In the intricate world of molecular modeling, selecting specific atomic or molecular structures within your dataset can often feel like finding a needle in a haystack. Whether you are looking to analyze subsets of atoms, residues, or bonds, a streamlined method for selection can save considerable time and effort.
This is where the Node Specification Language (NSL) in SAMSON shines. NSL allows users to filter and interact with nodes (e.g., atoms, residues, structural groups) tailored to your goals. This post will introduce how NSL expressions can be used for filtering nodes effectively, including a tip on leveraging AI for additional assistance.
What Makes Node Filtering Powerful?
Imagine working on a large molecular system where you want to analyze specific structural groups or atoms with distinct properties. For instance, selecting all amino acid residues surrounding a particular ligand, or pinpointing sulfurs within specific chains. Manual methods can be slow and error-prone, especially for complex systems—but NSL simplifies this process dramatically.
Using NSL, you can precisely filter your data in the Document View of SAMSON by applying targeted queries. This enables clarity and targeted analysis, ensuring you focus only on the components relevant to your research.
How Does it Work?
Here’s how you can filter nodes with NSL in document view:
For example, to filter structural groups (e.g., rings or secondary structures):
|
1 |
n.t sg</code> <em>(short for: node.type structuralGroup)</em> |
Filter hydrogens belonging to arginine residues:
|
1 |
H in r.t ARG |
Want to find all nodes within 5 Å of a structural group?
|
1 |
"CA" within 5A of S |
Each of these expressions can be entered directly in SAMSON’s document view. To select the filtered nodes, simply press Enter after entering the NSL query. This seamless filtering can substantially improve productivity as you iterate on your molecular design.
AI Assistance for Node Filtering
One standout feature of NSL in SAMSON is its integration with the AI Assistant. Stuck on how to frame an NSL query for a particular scenario? Just ask for help! By clicking on the Ask AI button (depicted with ![]() ), you can generate a custom NSL expression tailored to your active document hierarchy.
), you can generate a custom NSL expression tailored to your active document hierarchy.
The AI Assistant is particularly useful when dealing with extensive molecular structures or when you’re unsure how to specify certain nodes. Employing this feature ensures you have the exact query you need without trial and error.
Visual Example
Here’s an example where a structural group is filtered in document view using NSL:
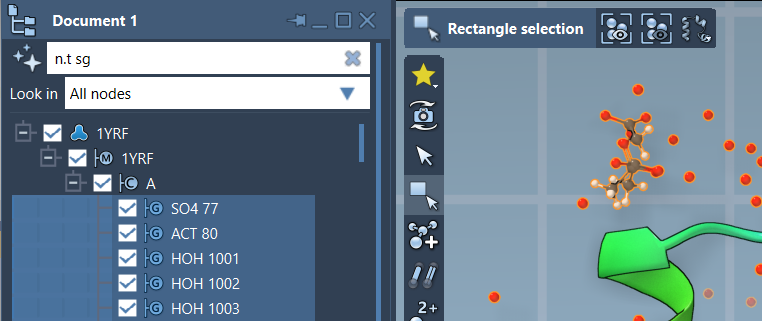
In the above example, the expression n.t sg has been used to select structural groups effectively.
Conclusion
NSL brings precision and efficiency to molecular modeling. With advanced filtering capabilities and integration with AI, SAMSON ensures that you focus on the data that matters without unnecessary hassle. For more detailed information on how you can leverage NSL for node specification, visit the official SAMSON documentation page.
SAMSON and all SAMSON Extensions are free for non-commercial use. You can get SAMSON at samson-connect.net.





