Molecular modelers often need an efficient way to generate and test new analogs of a molecule. Determining how even small changes to a molecule can impact its properties such as binding affinity or interactions with a target protein can be a time-consuming process without the right tools. Enter the SMILES Manager in SAMSON—an intuitive tool to perform Positional Analogue Scanning (PAS). This blog post focuses on a practical approach to getting started with PAS and automating the process of generating analogs efficiently.
Why Positional Analogue Scanning Matters
In molecular design, modifying specific parts of a molecule can significantly alter properties like solubility, binding energy, and interaction with proteins. PAS allows scientists to systematically scan positions within a molecule and evaluate substitutions to uncover analogs with better therapeutic, chemical, or physical properties. The SMILES Manager in SAMSON makes this traditionally tedious task faster and accessible within an integrative platform.
Getting Started with a Single Molecule
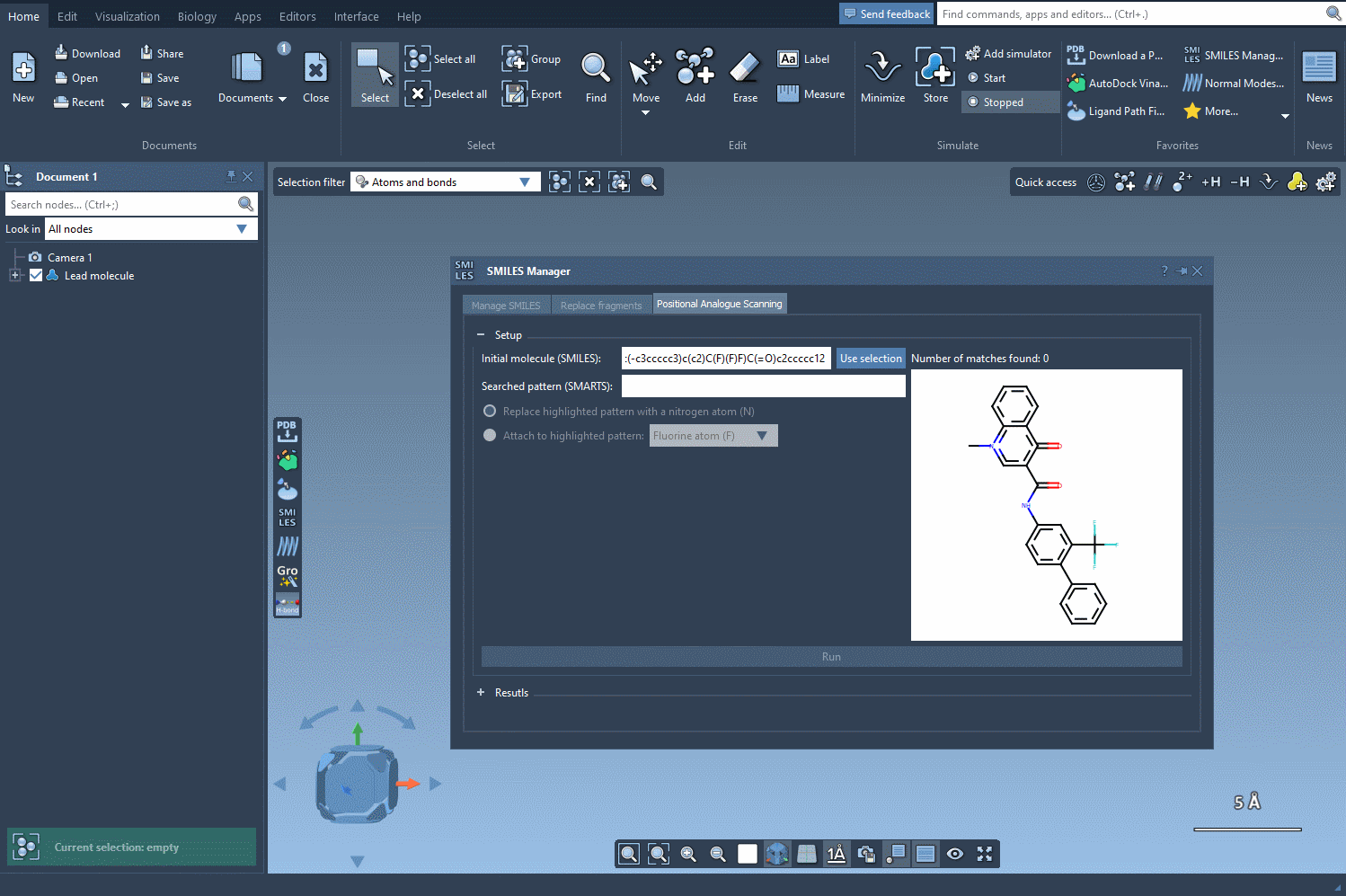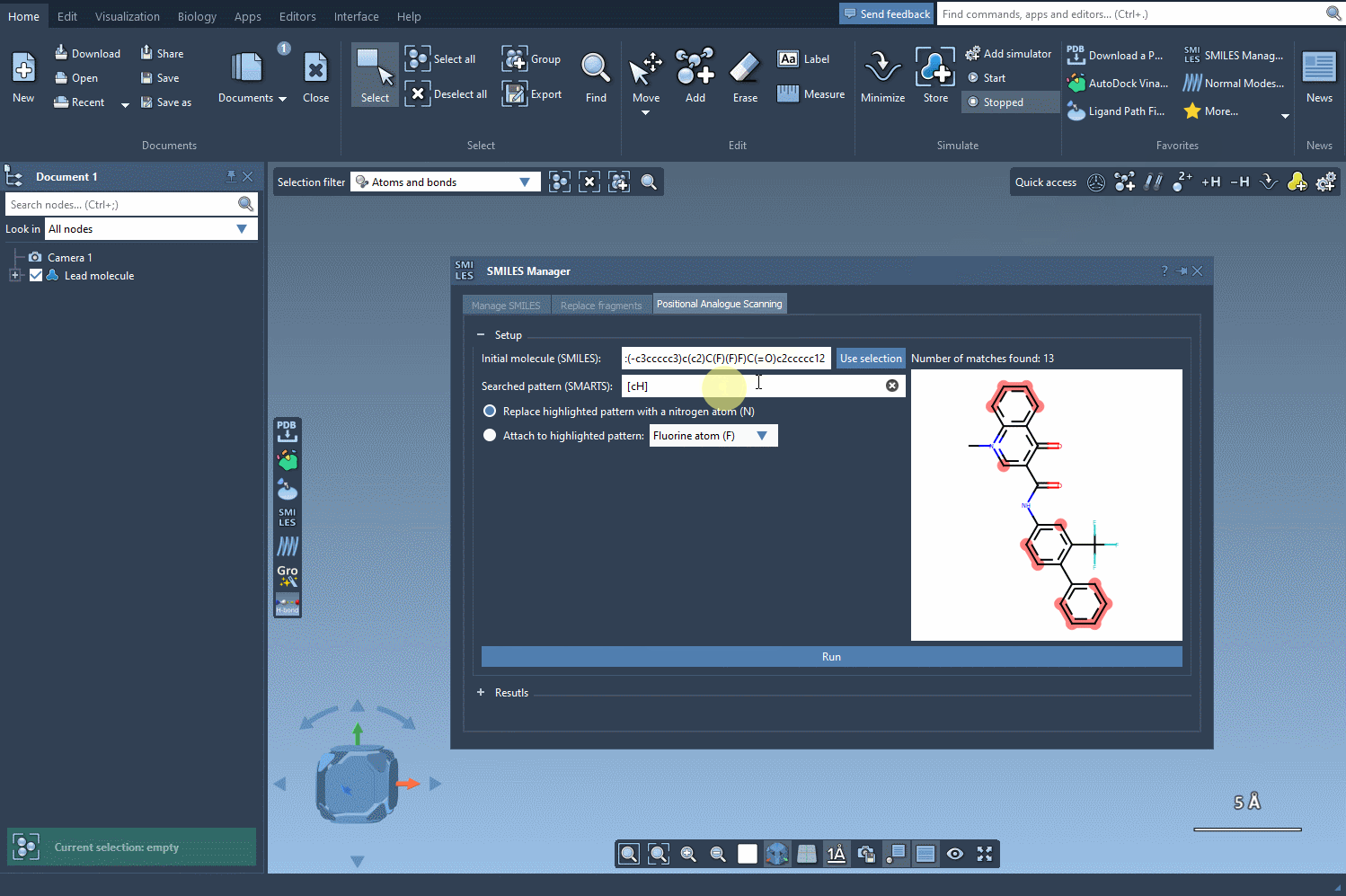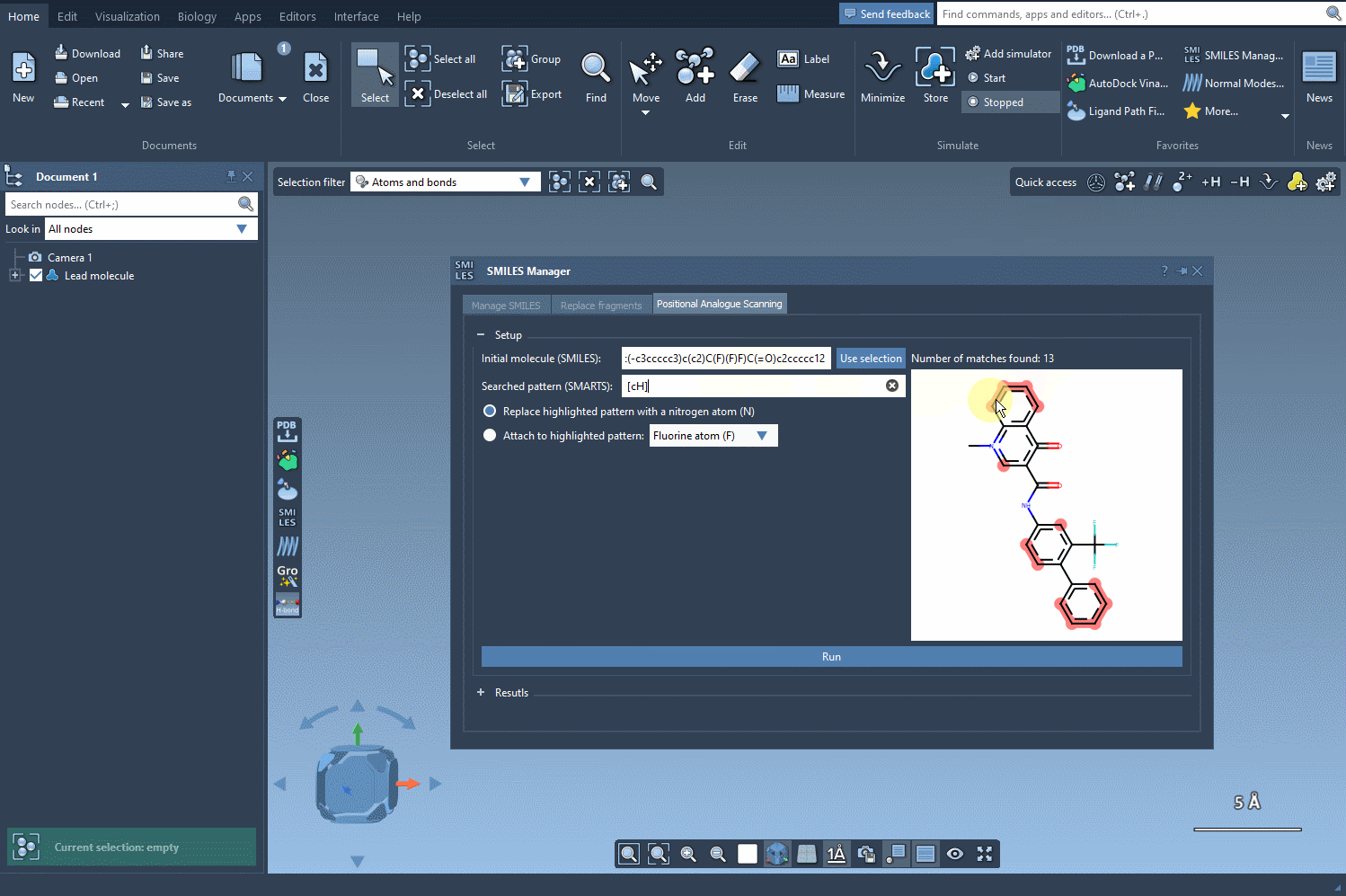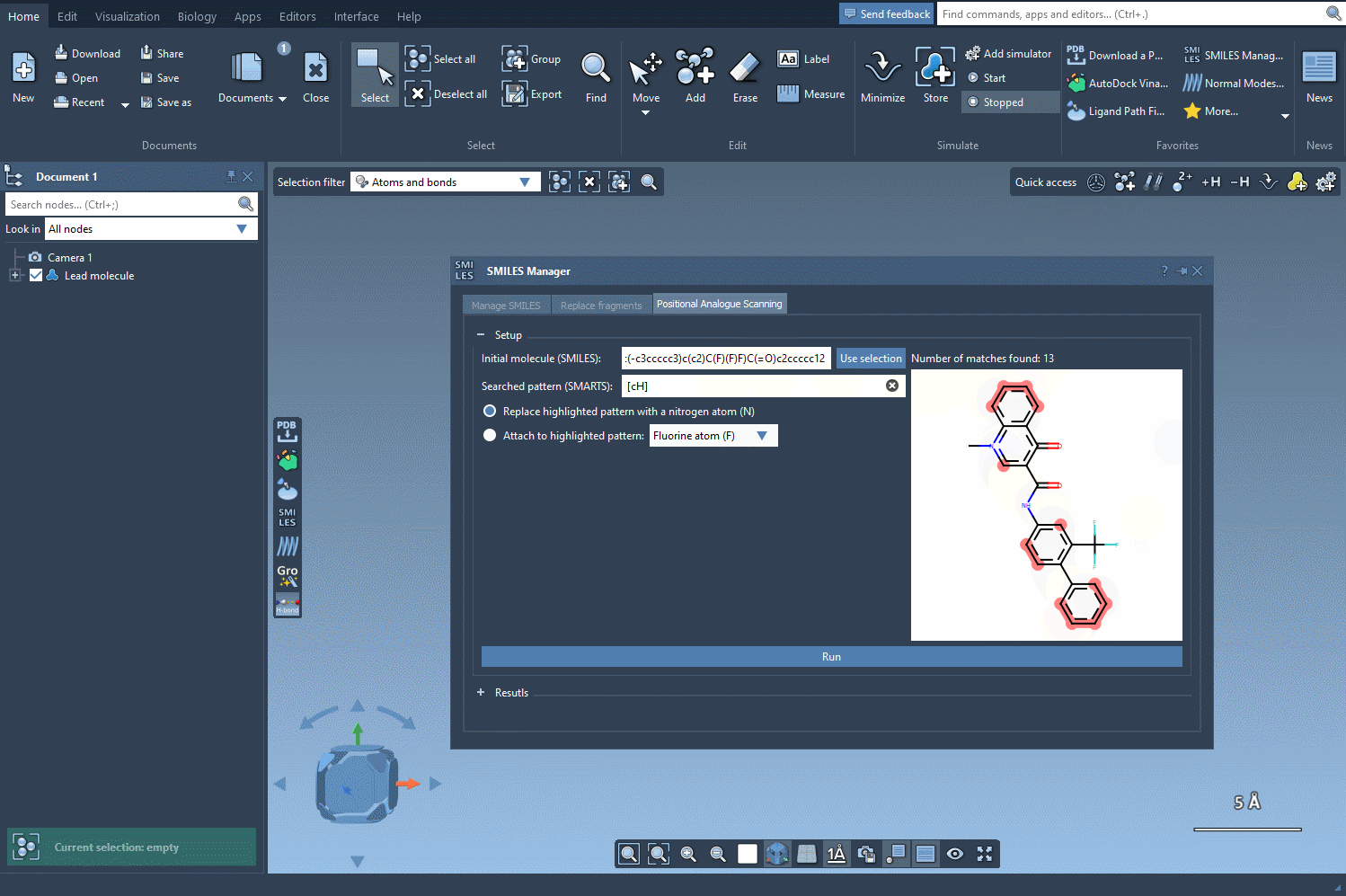
To begin scanning, you’ll first need to define your starting molecule. SMILES Manager provides two simple ways to do this:
- Directly enter the SMILES code of your molecule.
