Molecular modelers often face the challenge of exploring how small changes to a molecule’s structure can impact its properties or interactions. Whether looking to enhance binding affinity, optimize pharmacokinetics, or explore new analogs, this task can be daunting without the right tools. If you want an intuitive and efficient way to generate and evaluate molecular analogs, SAMSON’s SMILES Manager offers an excellent solution for performing positional analogue scanning.
What Problem Does This Solve?
Positional analogue scanning addresses the need to evaluate the structural impact of substituting specific atoms or groups in a molecule. For instance, if you’re working with a lead compound, changing one group to a different functional group across several positions can help identify areas for further optimization. This is especially useful for tasks like improving binding affinity or preserving interactions with a target protein. Instead of manually creating and analyzing each possible analogue, the SMILES Manager allows you to automate much of this work.
How It Works: A Step-by-Step Overview
The SMILES Manager makes this process straightforward with an intuitive workflow. Here’s how you can get started:
1. Setting up the Starting Molecule
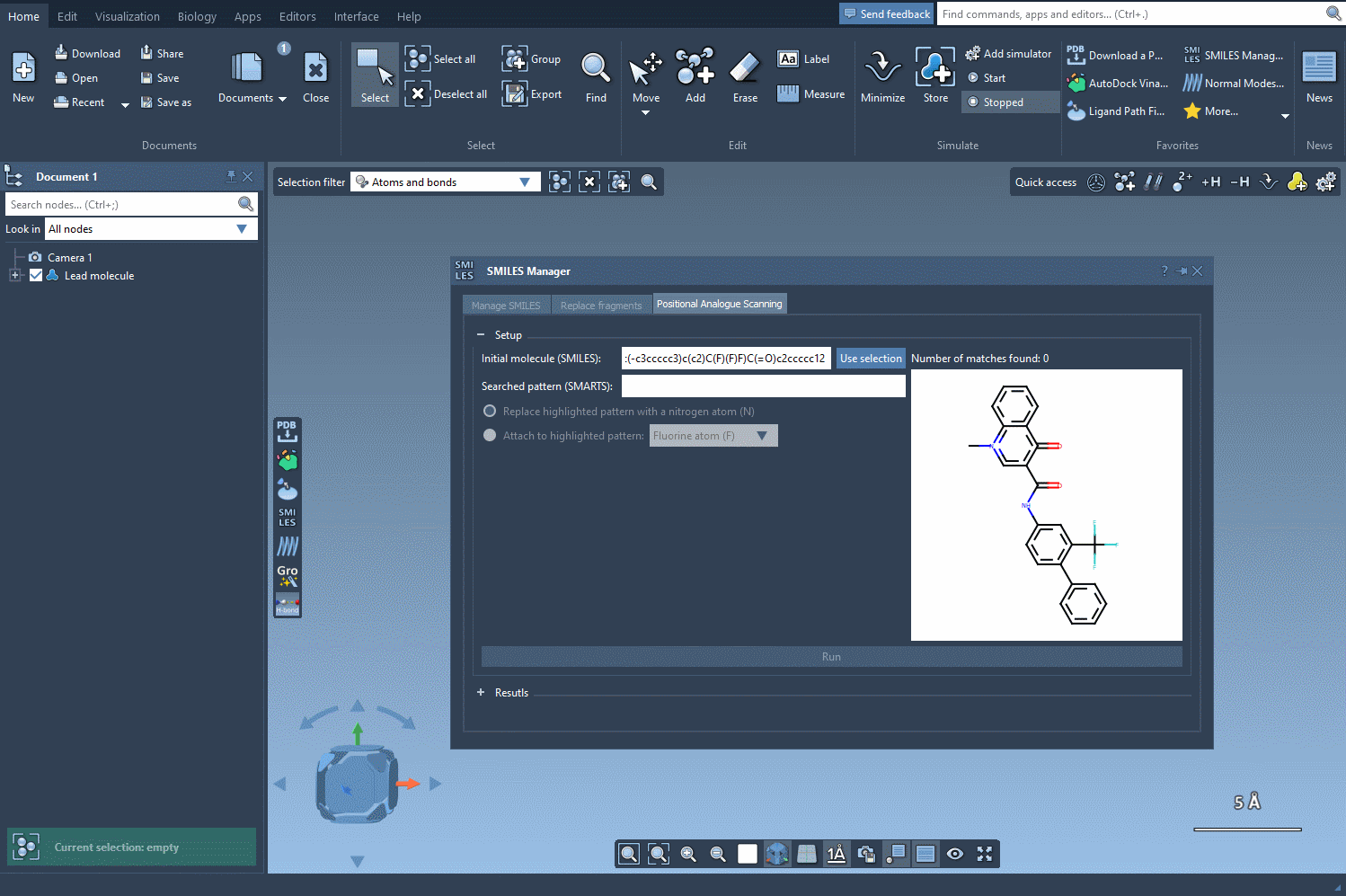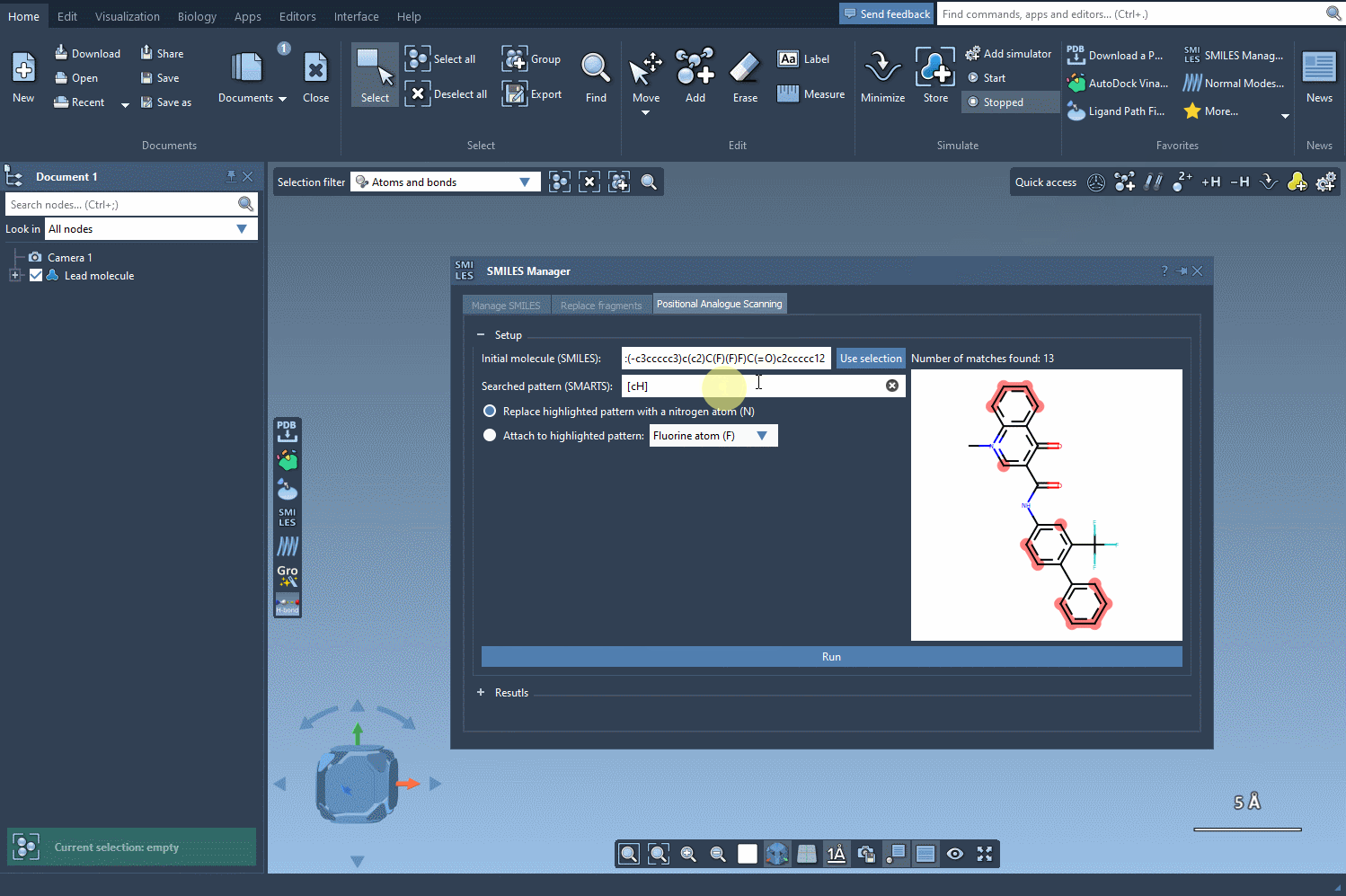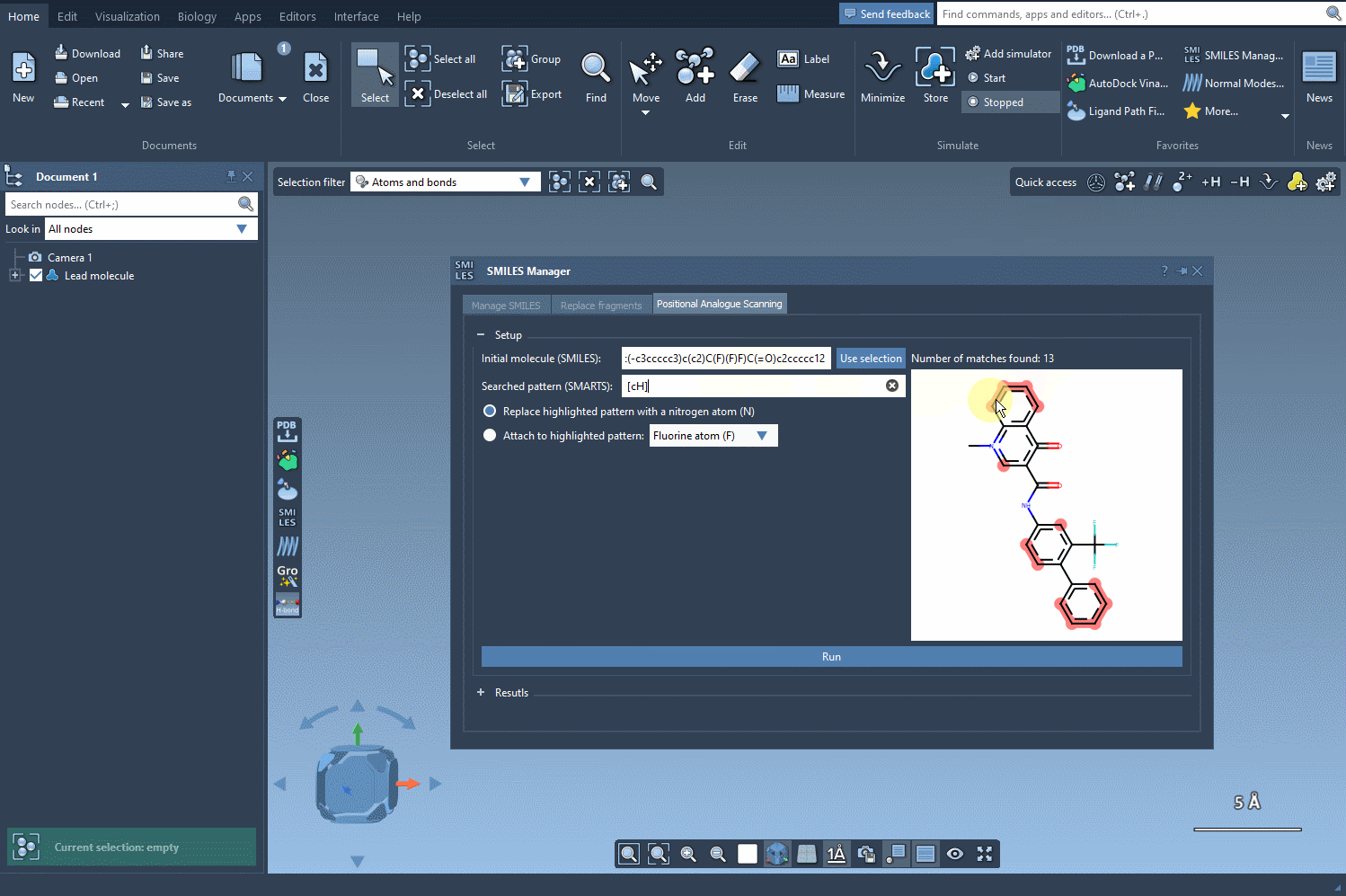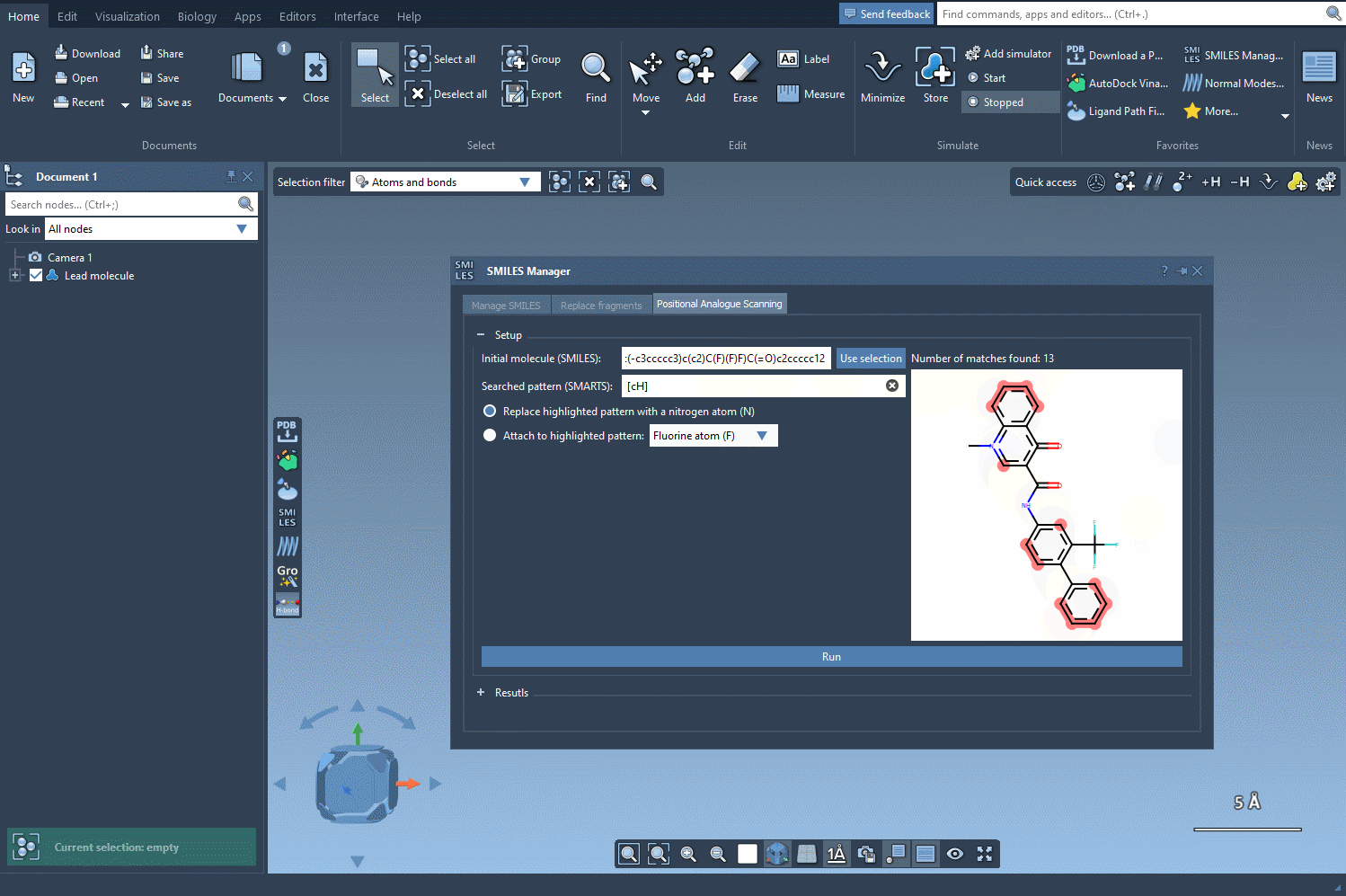
Before you begin, define the molecule you’d like to modify. You can either input the SMILES string directly or select a molecule you already have in your SAMSON workspace. Clicking on the Use