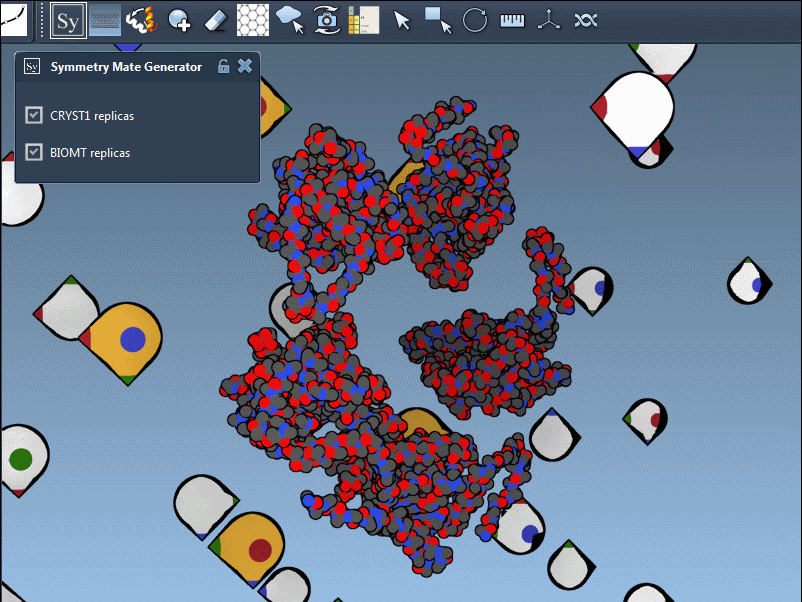
Applications for Molecular Modelers
By simplifying symmetry generation, this tool offers immense value for various molecular workflows, including:
- Reconstructing complete oligomeric assemblies from asymmetric units.
- Designing symmetric protein cages or scaffolds for nanotechnology applications.
- Analyzing repetitive binding sites across symmetric chains to understand interactions.
- Setting up simulations where symmetric structures play a critical role, such as molecular dynamics of quaternary assemblies.
Conclusion
The Symmetry Mate Editor is an indispensable tool for understanding and recreating the symmetric organization of proteins. With its intuitive features and the ability to quickly toggle between different symmetry sources, it caters to the needs of researchers working on structural biology and molecular design.
Explore the full documentation of the Symmetry Mate Editor here: https://documentation.samson-connect.net/tutorials/symmetry/generating-symmetry-mates/.
Note: SAMSON and all SAMSON Extensions are free for non-commercial use. Get started today by downloading SAMSON at https://www.samson-connect.net.