- Select the molecule from the SAMSON document and click the Use selection button.
The following visual provides a step-by-step demonstration of initializing the molecule selection:
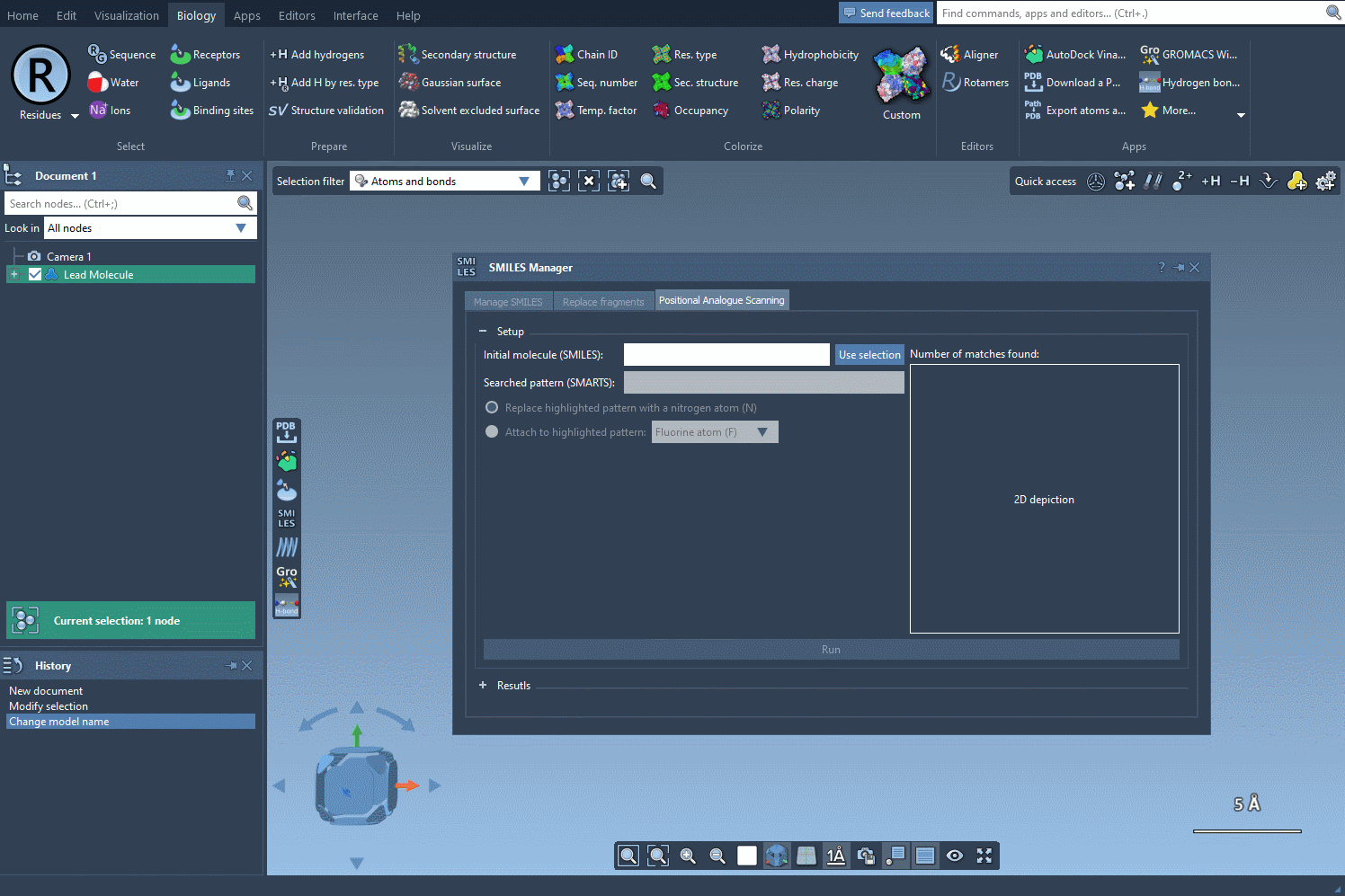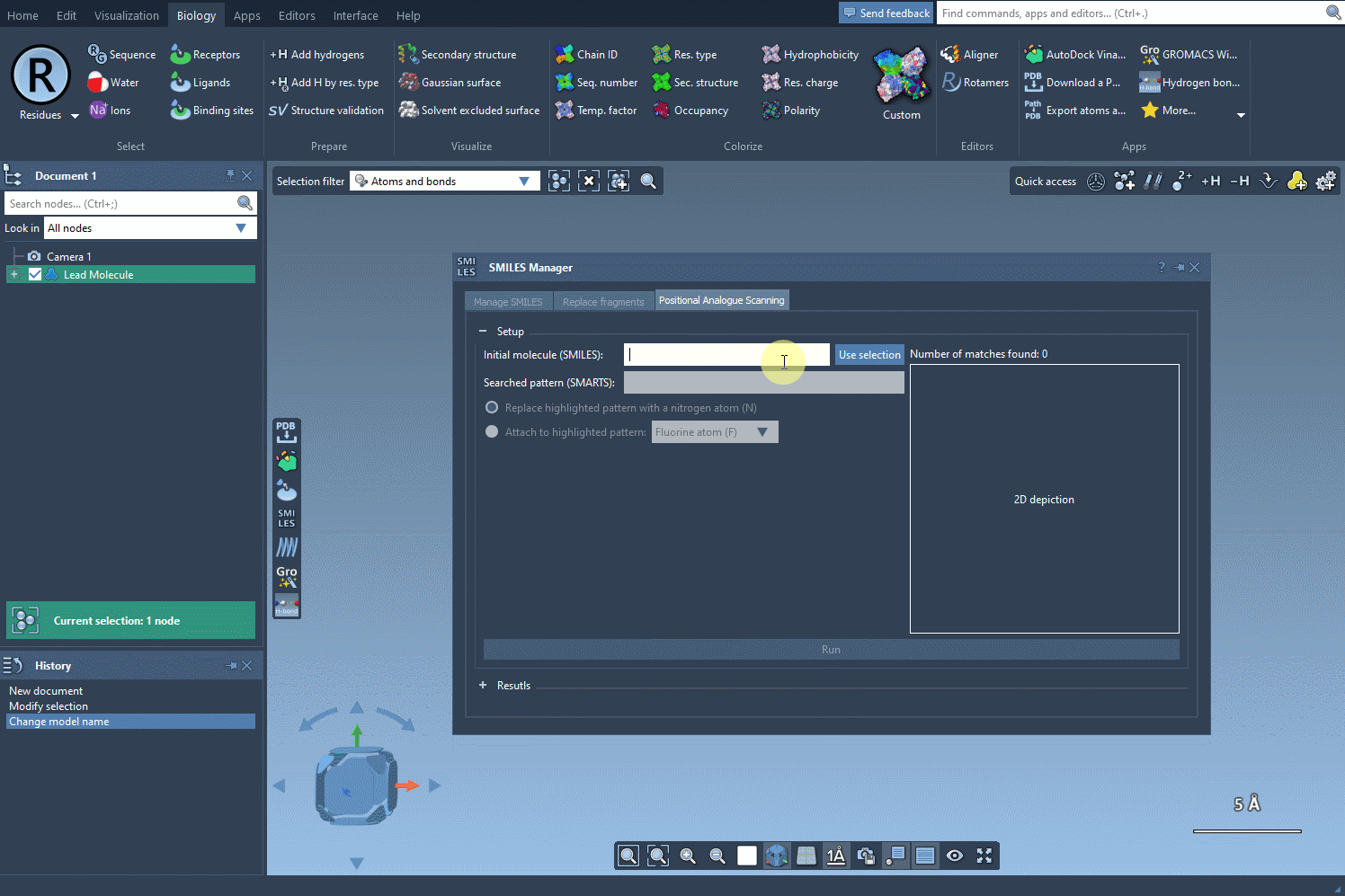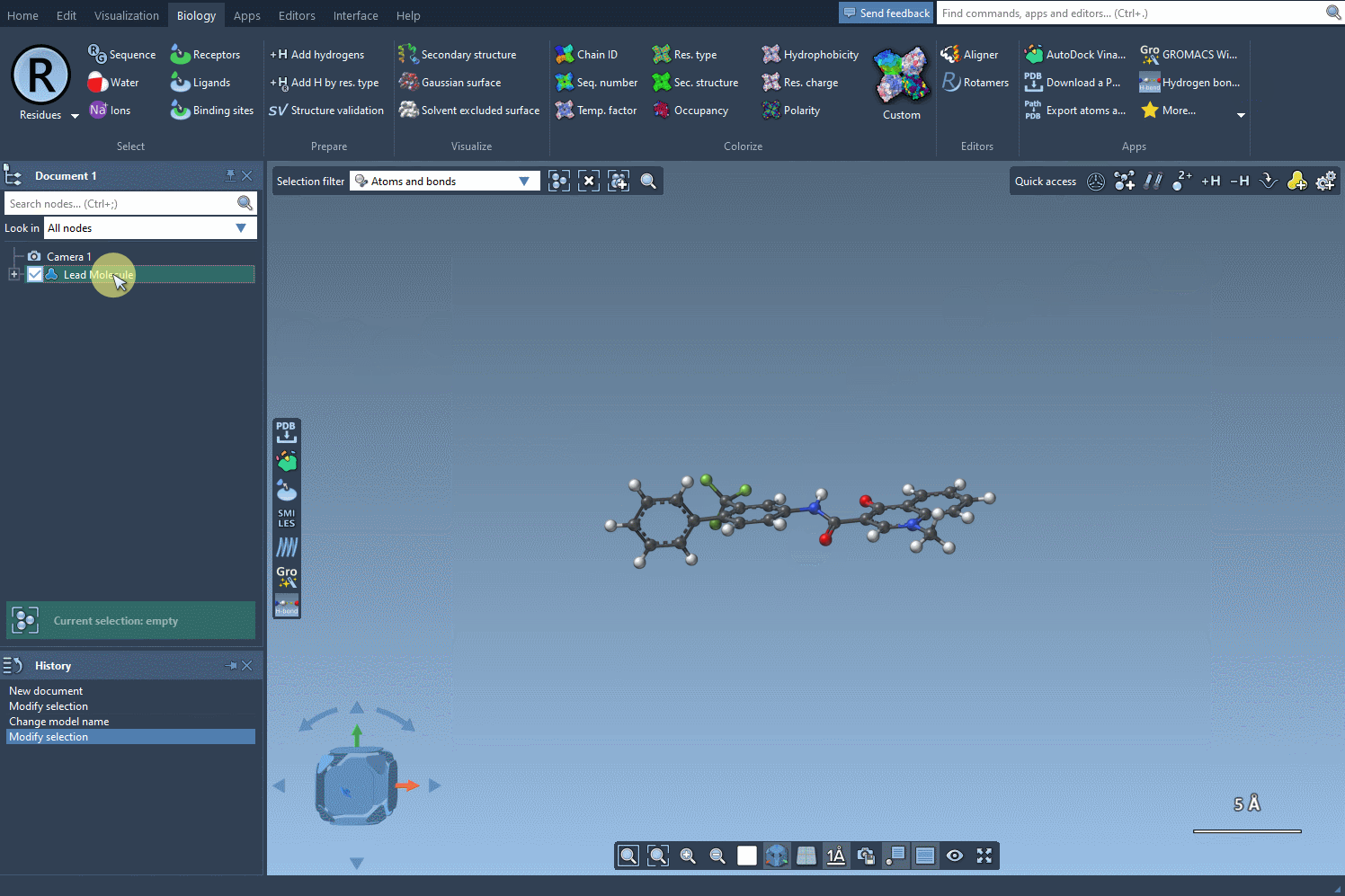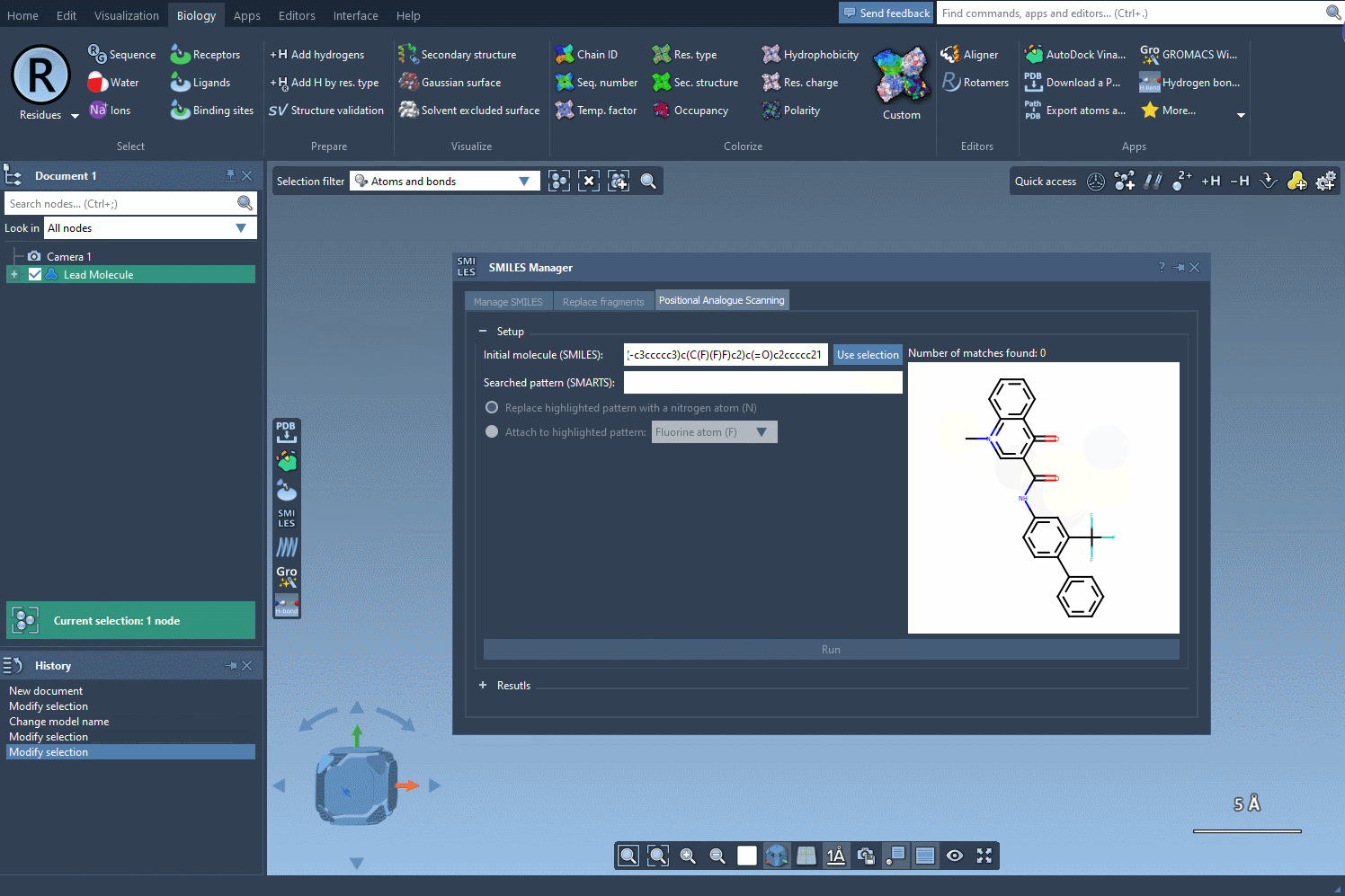
Focus Your Search with SMARTS
Once the molecule is loaded, you’ll want to define the specific pattern to modify. In