For molecular modelers exploring protein dynamics, understanding the movement of key structures like the SARS-CoV-2 spike protein is crucial for advancing research. The spike protein’s transition from a closed to an open state, enabling binding with the ACE2 receptor, is a fundamental step in the virus’s ability to infect human cells.
This blog post will guide you through the computational data and visualizations of the spike’s motion as derived using SAMSON, the integrative molecular modeling platform. The information presented here offers an insightful look at how such tools can help model and understand complex biomolecular interactions.
Why Study the Spike’s Motion?
The SARS-CoV-2 spike facilitates the virus’s entry into cells by transitioning into an open state, which allows it to bind to the ACE2 receptor. Understanding this motion is valuable for researchers developing vaccines, therapeutics, or inhibitors aimed at disrupting this binding mechanism. Despite the challenges of experimental validation, computational approaches offer models and trajectories to visualize and hypothesize the mechanics of this interaction.
The Spike in Motion: See It for Yourself
SAMSON’s advanced tools, including the ARAP (As-Rigid-As-Possible) Interpolation Path module and the P-NEB (Parallel Nudged Elastic Band) module, were employed to compute the transition of the spike protein from its closed state to its open receptor-binding state. The following visualizations demonstrate this motion:
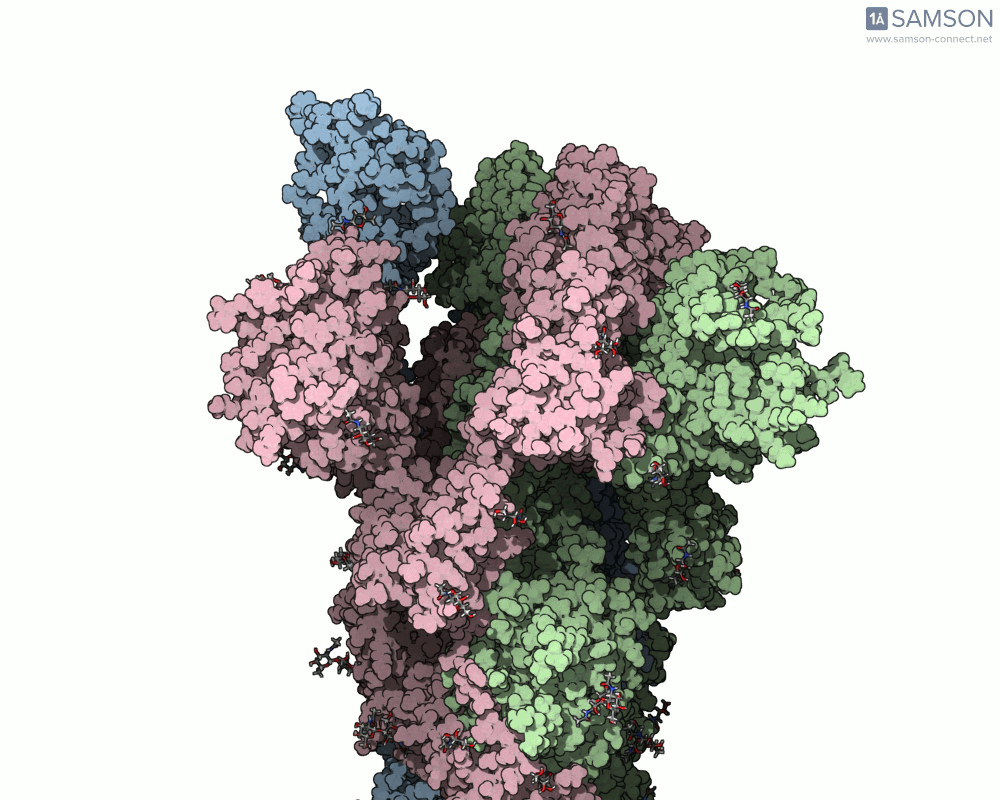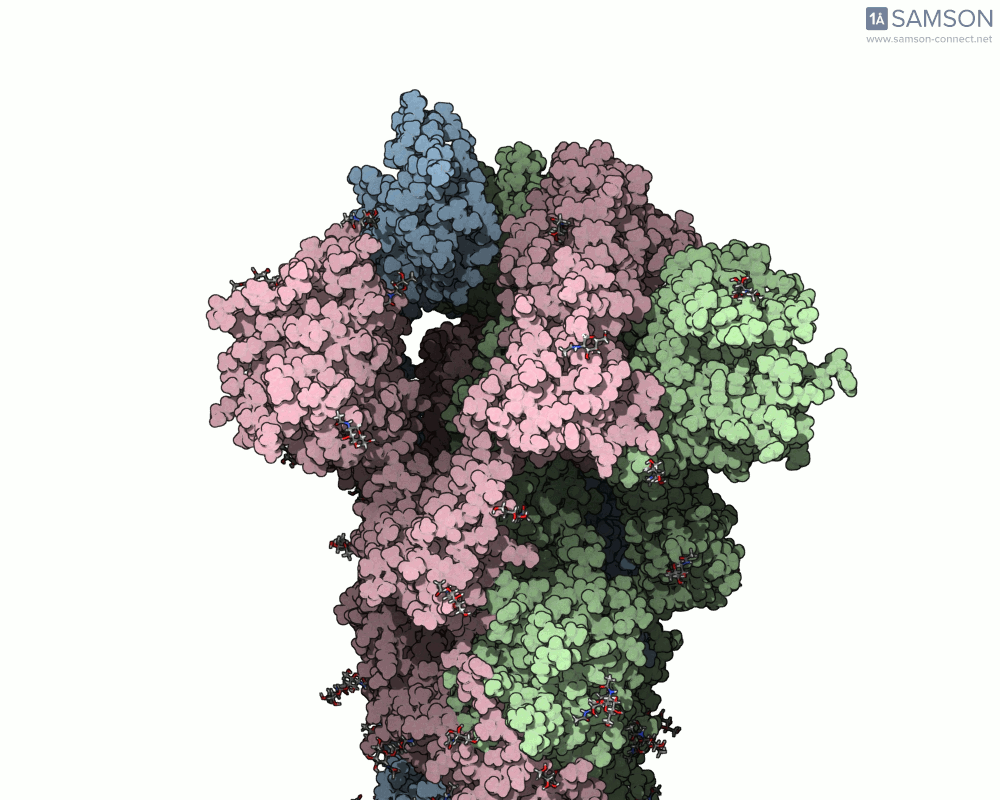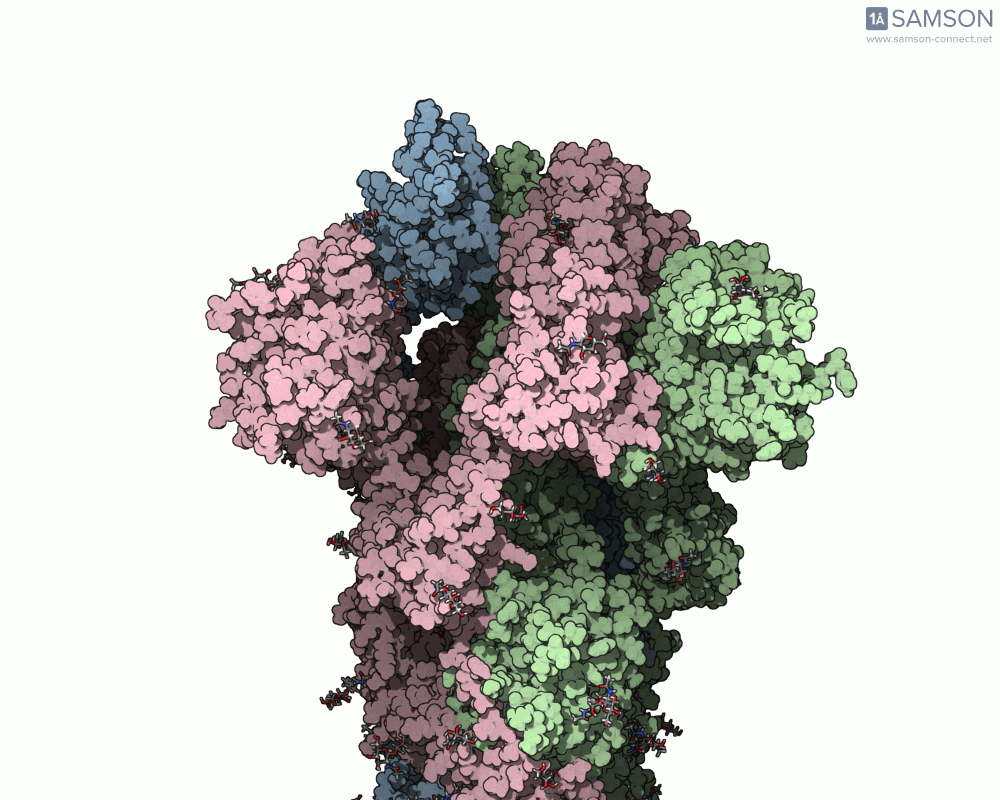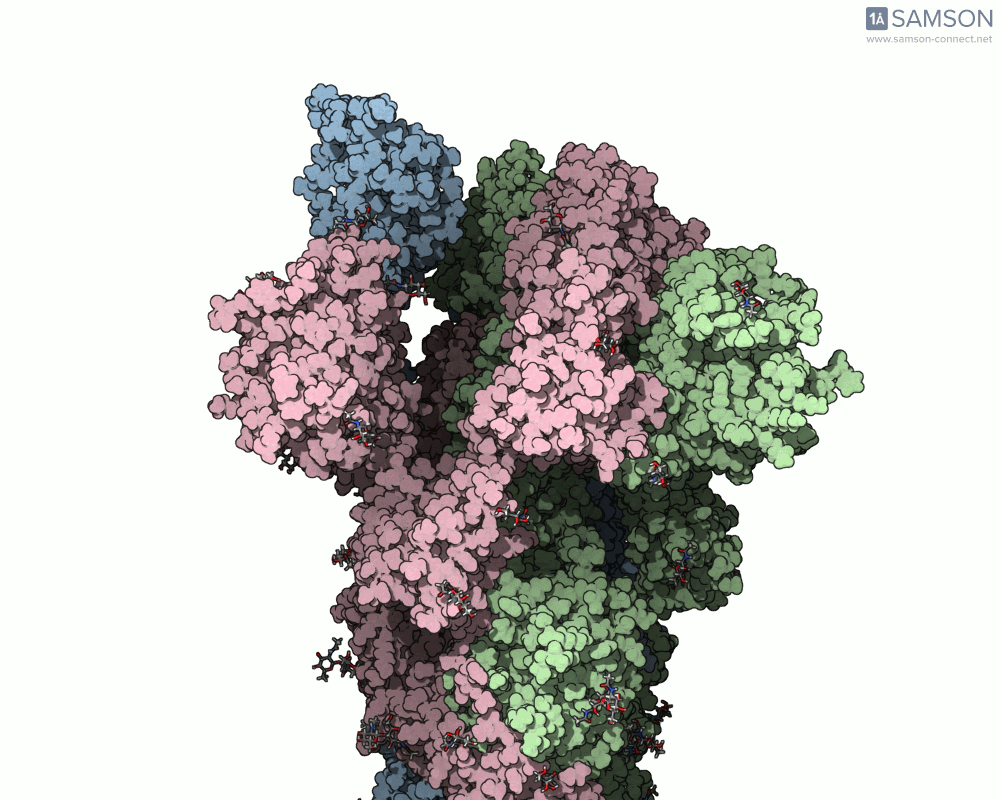
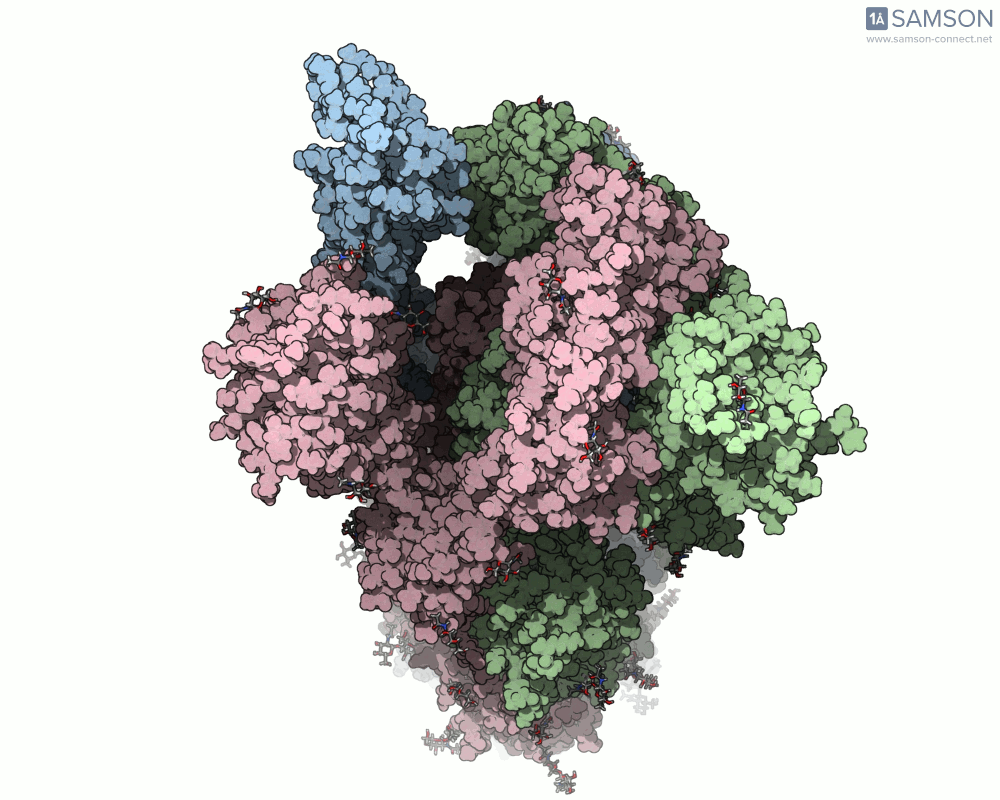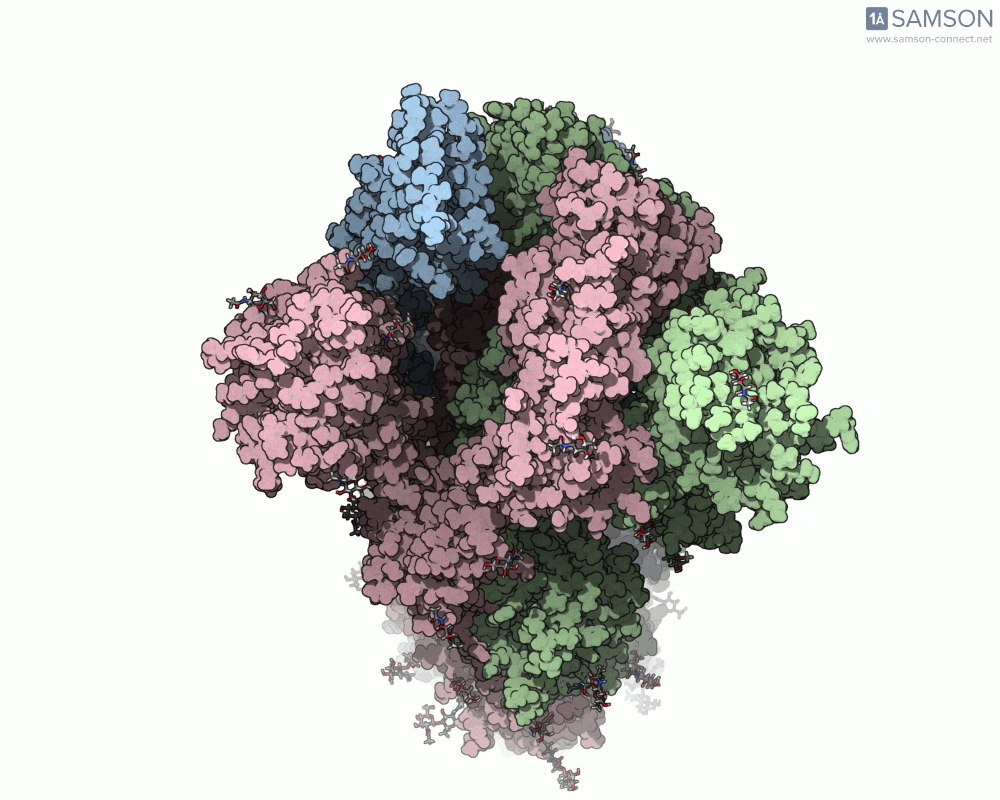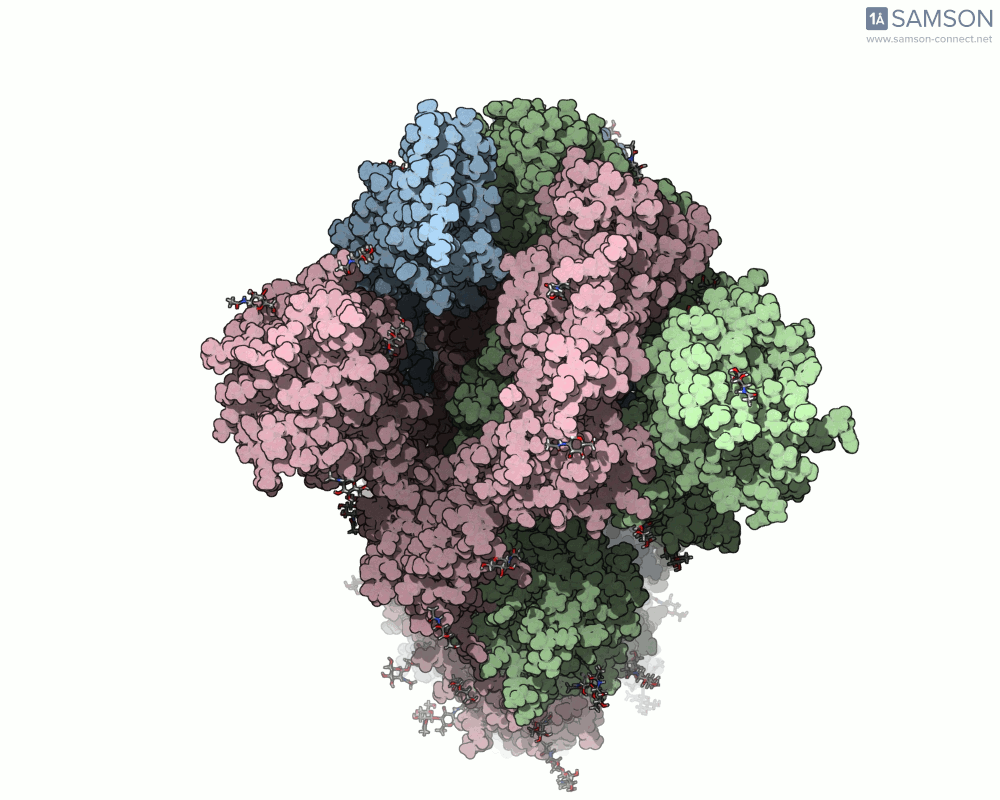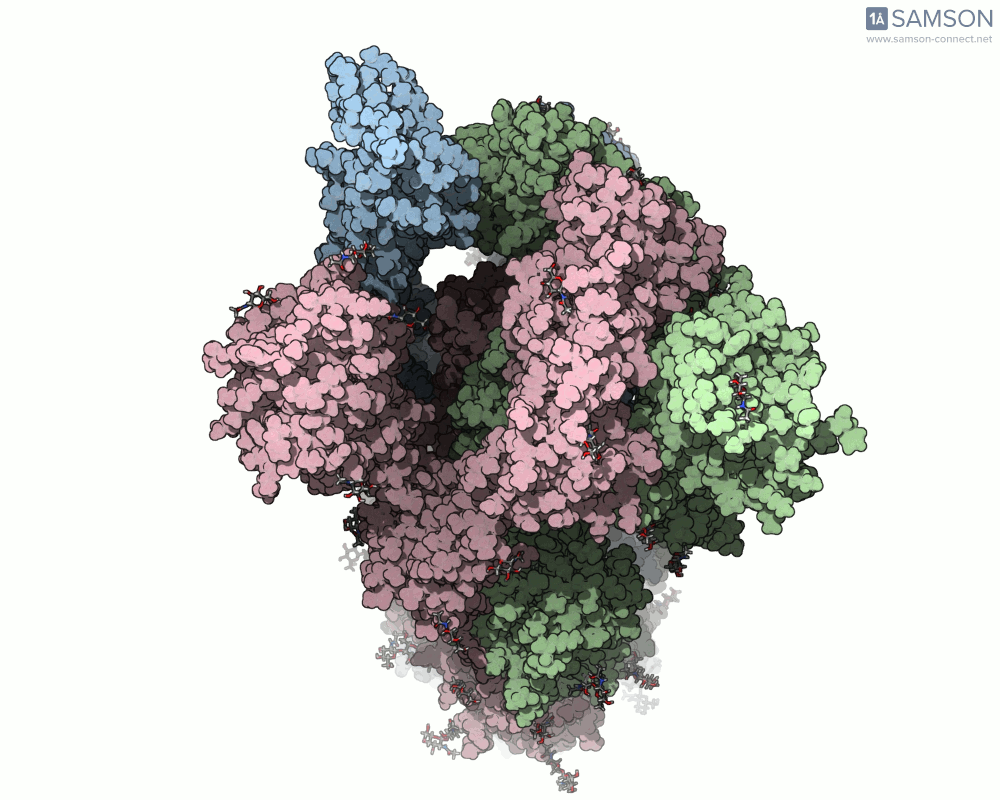
