Potential of Mean Force (PMF) analysis is a fundamental tool in molecular modeling for calculating free energy surfaces and understanding reaction pathways. For researchers diving into molecular dynamic simulations, PMF can sometimes appear daunting. If you’ve ever struggled with organizing your results or ensuring accurate reaction coordinate coverage, the GROMACS Wizard’s PMF analysis feature offers a streamlined workflow.
Here, we outline how to efficiently compute PMF using the Weighted Histogram Analysis Method (WHAM) in the GROMACS Wizard on the SAMSON platform. By following this guide, you’ll optimize your time and gain confidence in setting up your system for productive results.
Step 1: Setting Up Your Project Directory
The first crucial step is organizing your project files. The PMF analysis requires your project folder to contain numbered subfolders. Each subfolder should have simulation results corresponding to the same system and reaction coordinate. This systematic setup ensures that GROMACS Wizard can seamlessly process the data later. If you’ve previously conducted an Umbrella Sampling simulation or batch computations, you can easily auto-fill the path to your data folder by clicking the auto-fill button (![]() ).
).
Step 2: Configuring the Analysis
Once your project directory is set up, switch to the WHAM Analysis tab within the GROMACS Wizard. The system automatically loads necessary project data, such as reaction coordinate details, time, and temperature settings. Select your desired reaction coordinate from the interface. Additionally, you can make custom adjustments such as modifying bounds, time settings, and energy units depending on your analysis objectives.
Step 3: Running the Computation
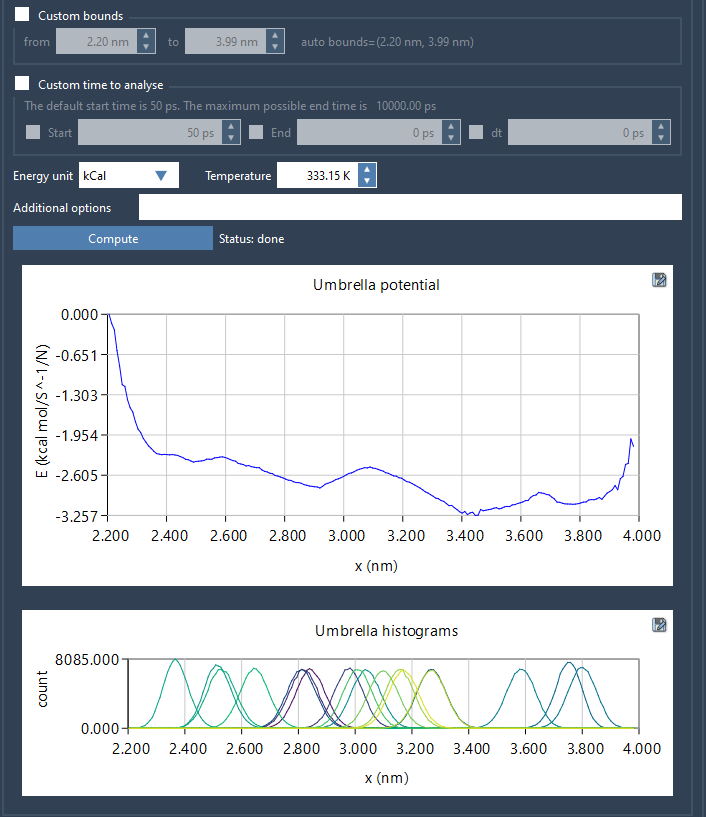
When ready, click the Compute button to start the PMF analysis. It takes only a matter of seconds or minutes, depending on the size of your trajectories. The results are presented as two main plots: the PMF graph and the histogram. The PMF graph gives a clear view of free energy variations along the reaction coordinate, while the histogram shows the coverage of reaction coordinate space. This histogram can help you identify regions requiring further simulations to improve data sampling.
Step 4: Managing Your Outputs
All analyzed data, including computed profiles, histograms, and plots, are automatically saved in the wham_results subfolder of your project directory. This convenient organization ensures that switching between reaction coordinates or tweaking analysis parameters is seamless—previously processed data is reused, greatly reducing computation times.
Conclusion
By leveraging the PMF analysis capabilities of the GROMACS Wizard, molecular modelers can simplify what initially might seem like a complex task. Accurate free energy surface computations are vital to many areas of integrative molecular design, and this tool offers an intuitive yet robust way to achieve that.
For more detailed guidance, visit the full documentation page.
SAMSON and all SAMSON Extensions are free for non-commercial use. You can get the platform and start exploring at https://www.samson-connect.net.