this example, an aromatic carbon is selected as the target, identified using the [cH] SMARTS notation. The SMILES Manager conveniently highlights all occurrences of this pattern in your molecule, offering a straightforward way to see potential modification sites. Here’s a visual representation of this step in action:
Substitution Made Easy
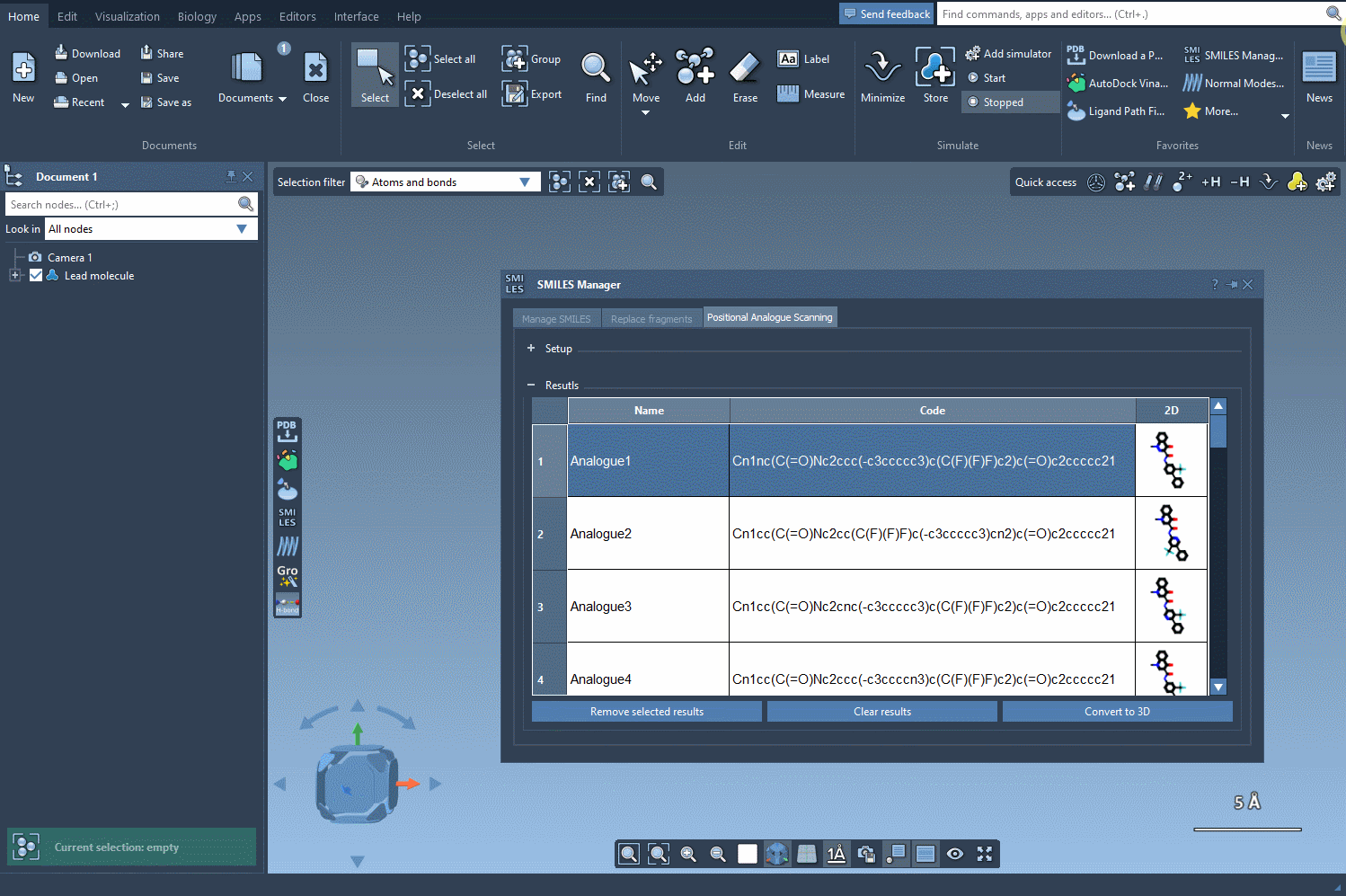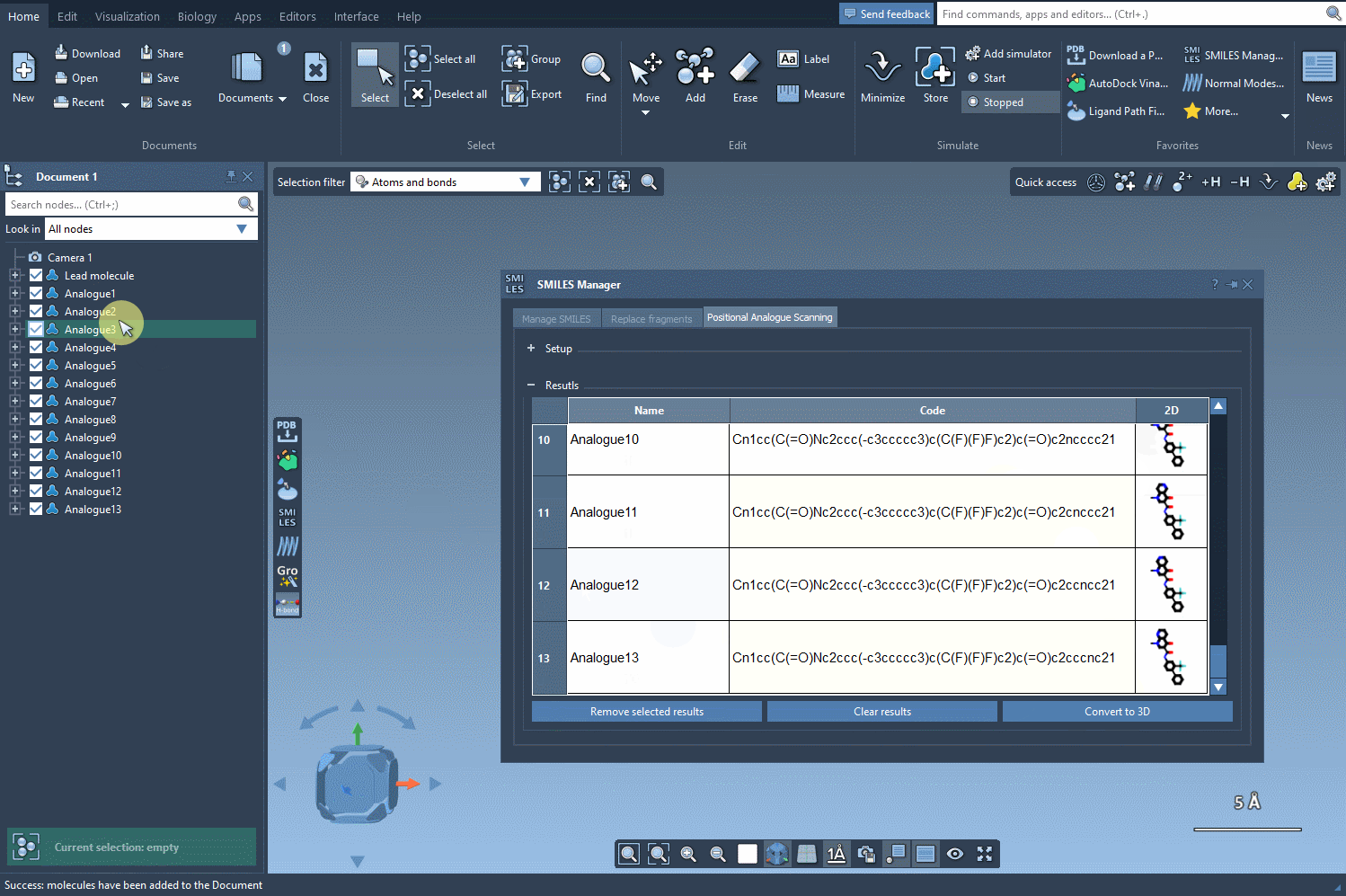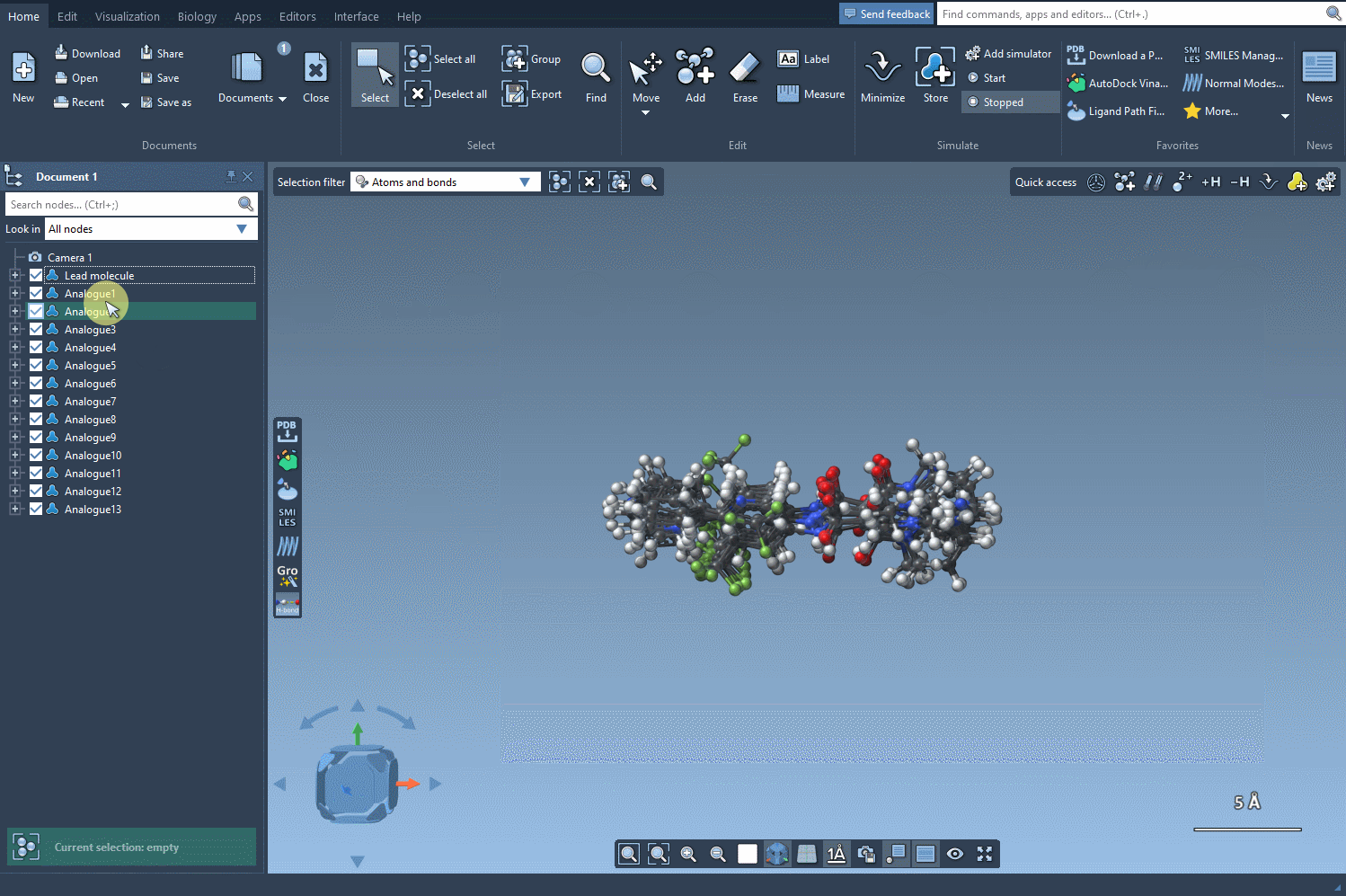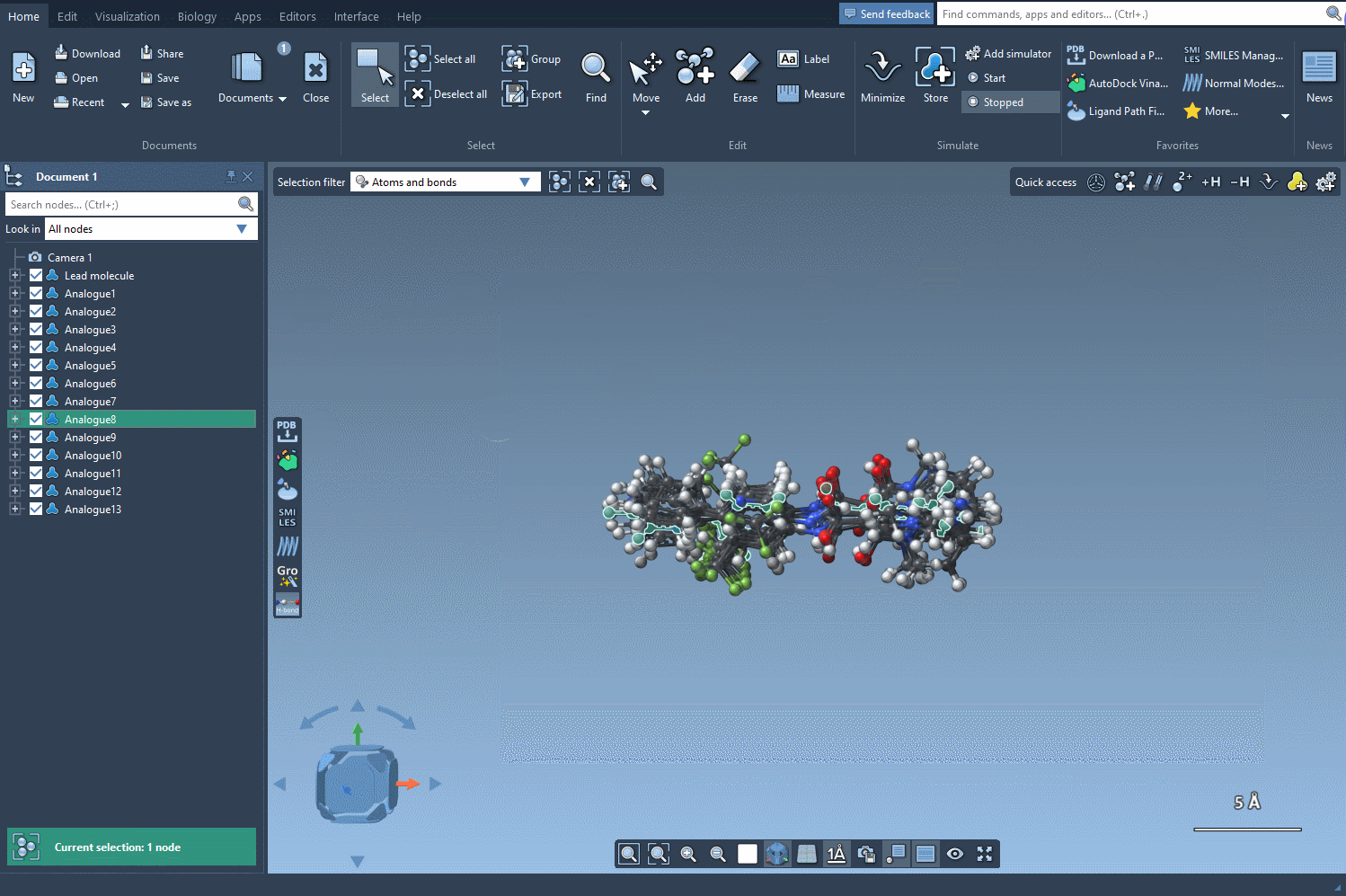
