Molecular modeling often involves exploring variations of a molecule to find optimal properties, improve interactions with targets, or test new hypotheses. One effective approach to achieve this is Positional Analogue Scanning (PAS), which lets you systematically modify specific parts of a molecule to create an analogue series. With SAMSON's SMILES Manager, PAS can be performed directly and seamlessly. Here’s how this feature can save you time while providing meaningful insights into molecular design.
What is Positional Analogue Scanning?
Simply put, PAS involves identifying specific patterns or positions on a molecule where substitutions or attachments can be made. These substitutions help generate analogues, allowing researchers to study the effects of different modifications on molecular behavior. For instance, chemists might replace an aromatic hydrogen with a fluorine atom to explore how this impacts binding to a protein target.
Performing PAS in SAMSON
The SMILES Manager extension in SAMSON makes the PAS workflow intuitive. You start with a base molecule and define a SMARTS pattern (a specialized search string) to identify the pattern or location you want to modify.
For example, let’s take the molecule CN1C=C(C(=O)Nc2ccc(-c3ccccc3)c(c2)C(F)(F)F)C(=O)c2ccccc12 as a starting point. Using the SMARTS pattern [cH] (aromatic hydrogen), we can systematically replace this hydrogen with new groups such as nitrogen (N), fluorine (F), or methyl (CH3).