selection button will initialize it immediately. Below is an example gif demonstrating this process:
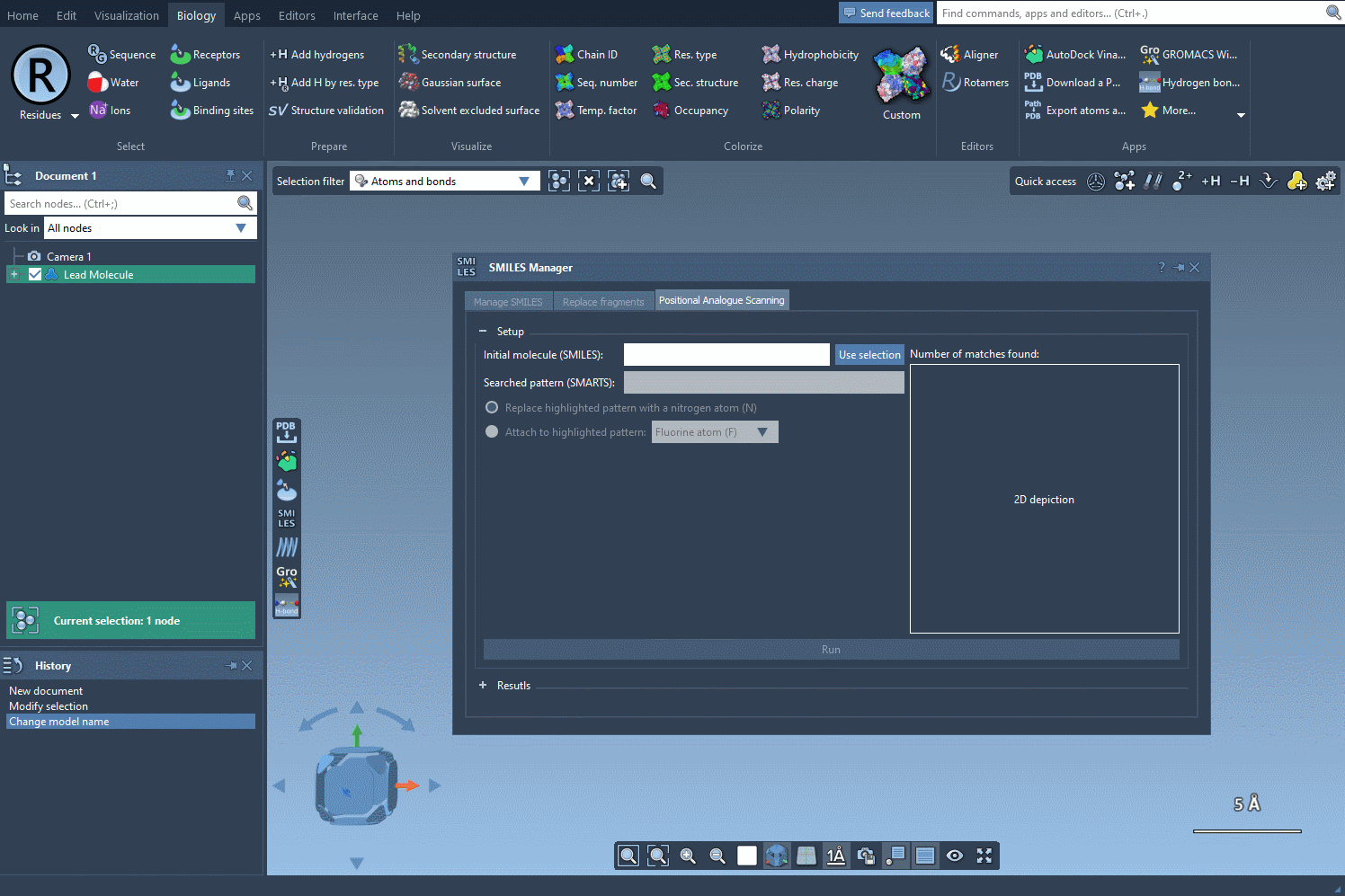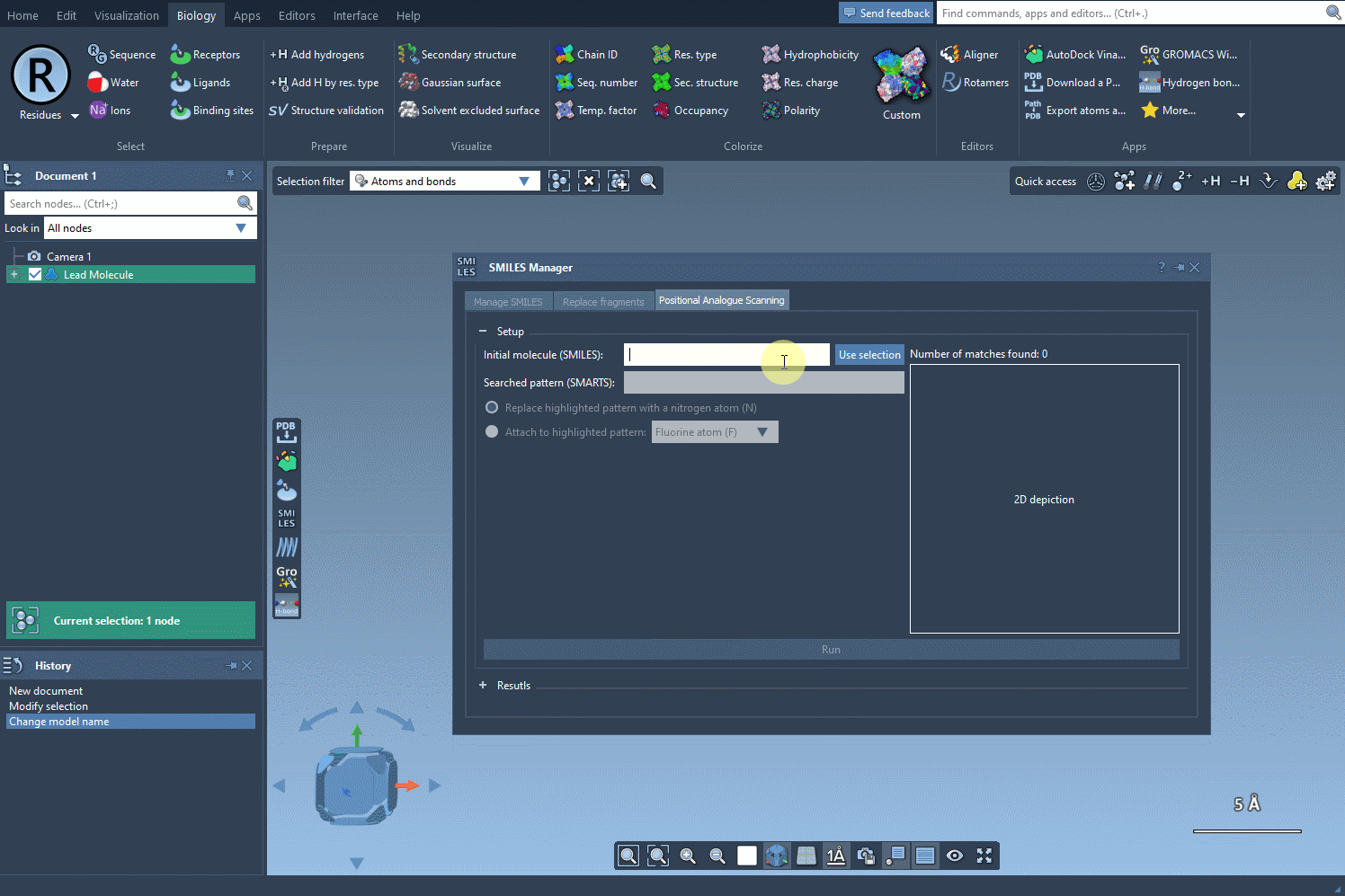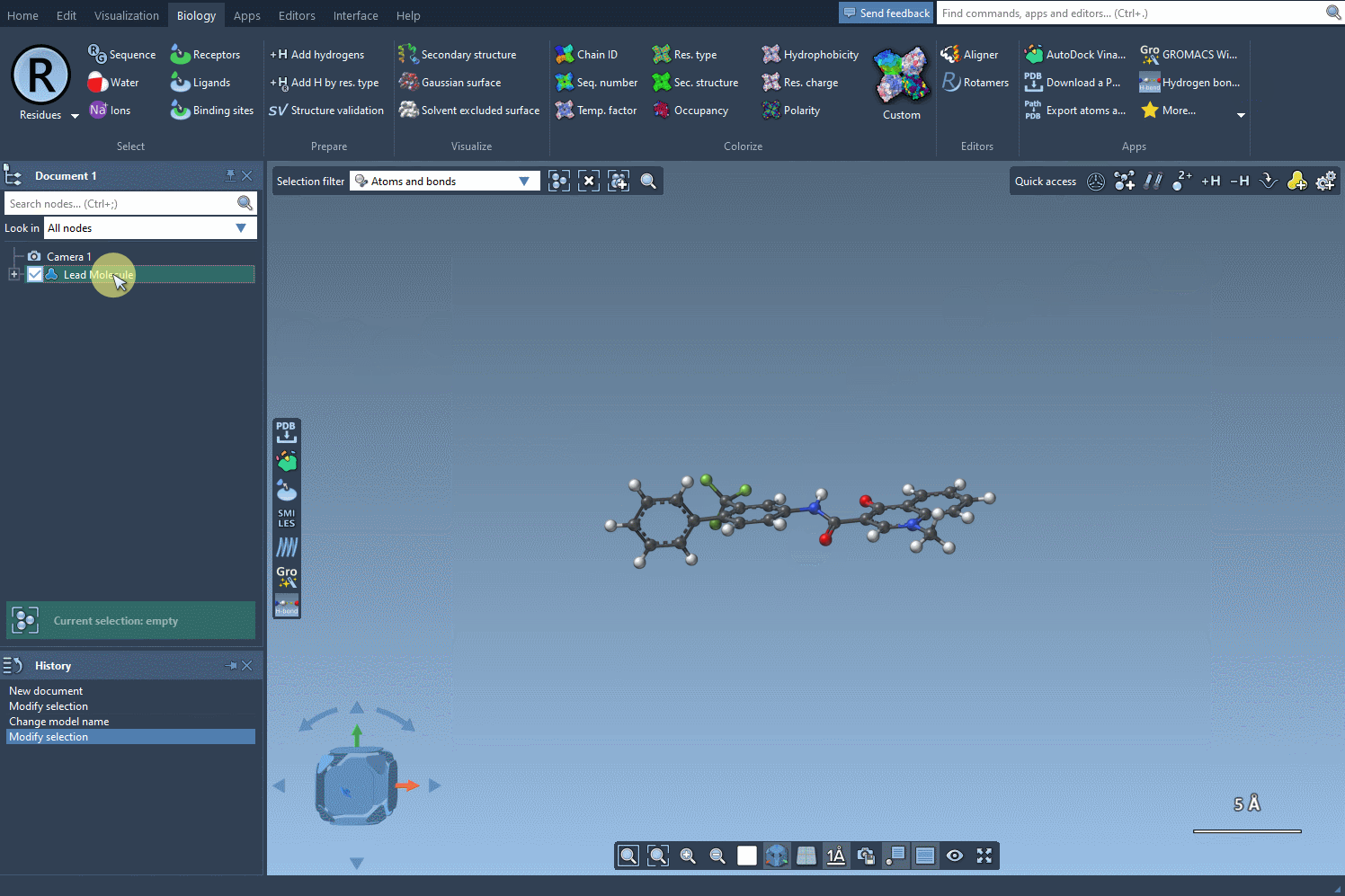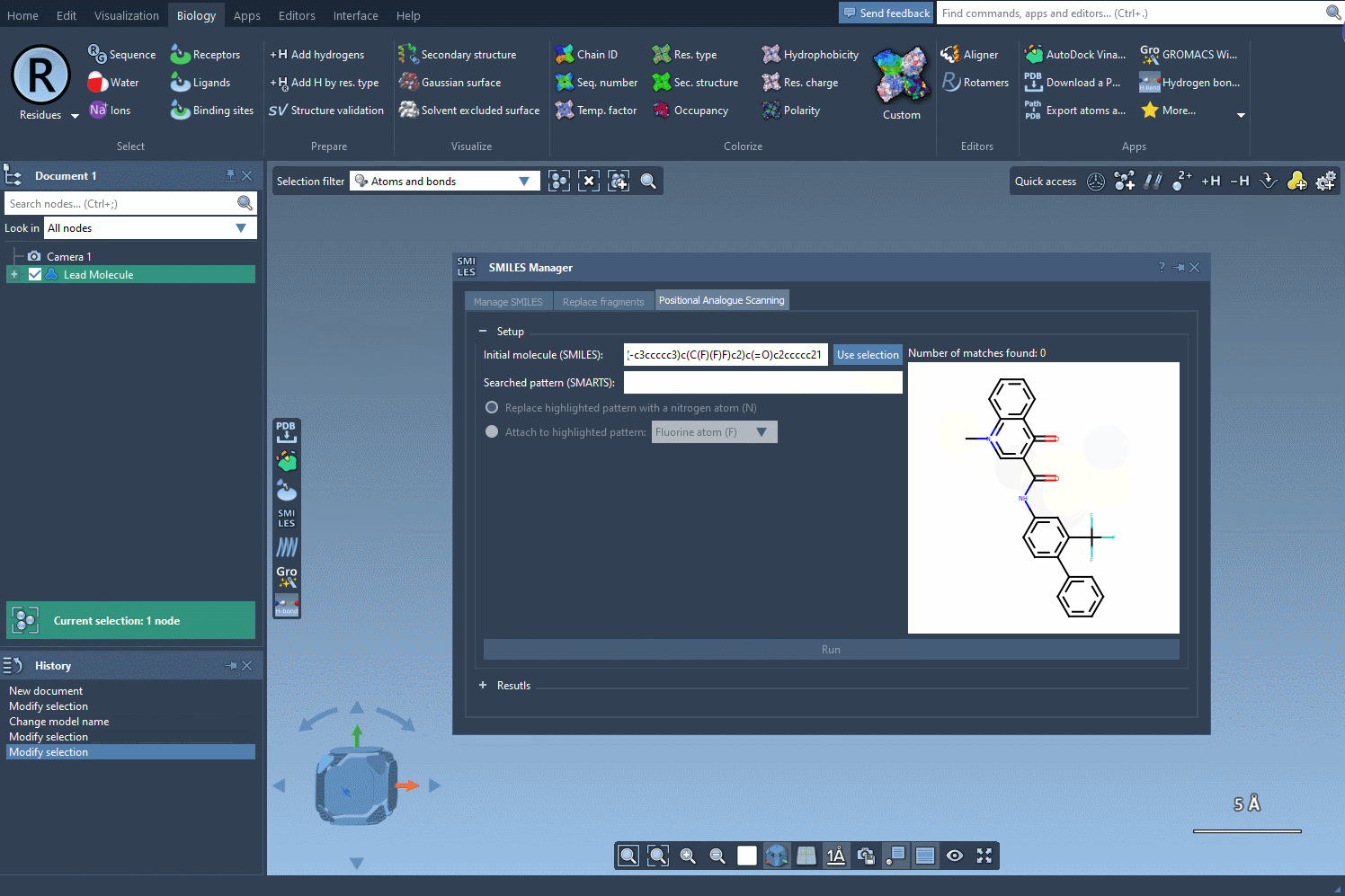
2. Defining the Replacement or Attachment Pattern
Once your molecule is set, the next step is to identify the pattern you’d like to change. For example, let’s say you want to replace aromatic hydrogens in your molecule. You can input the corresponding SMARTS code, [cH], to locate