For molecular modelers, understanding protein dynamics is crucial, especially when working with viral structures like the SARS-CoV-2 spike protein. The spike protein plays a key role in viral entry into human cells by transitioning from a closed state to an open state, where it binds to the ACE2 receptor. But analyzing these motions often requires heavy computational resources or specialized tools. Here, we explore how SAMSON facilitates such research with an efficient methodology to compute the SARS-CoV-2 spike’s opening motion.
The Challenge: Investigating a Key Movement
When studying SARS-CoV-2 and developing therapeutics, it’s essential to capture transitions between critical protein conformations accurately. The spike protein starts in a closed state and transitions to an open state, exposing its receptor-binding domain to the human ACE2 enzyme. However, obtaining an accurate trajectory for this motion, especially if residues differ between states, is non-trivial and time-intensive when using conventional methods.
SAMSON presents a highly efficient solution to calculate this motion through a pipeline involving two advanced modules: the As-Rigid-As-Possible (ARAP) Interpolation Path module and the Parallel Nudged Elastic Band (P-NEB) module. These modules accelerate the process significantly while maintaining precision, enabling researchers to investigate the underlying mechanics of the spike’s opening.
The Computational Pipeline
The computation of the SARS-CoV-2 spike’s opening motion begins with two known structures:
Since these structures vary in residues, the pipeline includes several preparatory steps:
- Adjust bond orders and add hydrogens for both structures using SAMSON’s Python scripting capabilities.
- Generate an interpolated path using the ARAP module. This module quickly processes transitions with atomic precision.
- Select a refined closed-state conformation from the ARAP trajectory and minimize it.
- Further refine the path with the P-NEB module. This step reduces energy barriers, yielding an optimized trajectory from the closed state to the open state.
In total, while running on a typical laptop, the ARAP module requires just under 30 seconds, while the P-NEB step takes about 15 minutes—making this approach highly accessible and efficient. Notably, the computed trajectories are visually inspected to ensure accuracy.
Visualization and Applications