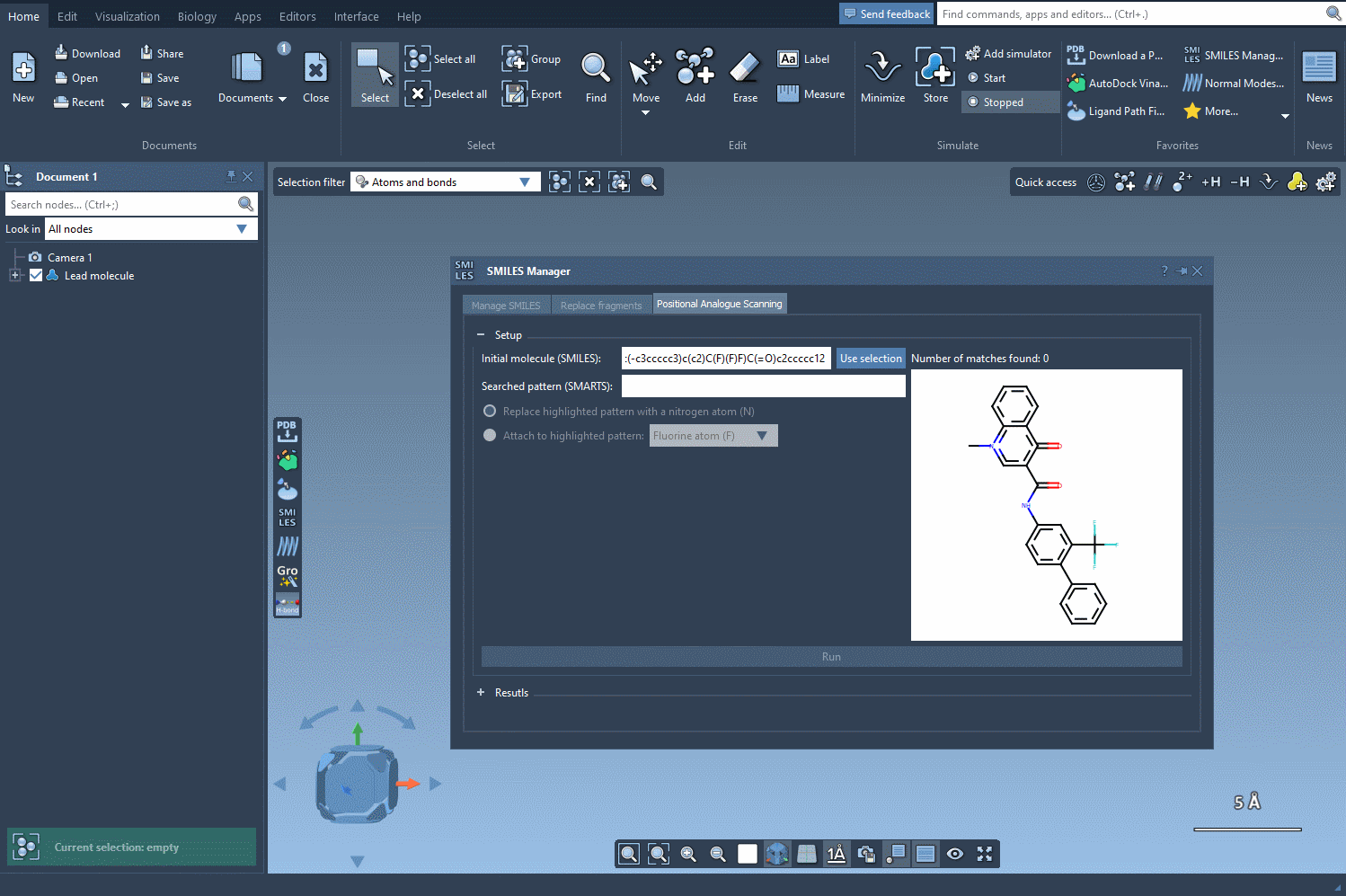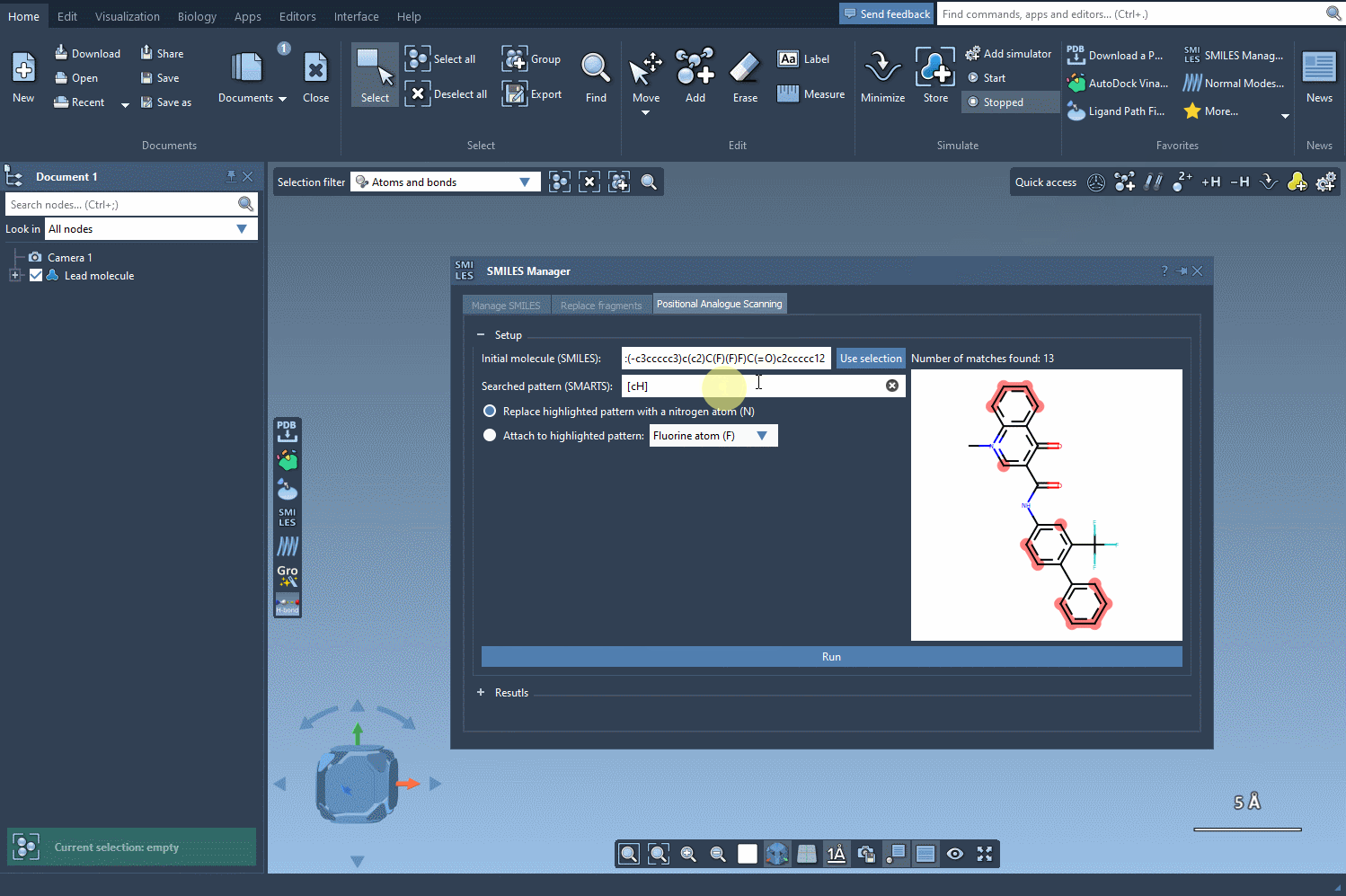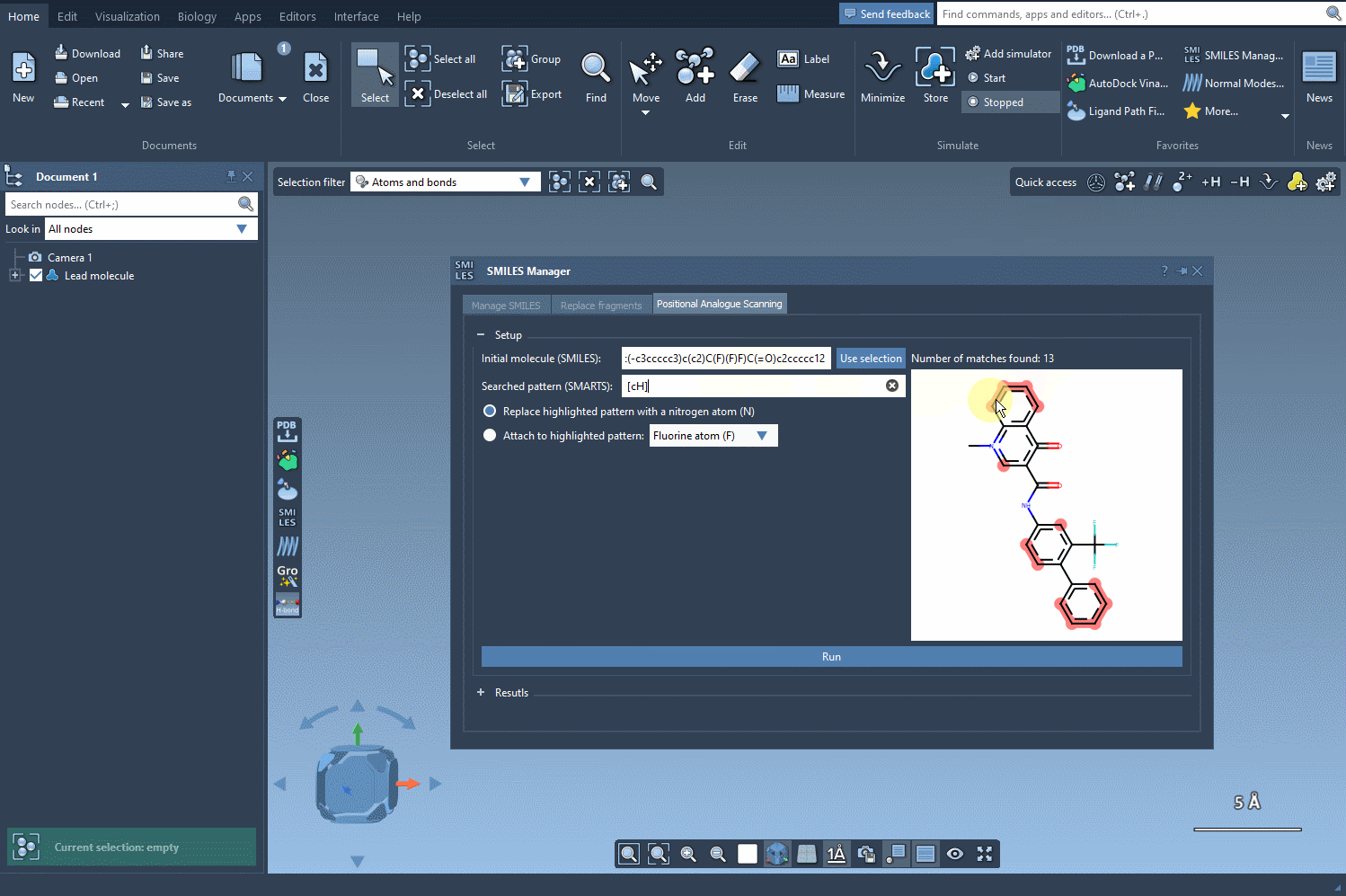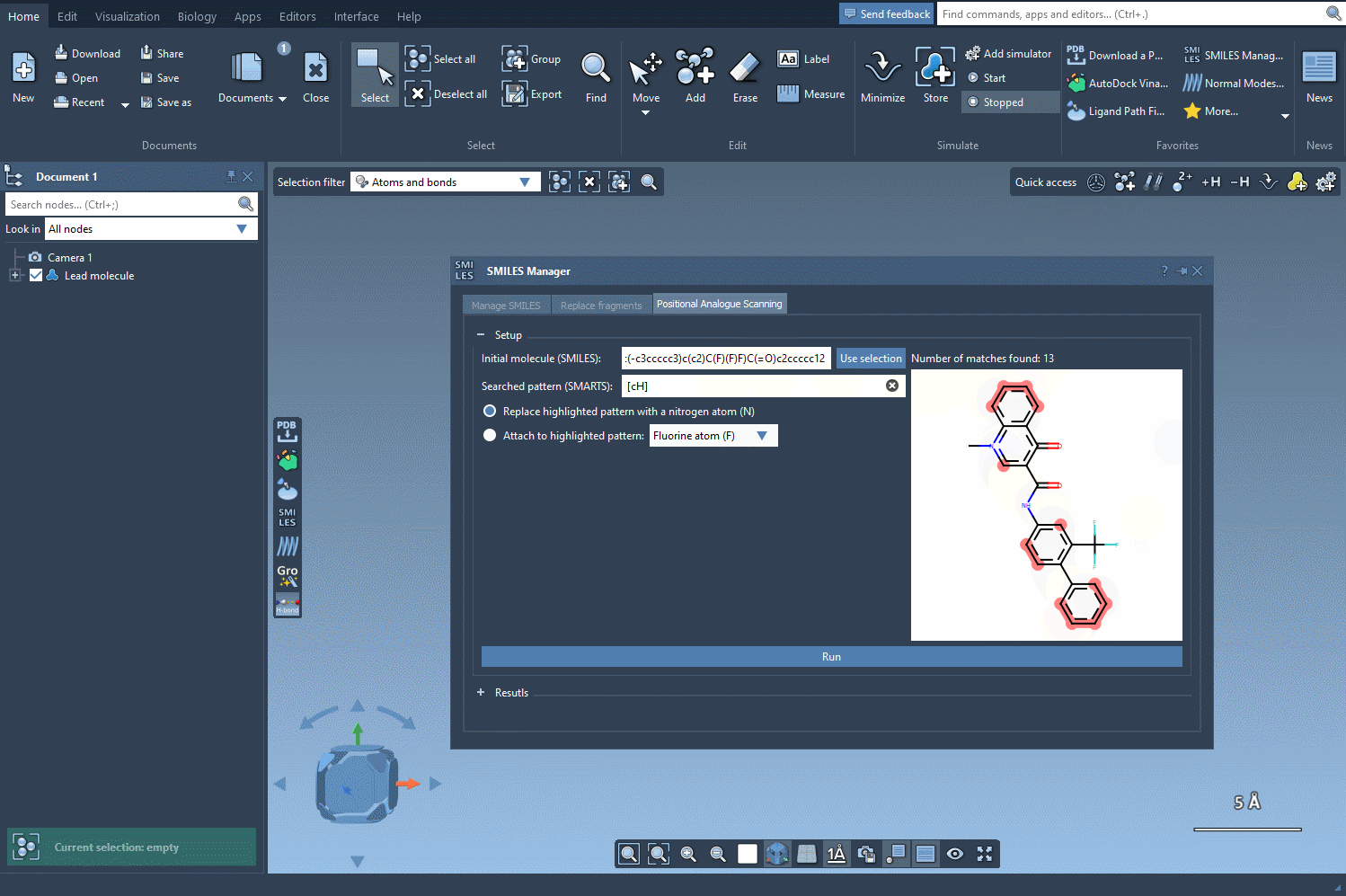
Exploring Analogues with Just a Click
Once the SMARTS pattern is defined, SAMSON highlights the relevant parts of the molecule where substitutions can be made. After selecting the modification type (e.g., replacing a hydrogen with nitrogen), all that’s left to do is hit the Run button. This action generates a series of analogues, including both SMILES representations and their 2D depictions.