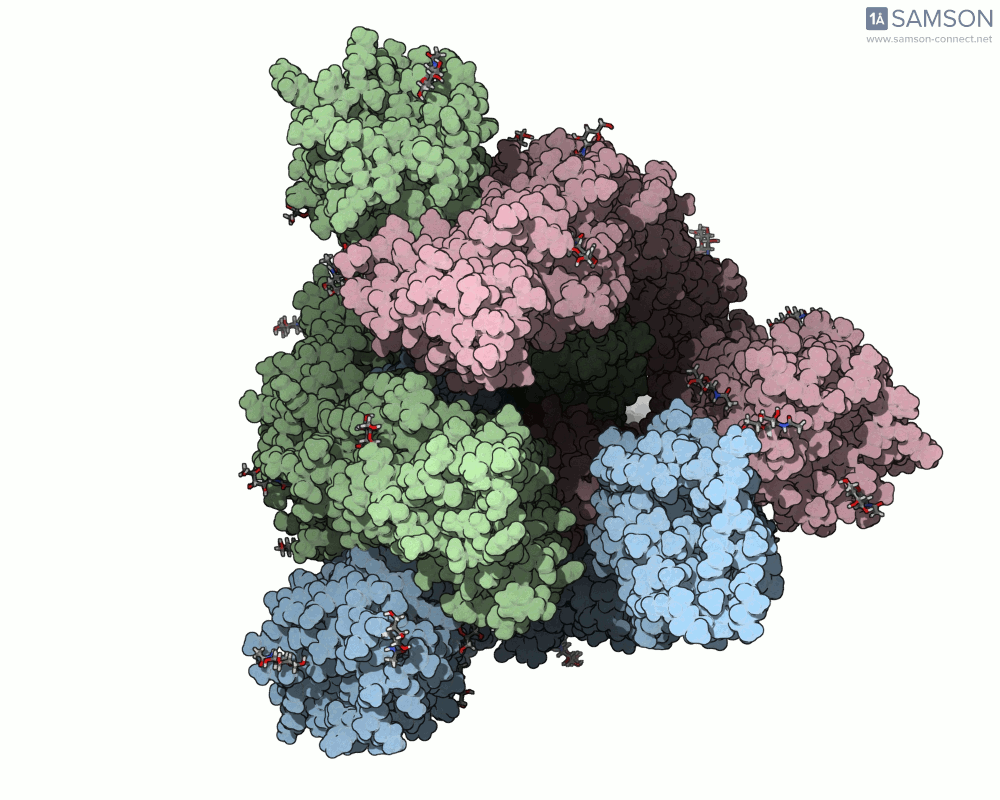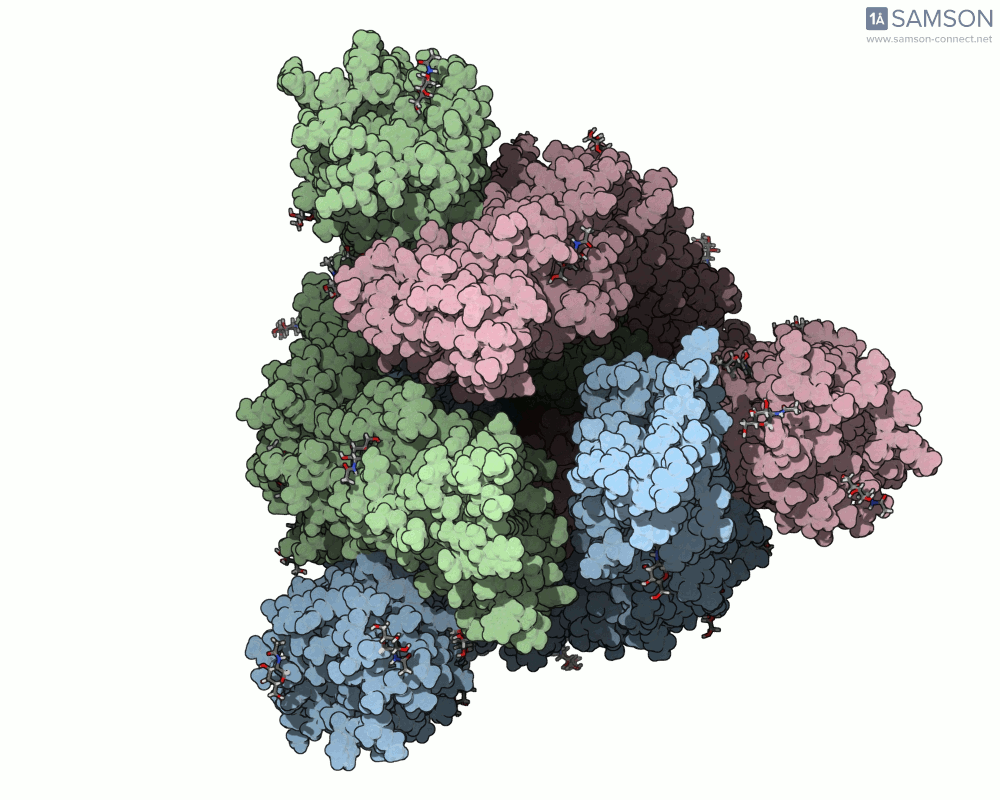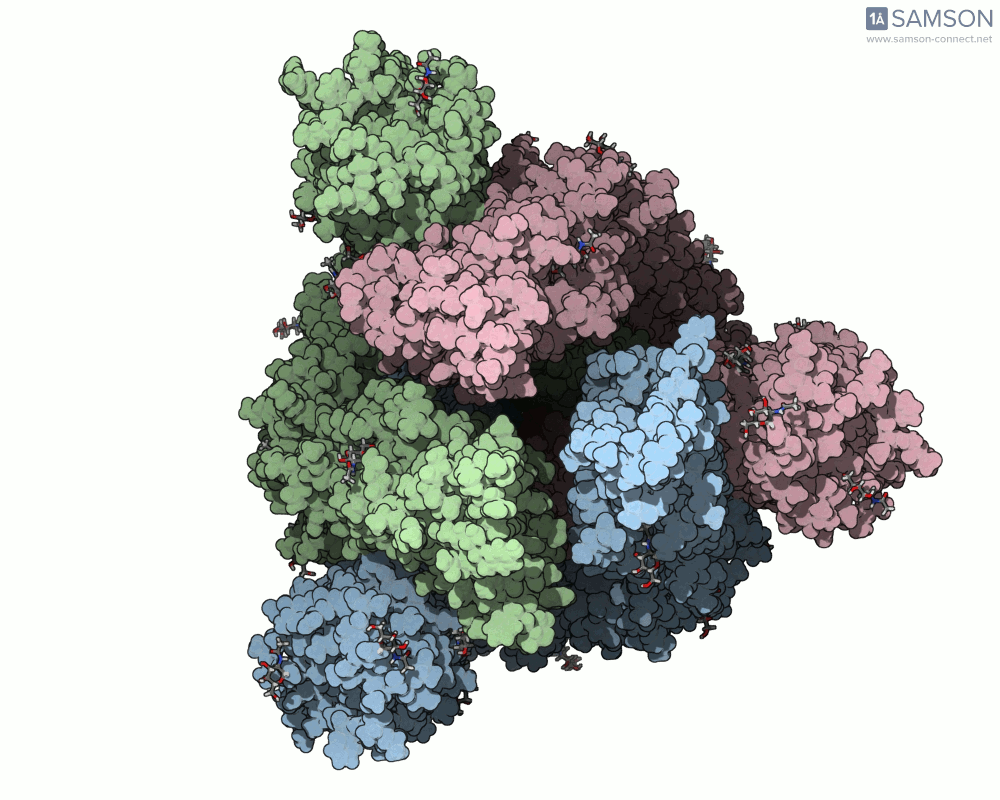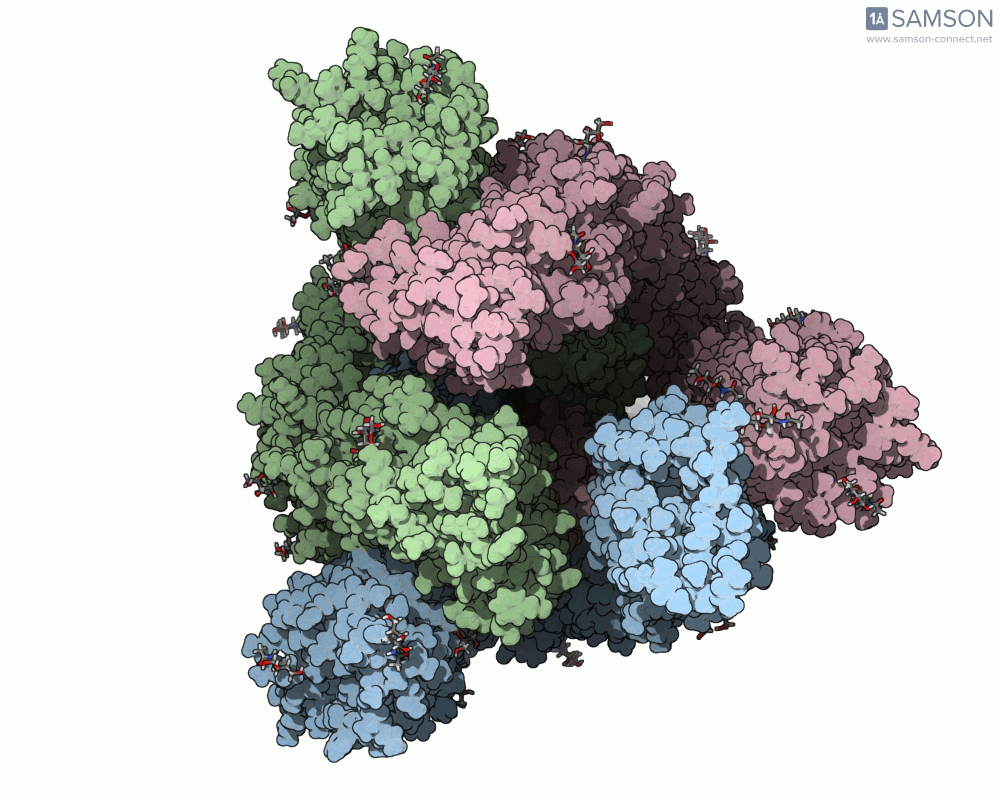
How Was This Motion Computed?
The computational pipeline for deriving the spike’s trajectory involved the following steps in SAMSON:
- Two known states of the spike protein were used as inputs: the closed state (PDB: 6VXX) and the open state (PDB: 6VYB).
- The ARAP Interpolation Path module provided a first approximation of the transition pathway.
- The P-NEB module was then applied to refine the computed trajectory, ensuring smooth transitions and lower-energy states.
This process took less than half an hour on a standard laptop and resulted in high-quality trajectories illustrating the spike’s transformation.
Accessing Data and Tools
The computed trajectory files are available for download in various formats, including PDB and SAMSON native formats. These can be used for further analysis or animations. SAMSON also provides free access to the necessary modules during the outbreak, allowing researchers worldwide to replicate or build upon this work.
You can download the trajectory files from the original documentation page.
Conclusion
Visualizing the SARS-CoV-2 spike protein’s motion provides critical insights into how the virus interacts with host cells. By leveraging SAMSON’s computational tools, researchers can generate trajectory data to model and study these dynamics, enriching our understanding of viral mechanisms. To learn more and explore the full methodology, visit the original documentation page: https://documentation.samson-connect.net/tutorials/sars-cov-2/coronavirus-computing-the-opening-motion-of-the-sars-cov-2-spike/.
Note: SAMSON and all SAMSON Extensions are free for non-commercial use. You can download SAMSON for free at https://www.samson-connect.net.