Once computed, the spike’s motion can be visualized in various forms, allowing researchers to gain insights into its dynamics. Below is an example of the computed trajectory:
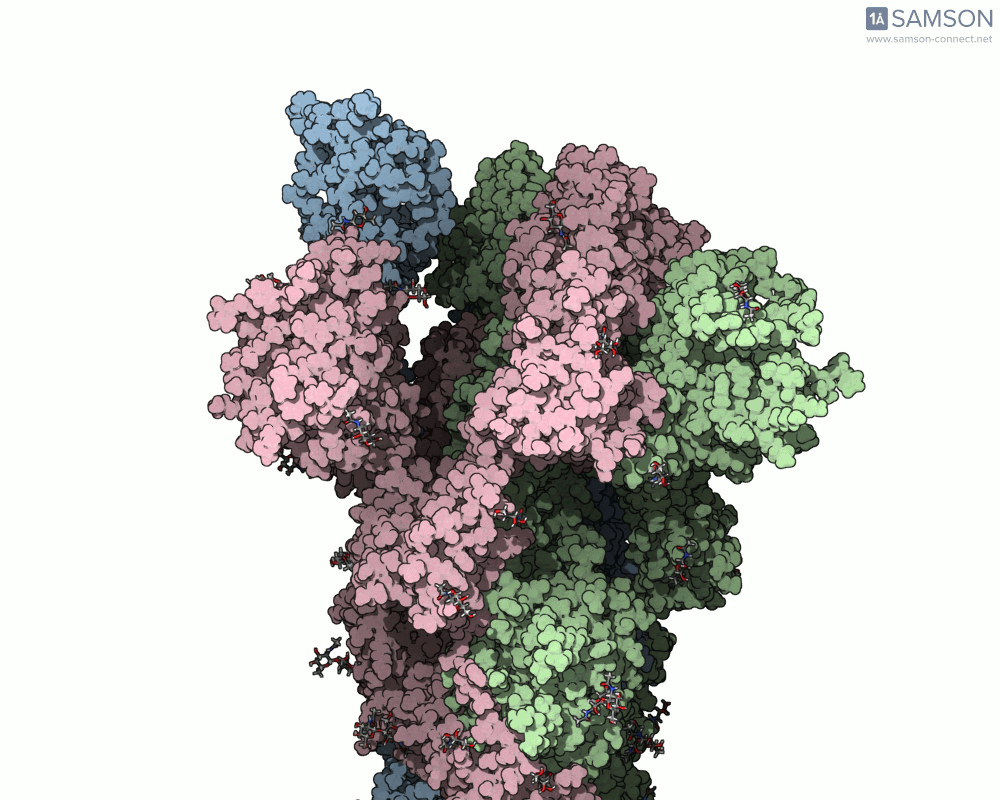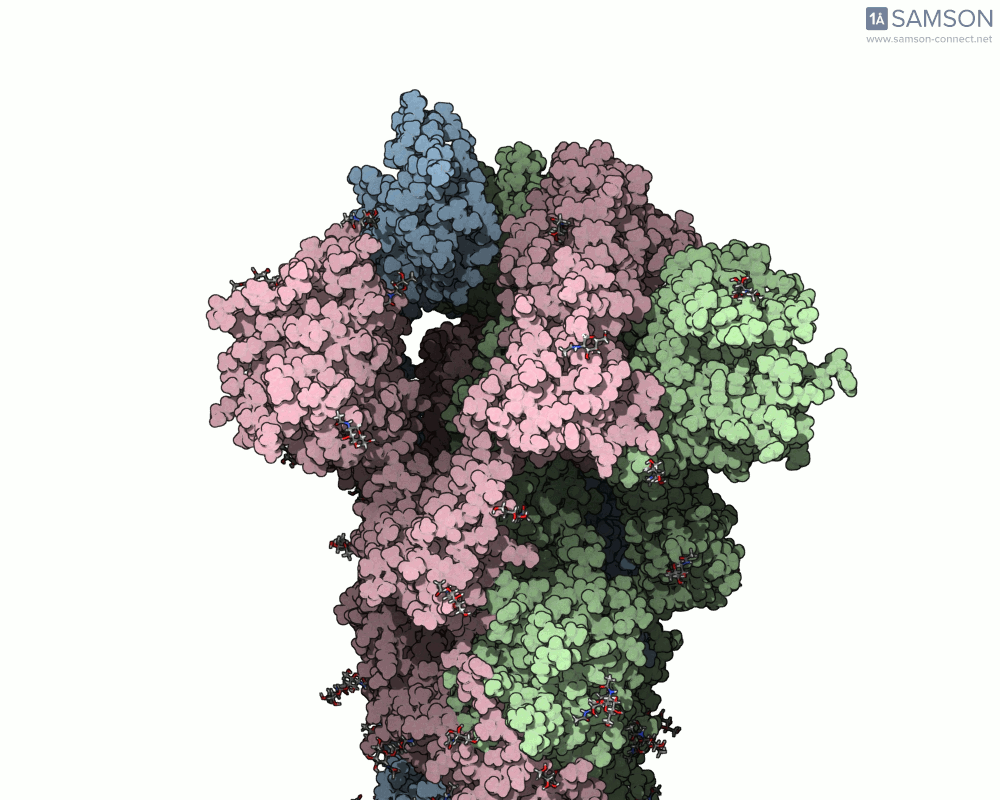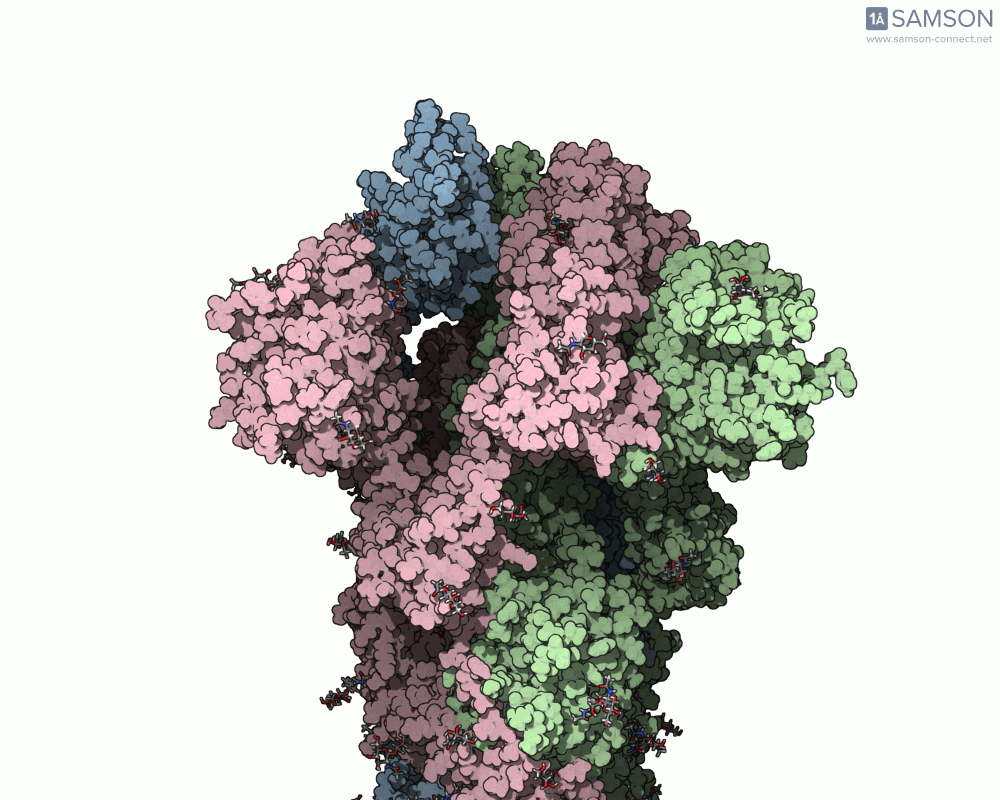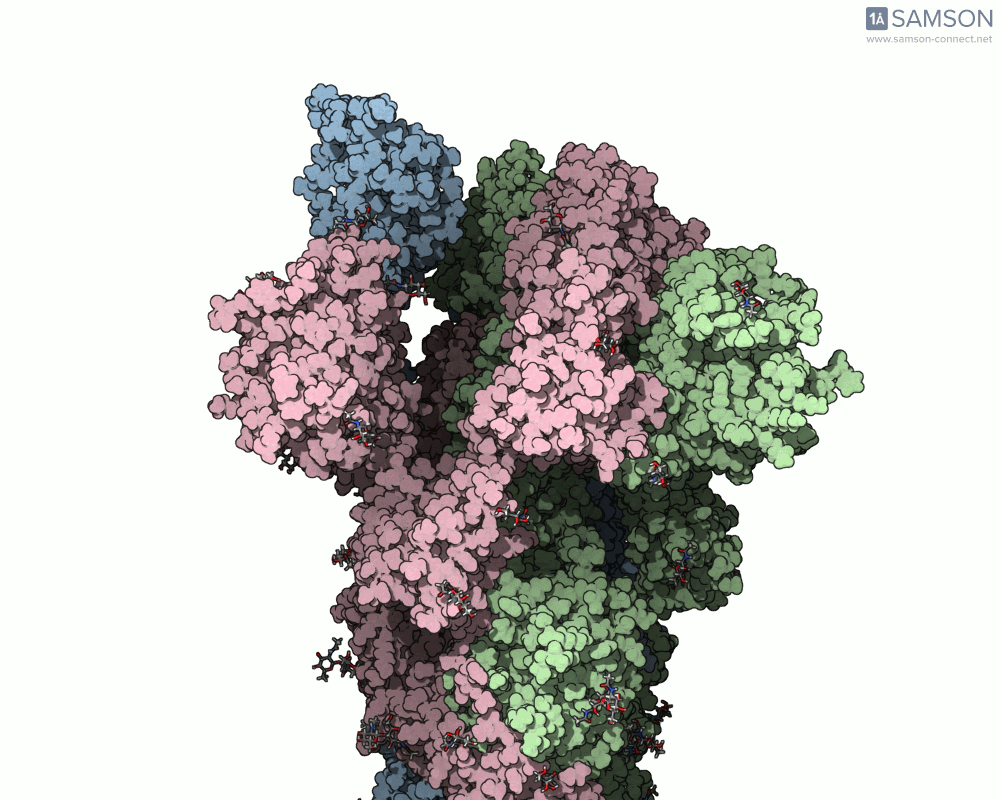
Such visualizations help identify critical conformations for drug targeting, optimize antibody binding, or investigate factors affecting spike stability. Additionally, SAMSON lets researchers export the trajectory in multiple formats (e.g., PDB and SAMSON files) for further analysis or use in simulations.
How to Access the Tools
Both the ARAP and P-NEB modules are free during the pandemic, as part of SAMSON’s initiative to support research against SARS-CoV-2. Here’s how to get started:
- Sign up on SAMSON Connect (it’s free).
- Download and install the SAMSON platform.
- Add the ARAP and P-NEB modules from their respective extension pages.
For more detailed guidance and additional resources, visit the full documentation here.
Please note: SAMSON and all SAMSON Extensions are free for non-commercial use. Get your free version of SAMSON at SAMSON Connect.