As demonstrated below, the process is both rapid and straightforward:
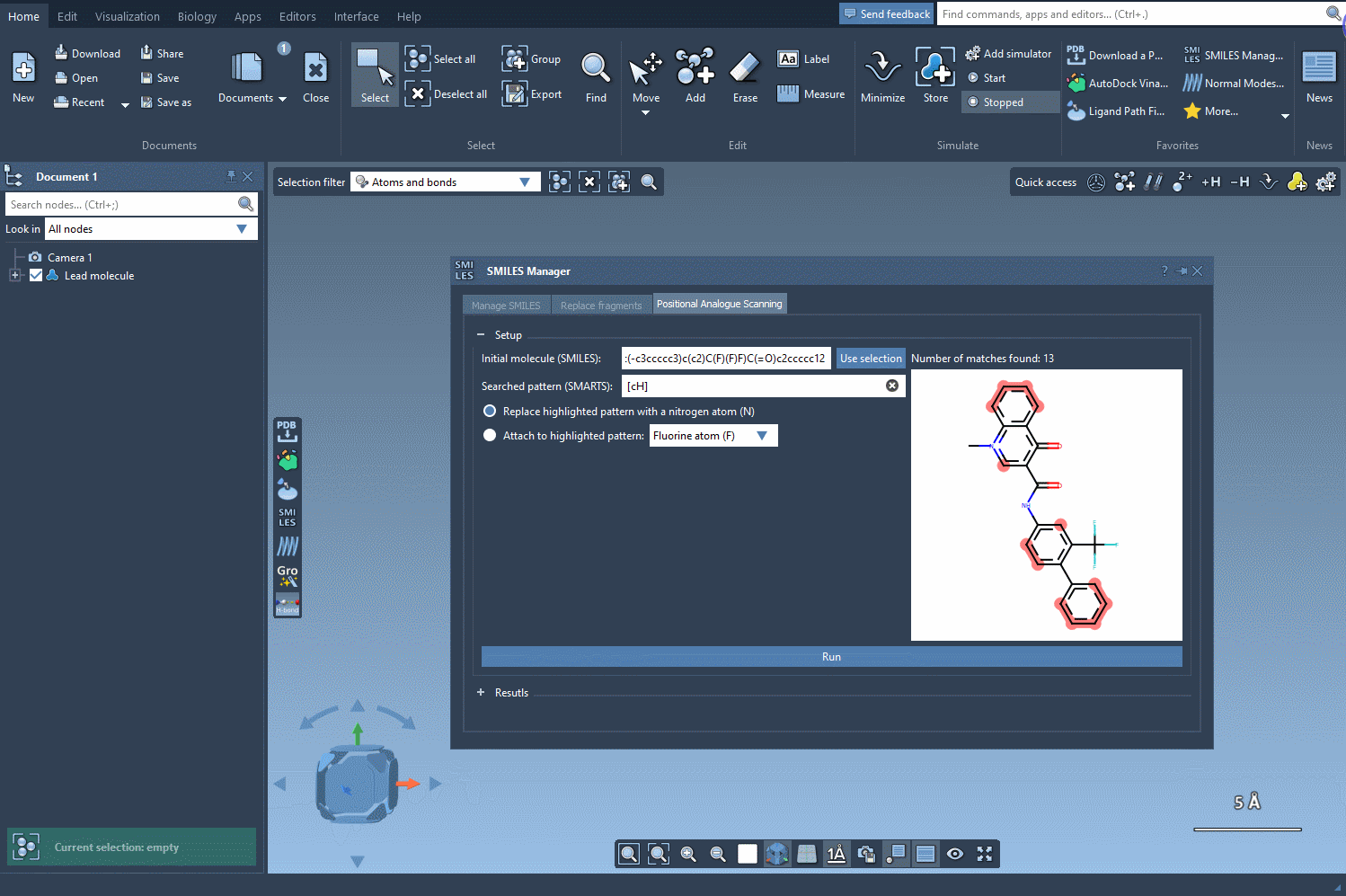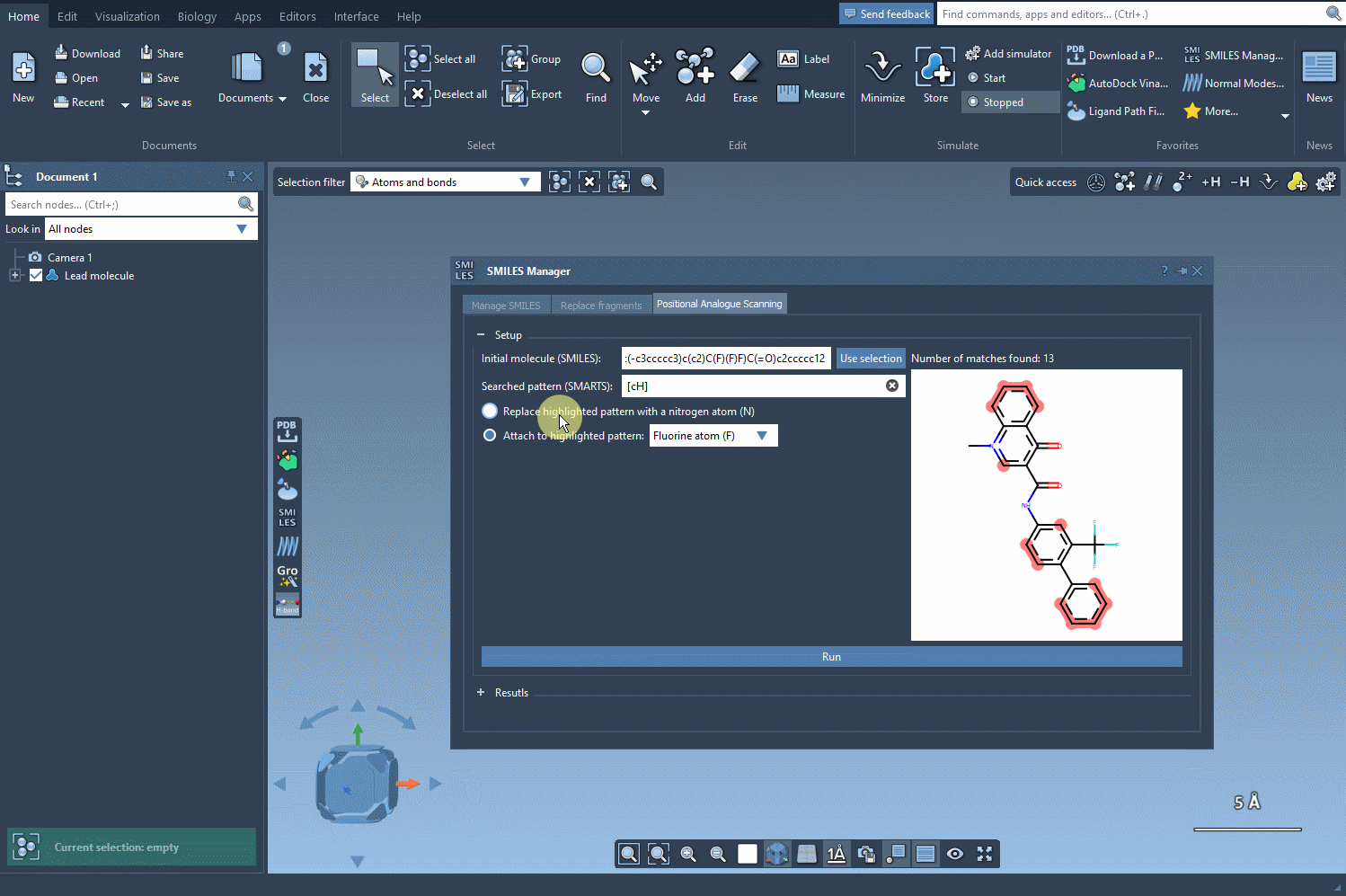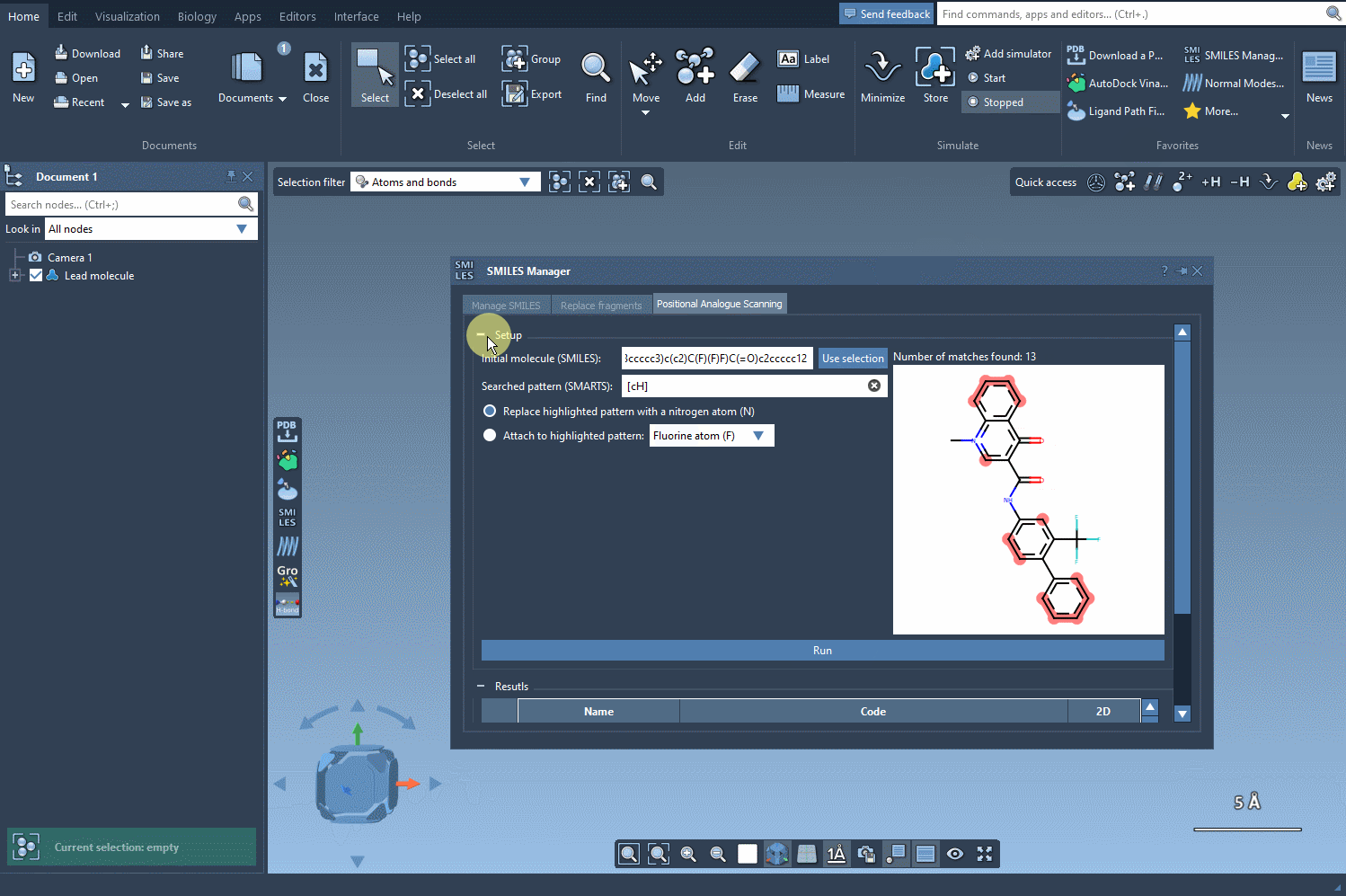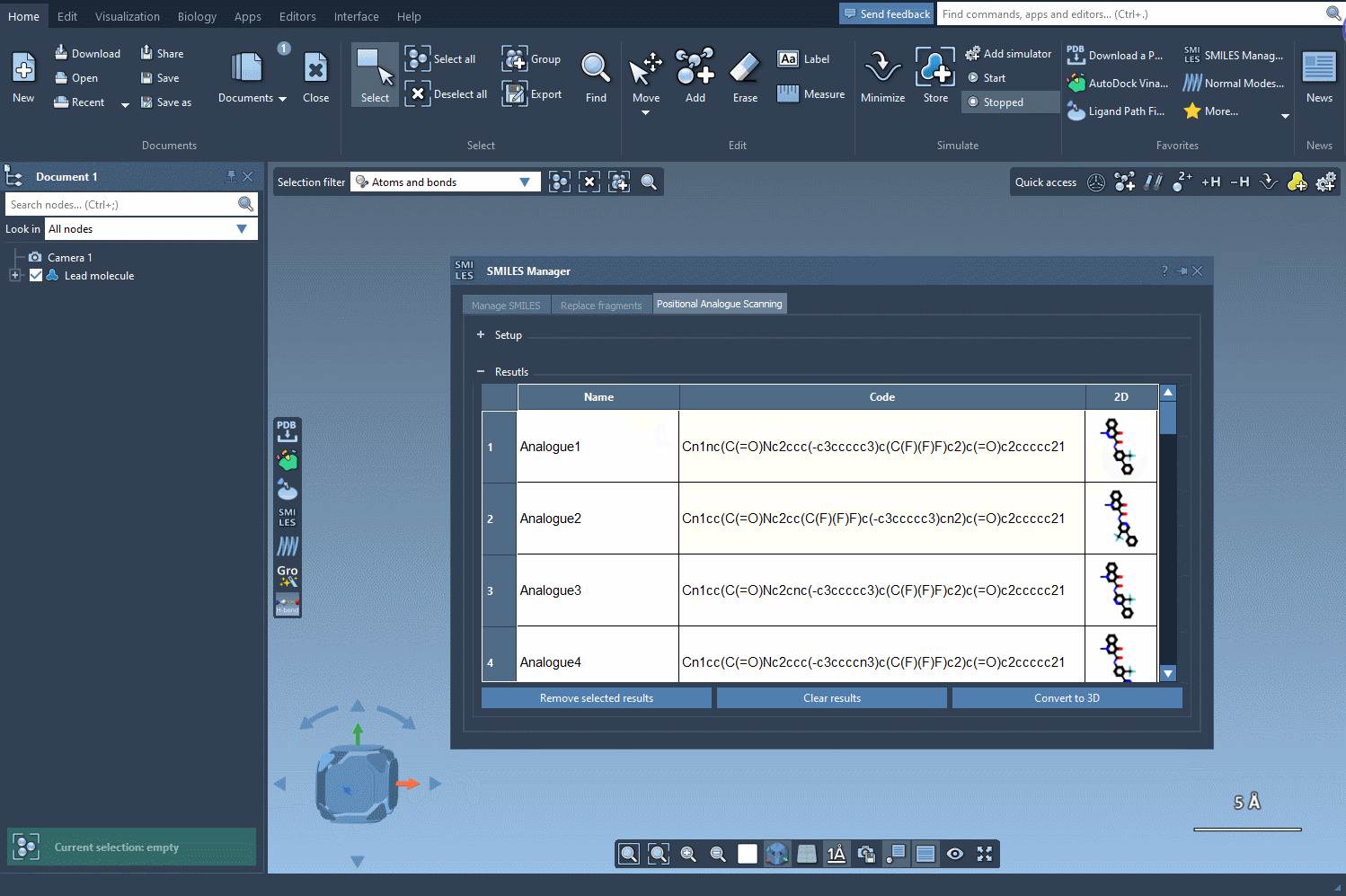
Modify and Refine Your Results
The results table in SMILES Manager offers several tools to refine and explore your analogues:
- Rename or adjust properties: Double-click the corresponding cell to modify SMILES or name values.
- Inspect 2D depictions: Open larger views of 2D images by double-clicking them or using the context menu.
- Generate 3D structures: For any analogue, right-click its 2D depiction and choose to convert it to a 3D structure.
You can even clear out or selectively remove analogues to streamline your study further.
From Analogues to Molecular Insights
After generating your analogues, the next logical step is structural or interaction analysis. For instance, you can use SAMSON’s Autodock Vina Extended extension to perform docking studies of the analogues with a target protein. This helps examine how your structural modifications influence binding or create new interactions, aiding drug design or similar molecular investigations.
Learn More
Want a step-by-step guide on how to perform positional analogue scanning in SAMSON? Check out the original documentation at this link.
Note: SAMSON and all SAMSON Extensions are free for non-commercial use. You can get SAMSON at https://www.samson-connect.net.