all the matching patterns. The SMILES Manager will highlight the targeted positions in your molecule:
3. Running the Positional Scanning
After defining the pattern, you can specify how you’d like to modify it. Whether it’s replacing a pattern with a nitrogen atom (N) or attaching functional groups like fluorine (F) or a methyl group (CH3), the interface allows for easy customization. Simply click Run to generate all possible analogs, which will appear in a dedicated results section. It doesn’t get much simpler than this!
Customizing and Analyzing Results
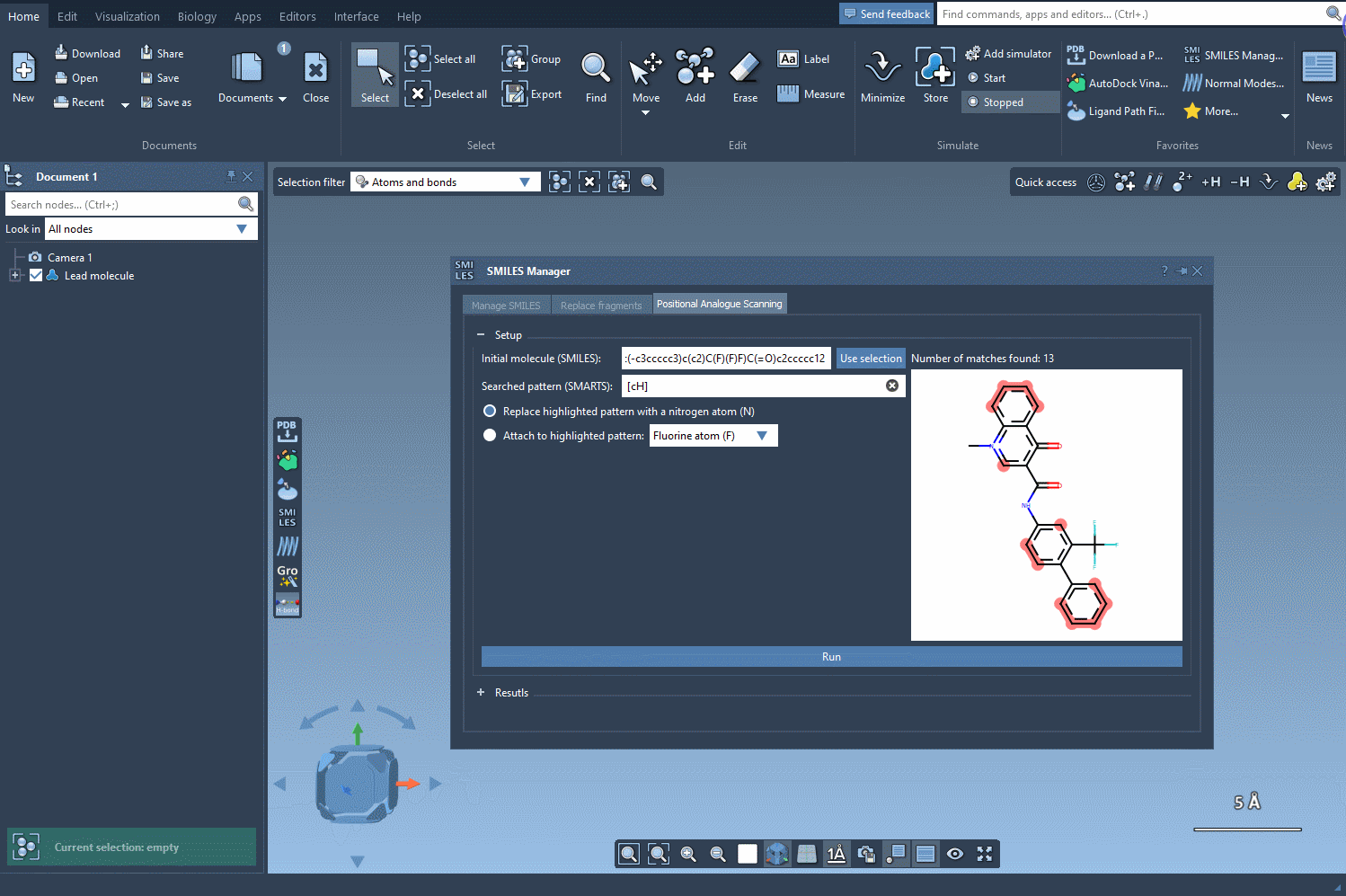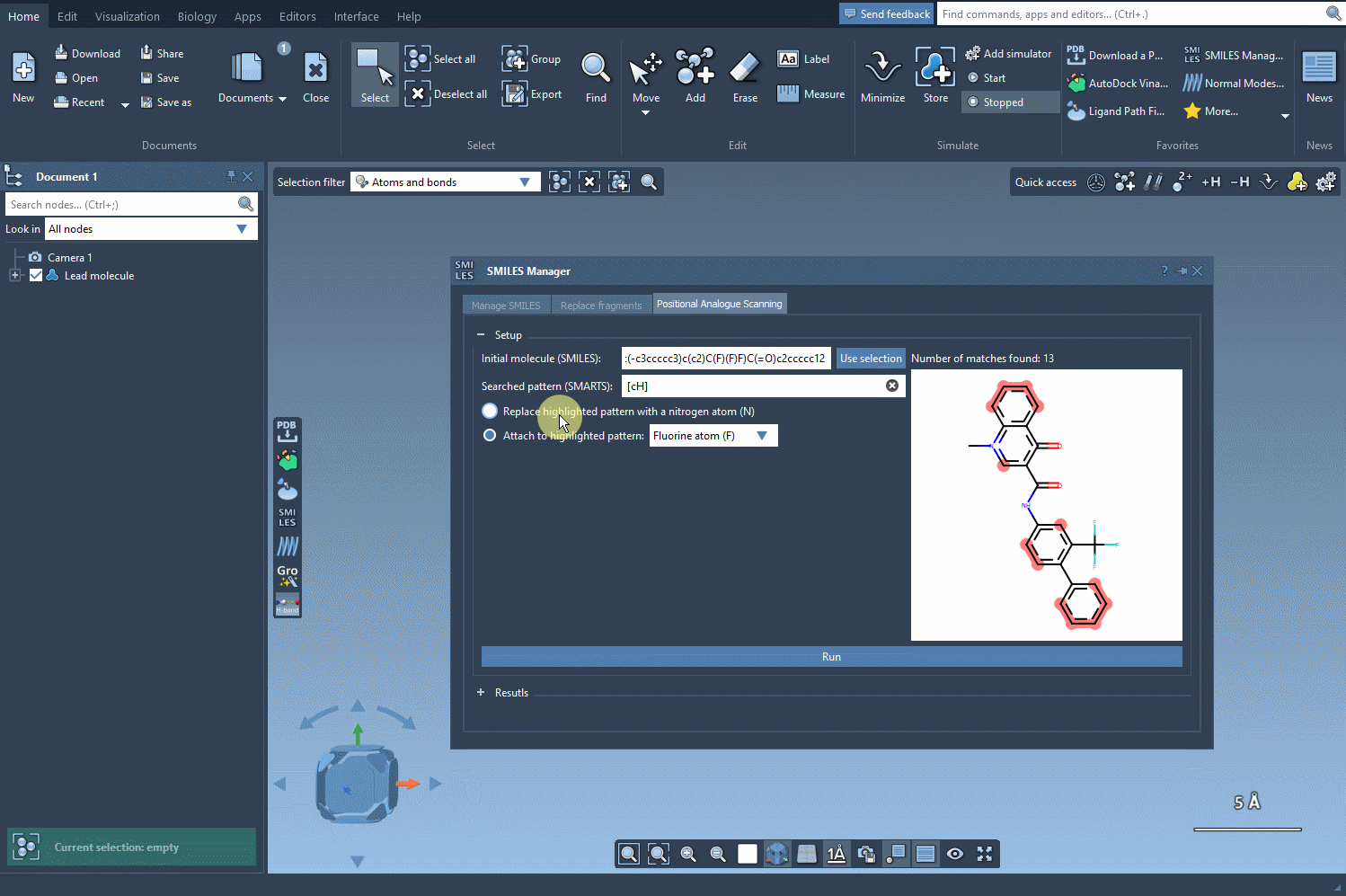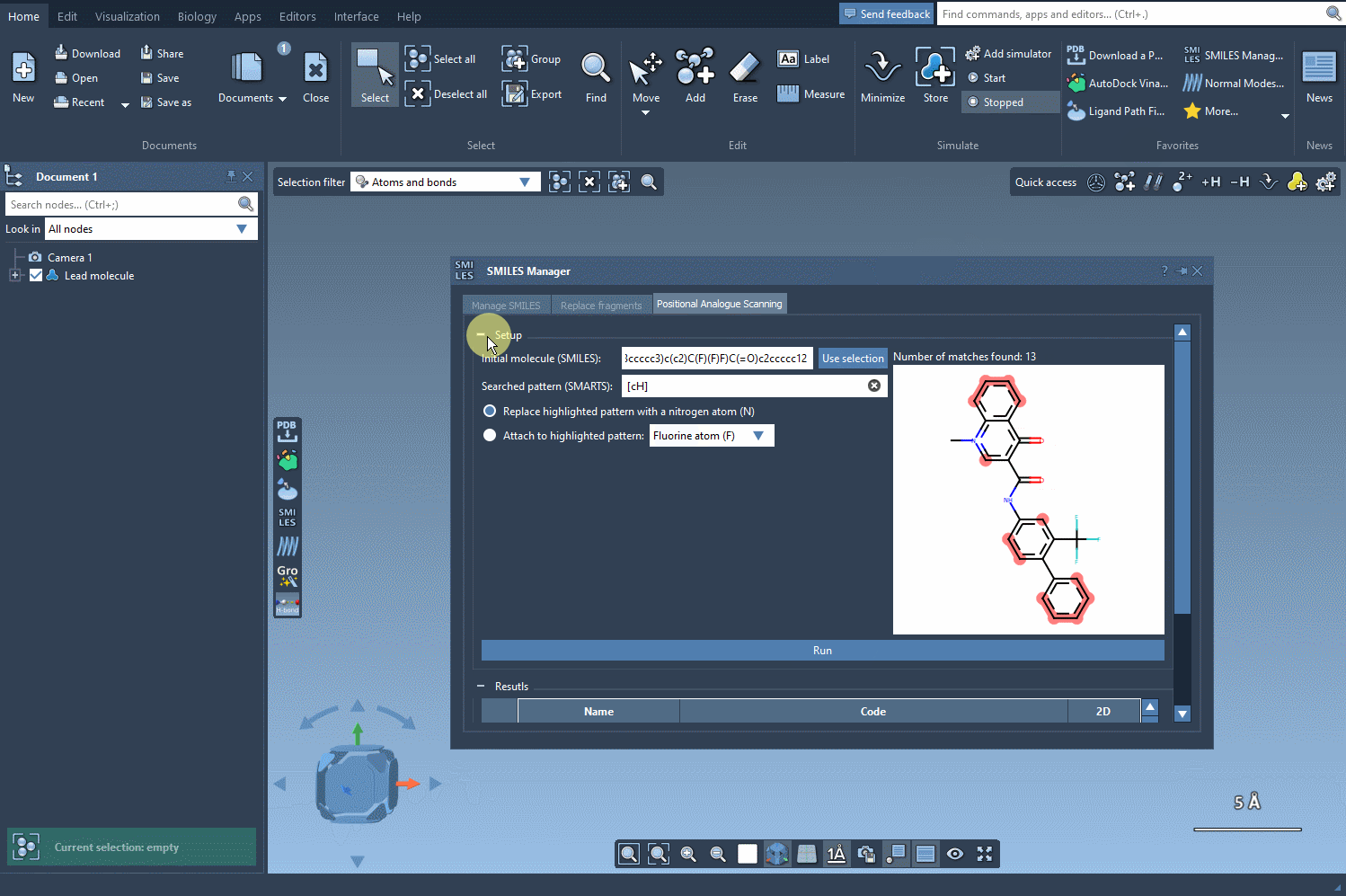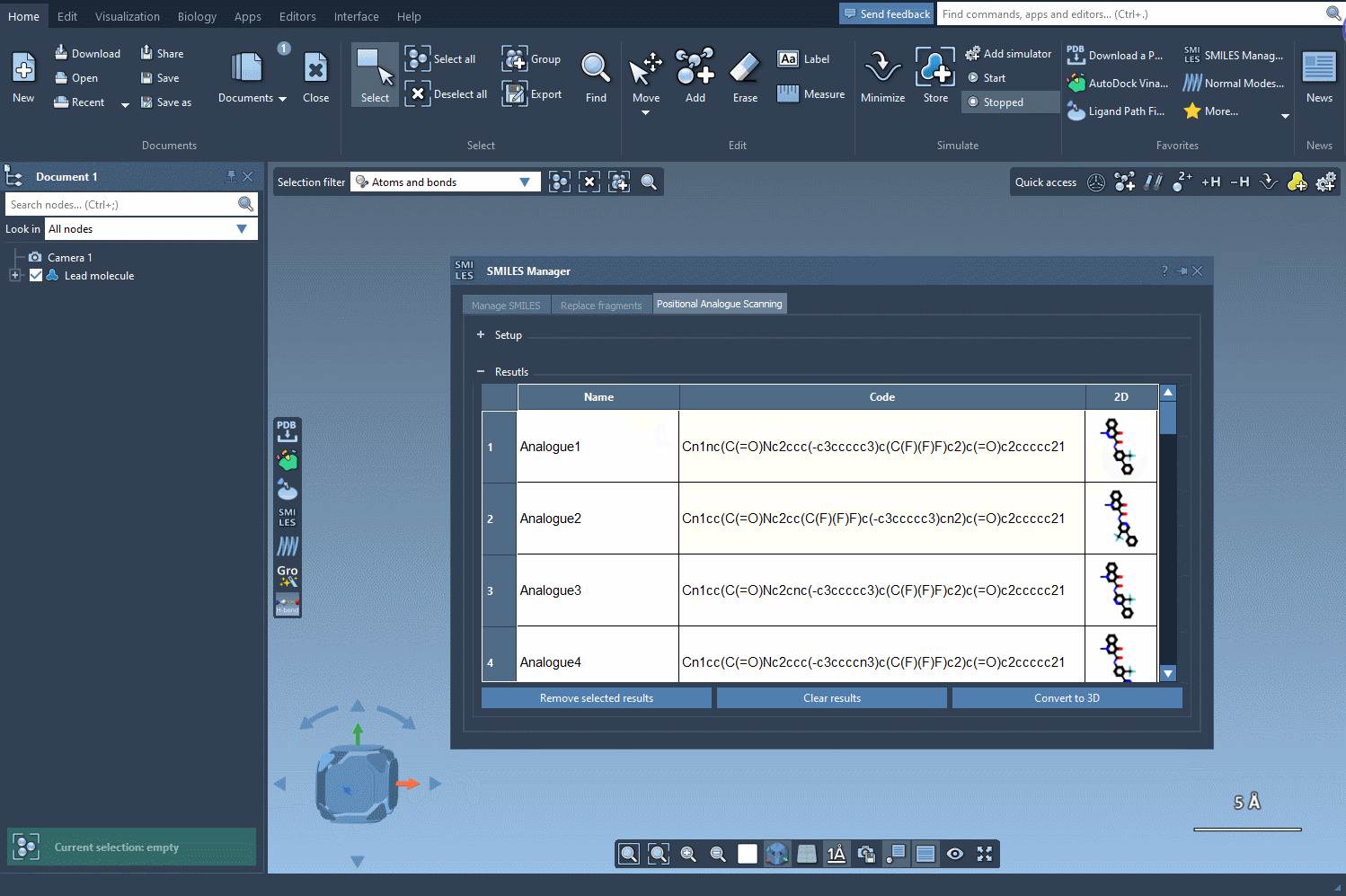
The results section offers several ways to customize and analyze your data:
- Editing: Double-click the SMILES code or name of an analogue to make changes.
- Visualizing 2D Structures: Open larger 2D depictions of your analogs to better analyze their structure.
- Generating 3D Models: Right-click an analogue to immediately convert it to a 3D structure for further study.
- Removing Analogues: Select and remove unwanted results or clear the results table entirely.
Ready for Deeper Analysis?
Once you’ve generated and refined your analogs, you can take your molecular study even further in SAMSON by performing docking studies. Extensions like Autodock Vina Extended allow you to evaluate how structural modifications influence binding interactions with target proteins. This can guide your decisions towards designing better compounds.
Conclusion
The SMILES Manager in SAMSON streamlines positional analogue scanning, saving you time and effort in generating and evaluating molecular analogs. Whether you’re focused on optimizing binding affinities or exploring new structure-activity relationships, this tool provides a powerful approach to iteratively improve your designs.
To get a full walkthrough of the SMILES Manager and its features, visit the official documentation at this link.
Note: SAMSON and all SAMSON Extensions are free for non-commercial use. Learn more and get started today by visiting SAMSON’s website.





