Tracking atom trajectories through molecular pathways forms the backbone of simulations in molecular modeling. However, many researchers face challenges when trying to extract atomic coordinates along defined paths for advanced analyses like reaction coordinate studies or free energy calculations. If you’ve ever spent hours scripting or struggling to get the right data format, SAMSON’s Export Along Paths extension offers an intuitive, streamlined solution.
Why Do Molecular Modelers Need This?
In complex molecular systems, visualizing and analyzing atomic movements can shed invaluable insights on ligand interactions, reaction mechanisms, and thermodynamic profiles. From tracing ligand unbinding pathways to generating data for enhanced sampling methods, correctly exporting atomic trajectories can save time and allow precise downstream calculations. Unfortunately, traditional methods often involve tedious manual scripting or rigid frameworks that lack flexibility.
The Export Along Paths extension simplifies this by allowing you to effortlessly extract atomic coordinate sets, whether it’s the entire system or a subset of atoms, along predefined paths. Ready to dive in? Here’s how you can use this tool.
How to Export Trajectories with Ease
Here’s a straightforward guide to help you get started with this extension:
1. Load a Sample System
For tutorial purposes, you can begin with a sample system:
- Click Home > Download within SAMSON.
- Paste the following link into the download field: https://www.samson-connect.net/documents/046f1acd-c799-40f6-8185-cb4847eff795
- Hit Download to load a tutorial document containing the structure of Lactose permease along with its ligand (Thiodigalactoside) and two unbinding paths.

Within the Document view, observe that the system includes the protein (Protein_chain_A), the ligand (TDG), and predefined paths generated with the Ligand Path Finder extension.
2. Open the Export Along Paths App
Once the system is ready, open the tool:
- Access it via Home > Apps > All > Export Along Paths.
- You can also simplify navigation by pressing Shift+E.
The interface is designed for ease of use. You can manage advanced settings, choose export formats, and define atom subsets.
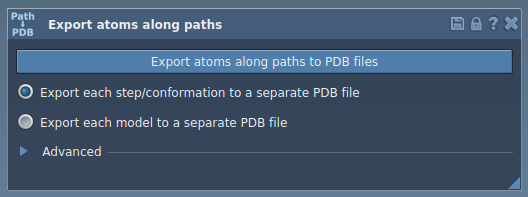
3. Export Atomic Coordinates
Option 1: Export All Atoms
If you want to export the entire atomic trajectory:
- Select one or more paths in the Document view.
- Choose your export format—either all frames in a single PDB file or each frame as separate PDB files.
- Click Export atoms along paths to PDB files and select a destination folder and file prefix.
Want to control frames? Expand the Advanced section to adjust frame export intervals.
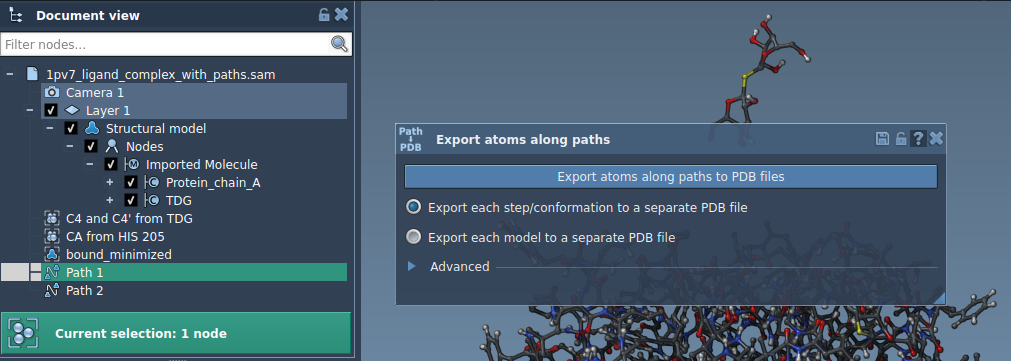
Option 2: Export a Subset of Atoms
Sometimes, your focus lies on a small subset of atoms—like ligands or specific residues. Here’s how:
- Expand the Advanced panel and select the subset (e.g.,
TDGligand). - Click Add to define this selection as a model for export.
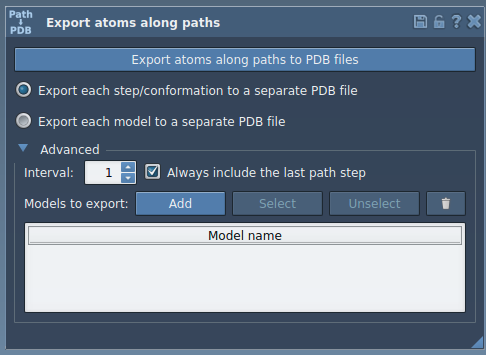
Define multiple models or rename them for clarity. Once your selection and paths are ready, click Export atoms along paths to PDB files.
Key Applications
This tool offers concrete ways to accelerate your modeling workflow:
- Generate reaction coordinate files for free energy profiling.
- Export ligand exit/entry trajectories for enhanced sampling simulations.
- Isolate intermediate states for custom visualization and insight.
- Focus on key subsets, like ligands or binding site residues, for precise analyses.
The Export Along Paths extension enables molecular modelers to save time while extracting actionable data tailored to specific systems and goals.
To learn more, visit the official documentation.
SAMSON and all SAMSON Extensions are free for non-commercial use. Get SAMSON today at SAMSON Connect.