With the defined pattern, you can decide on the type of substitution—replace the pattern with a nitrogen atom (N), attach a fluorine atom (F), or add a methyl group (CH3). Once you’ve configured the replacement or addition, simply click Run to generate a library of analogs. The tool’s interface will then display the SMILES codes and 2D visualizations of the generated analogs. This step is efficient and minimizes manual labor:
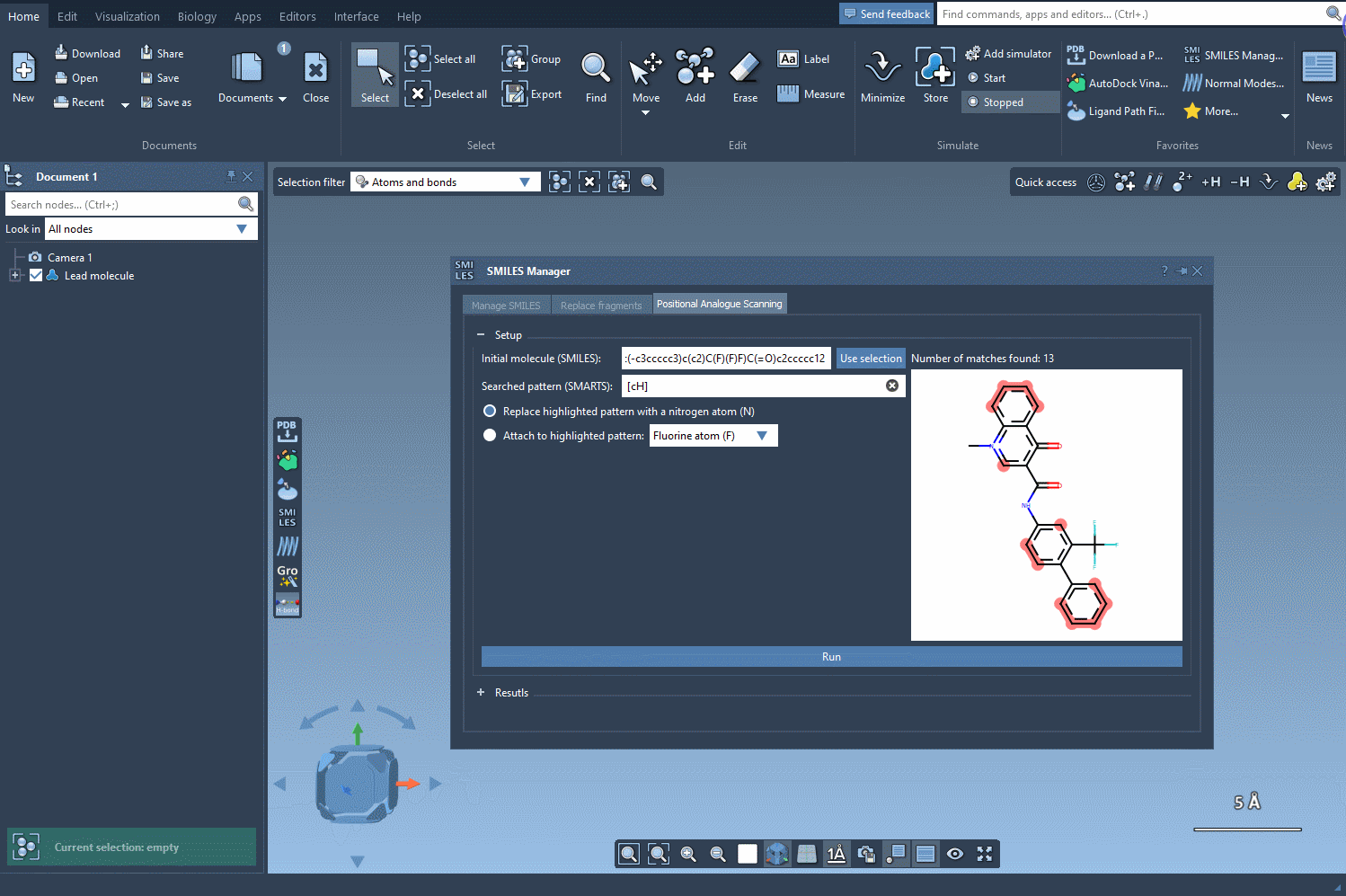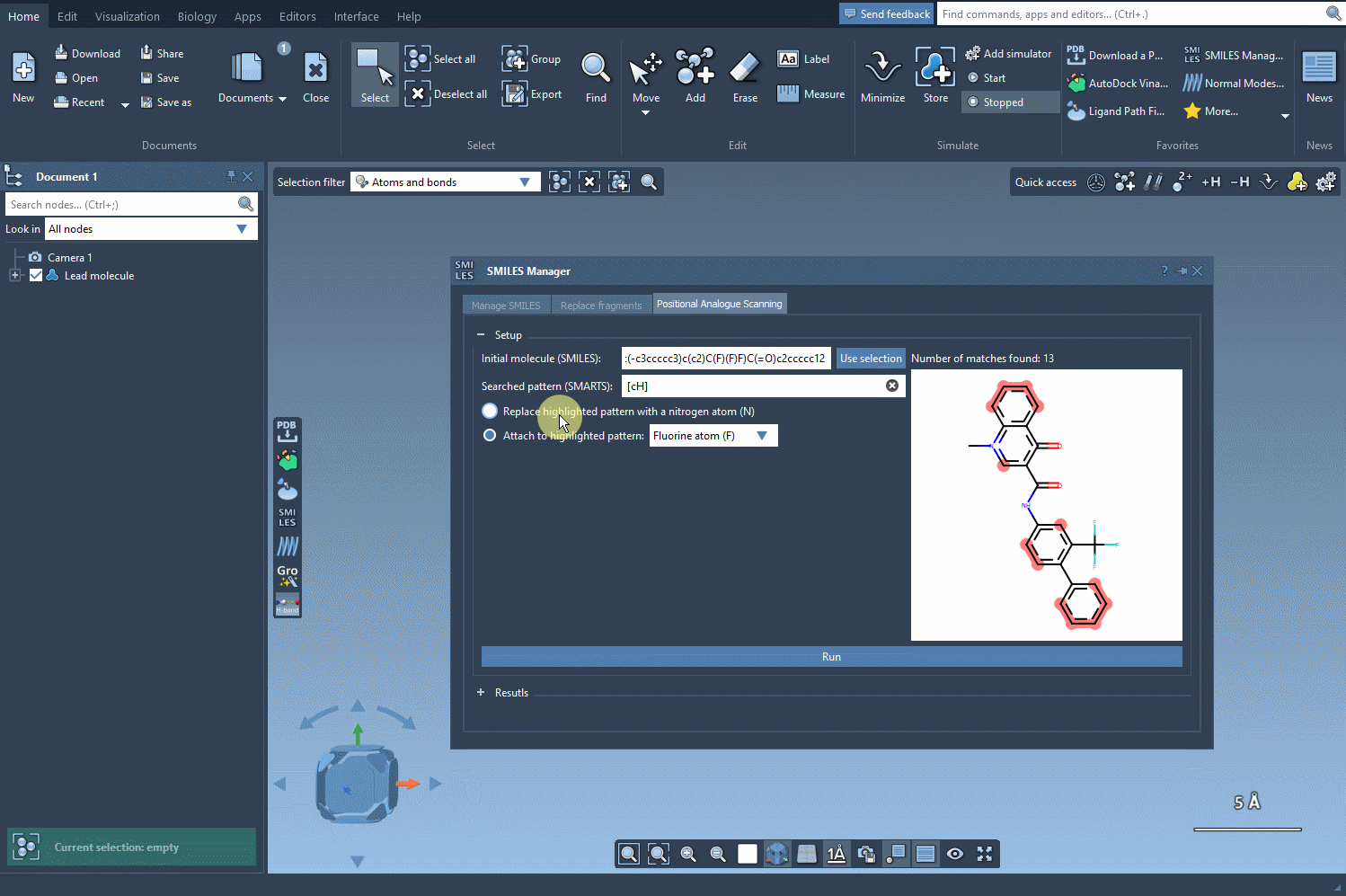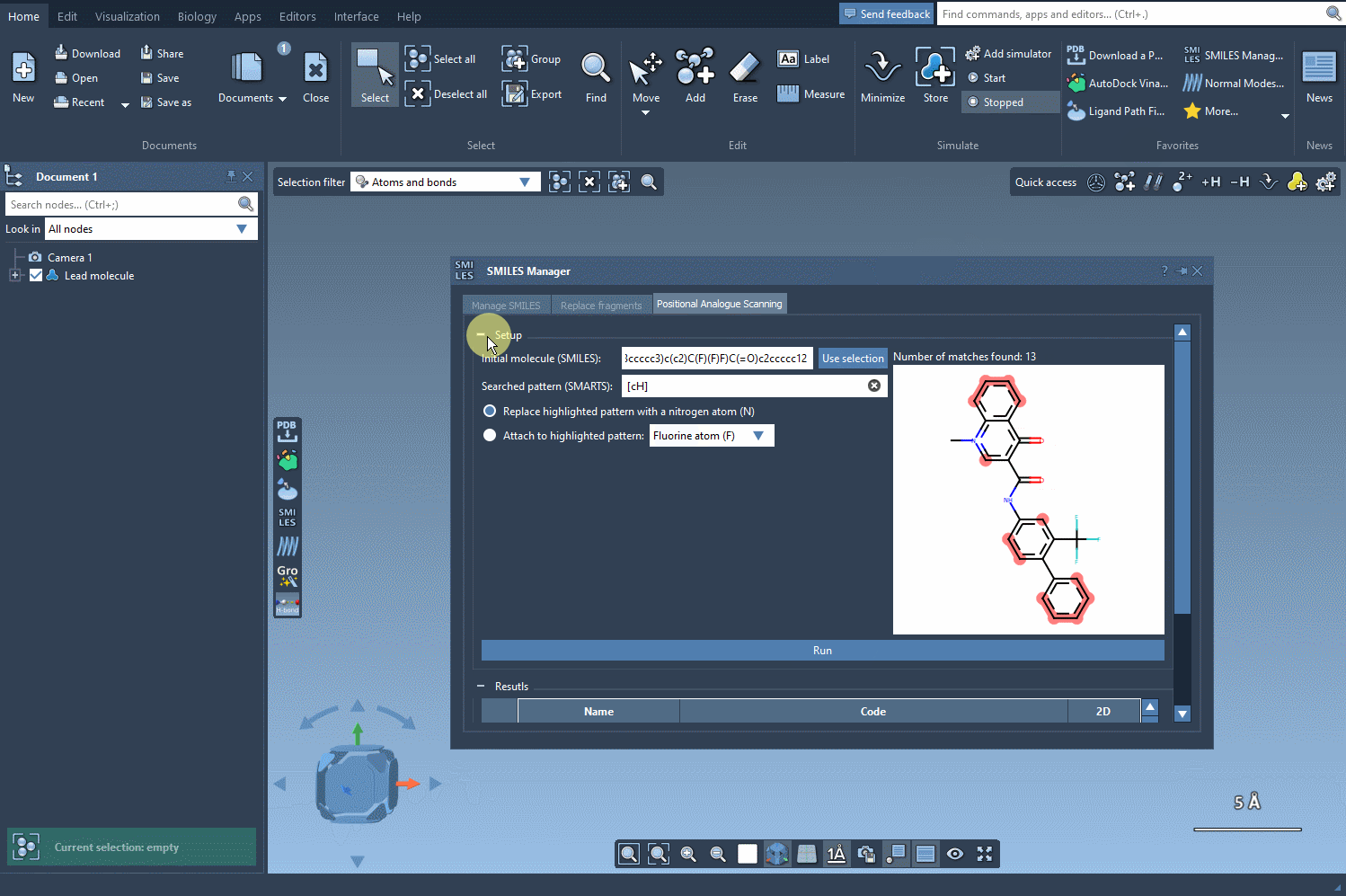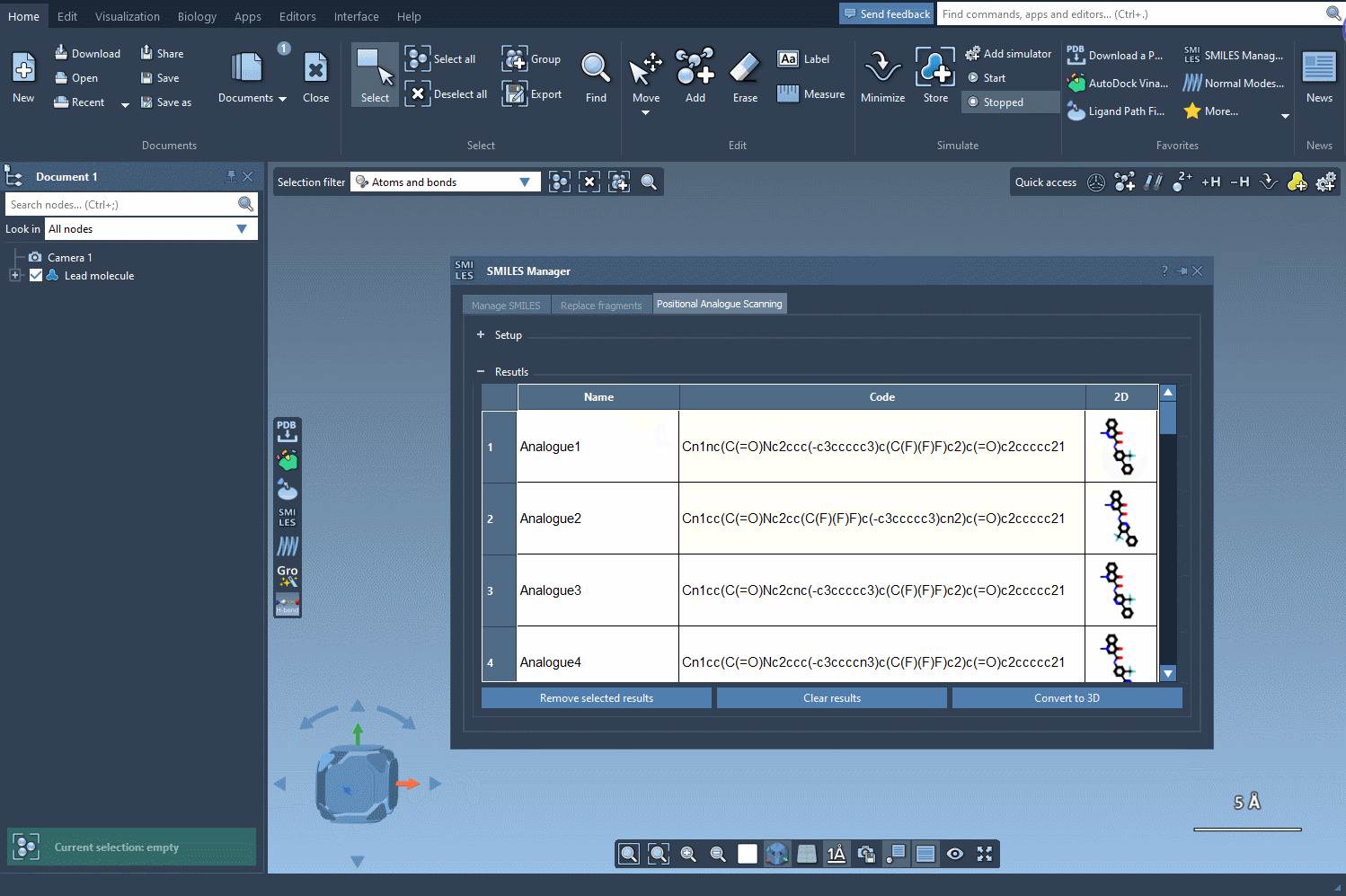
What Happens Next? Edit and Analyze
Exploring your results table offers even more flexibility:
- Modify analog names or SMILES codes directly by double-clicking.
- Open 2D depiction images in a larger window or toggle visibility.
- Generate 3D structures for a specific analog to prepare for further study.
3D analysis can be particularly useful when combined with SAMSON’s docking tools, such as the Autodock Vina Extended extension. This enables you to evaluate how modifications interact with a protein target, guiding you toward optimal molecule design.
Empower Your Molecular Design Workflow
Applying PAS through the SMILES Manager simplifies the molecular design process, boosting productivity and accuracy. Whether you are a seasoned molecular modeler or just beginning to explore molecule optimization, this tool will save you time and provide valuable insights into the behavior of your analogs.
Learn more about how to use Positional Analogue Scanning with the SMILES Manager by visiting the full documentation page.
Note: SAMSON and all SAMSON Extensions are free for non-commercial use. Start designing today by downloading SAMSON at samson-connect.net.





