For molecular modelers, understanding biomolecular dynamics can often feel like solving an intricate puzzle: the pieces are there, but the motion connecting them remains elusive. When it comes to studying a critical protein like the SARS-CoV-2 spike, which is central to the virus’s interaction with human cells, unlocking its motion is both a scientific and computational challenge.
Fortunately, SAMSON, the integrative molecular design platform, provides tools to compute, visualize, and explore the opening motion of this spike protein. This blog post delves into why this motion matters, how it was computed, and how SAMSON offers downloadable trajectories to support your molecular explorations.
Why Studying the Spike’s Motion Matters
The SARS-CoV-2 spike protein facilitates the virus’s entry into human cells by binding to the ACE2 receptor. Its opening motion, transitioning from a closed (down) state to an open (up) state, reveals how the spike structurally prepares to engage with host receptors.
The opening motion is more than a curiosity—it’s vital for understanding how antibodies and therapeutic molecules can target this protein, especially since part of the spike remains exposed for receptor recognition. Studying this motion could potentially inspire antiviral drug design or contribute to vaccine research.
How SAMSON Helps: Downloadable Trajectories
To make this motion accessible to researchers, SAMSON has computed a trajectory describing the spike’s full transition from closed to open states. What’s more, the trajectory is available for download in multiple formats:
Once downloaded, these trajectories can serve as an illustrative basis for further calculations, such as docking simulations, molecular dynamics refinements, or comparative studies of structural conformations.
The Spike in Motion: Visualizing the Transition
Visualizing the spike’s motion brings its dynamics to life. Below are animated views of the spike as it morphs between closed and open conformations, a transition critical for understanding its receptor-binding capabilities:
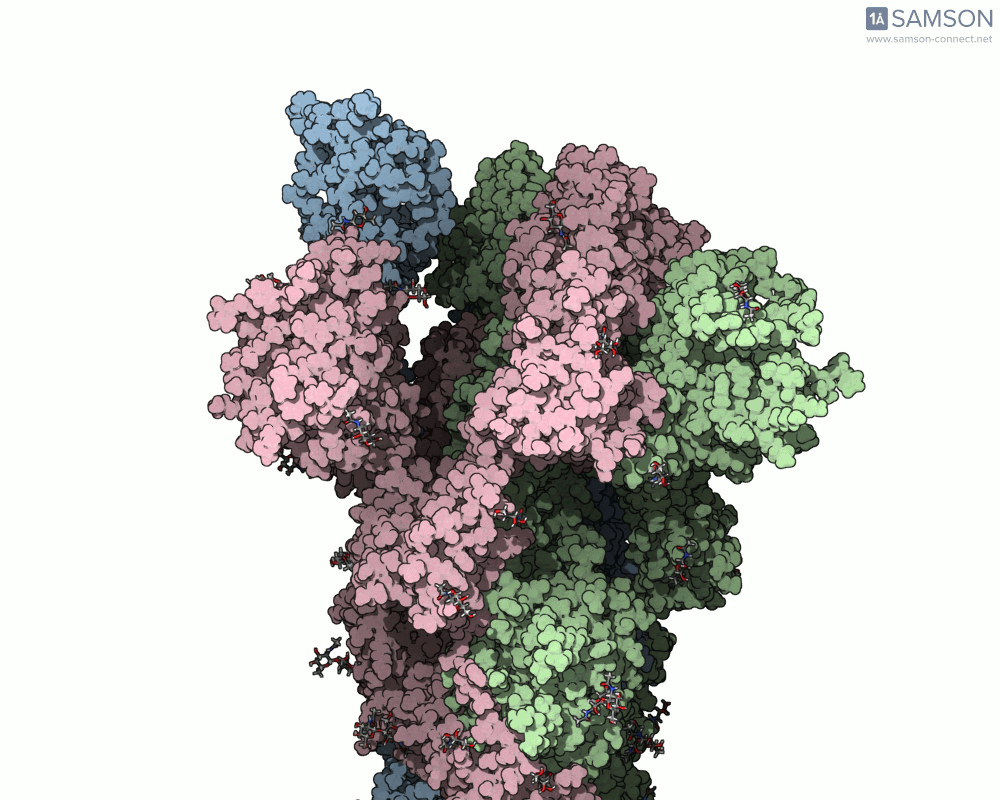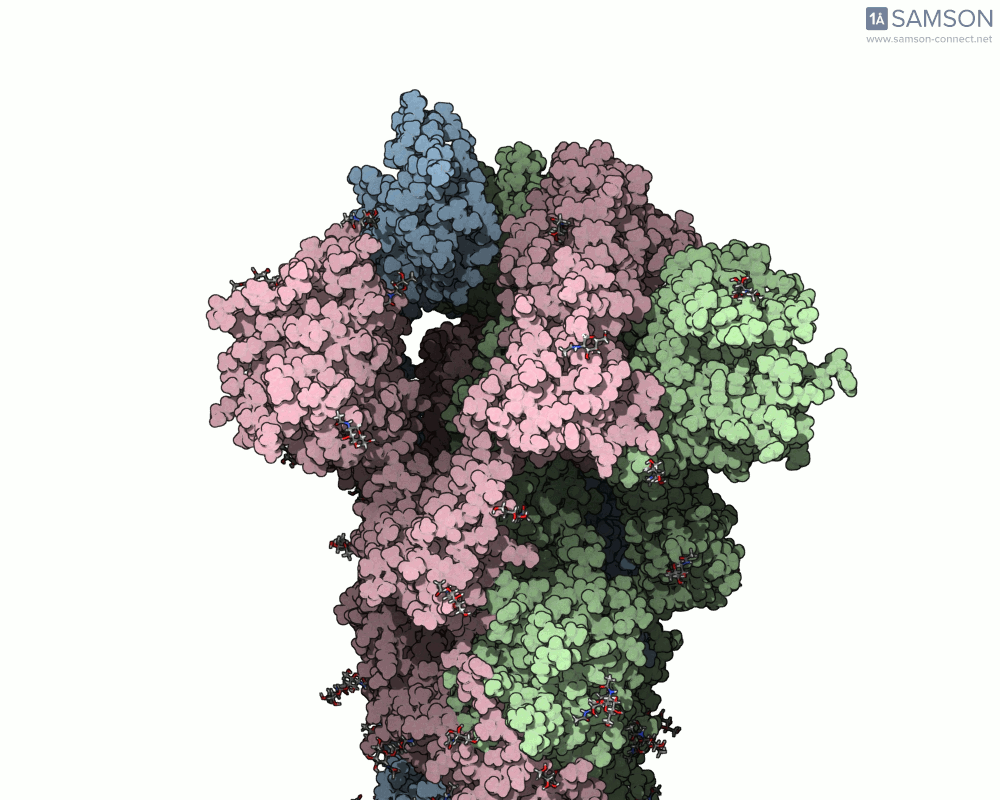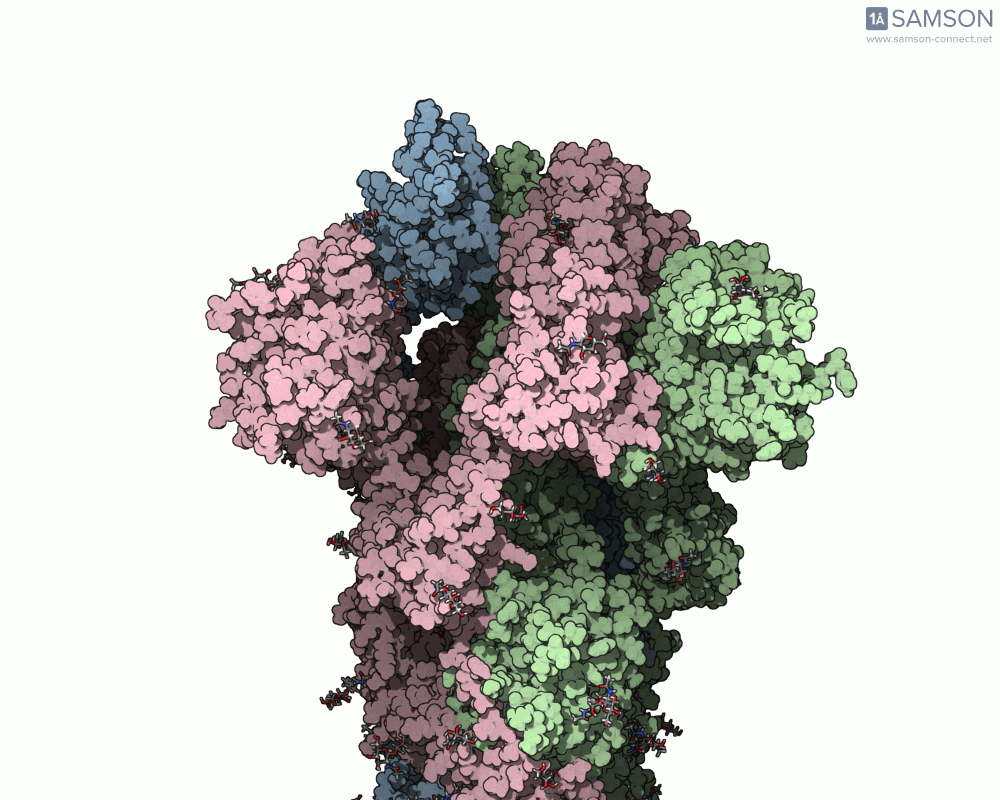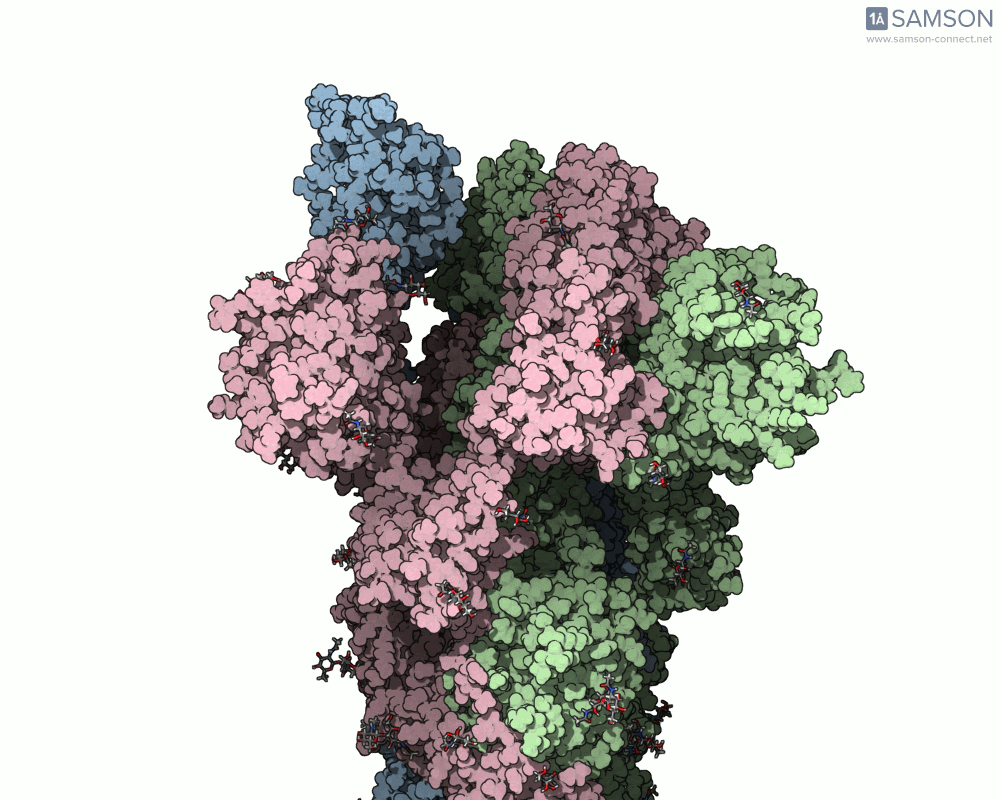
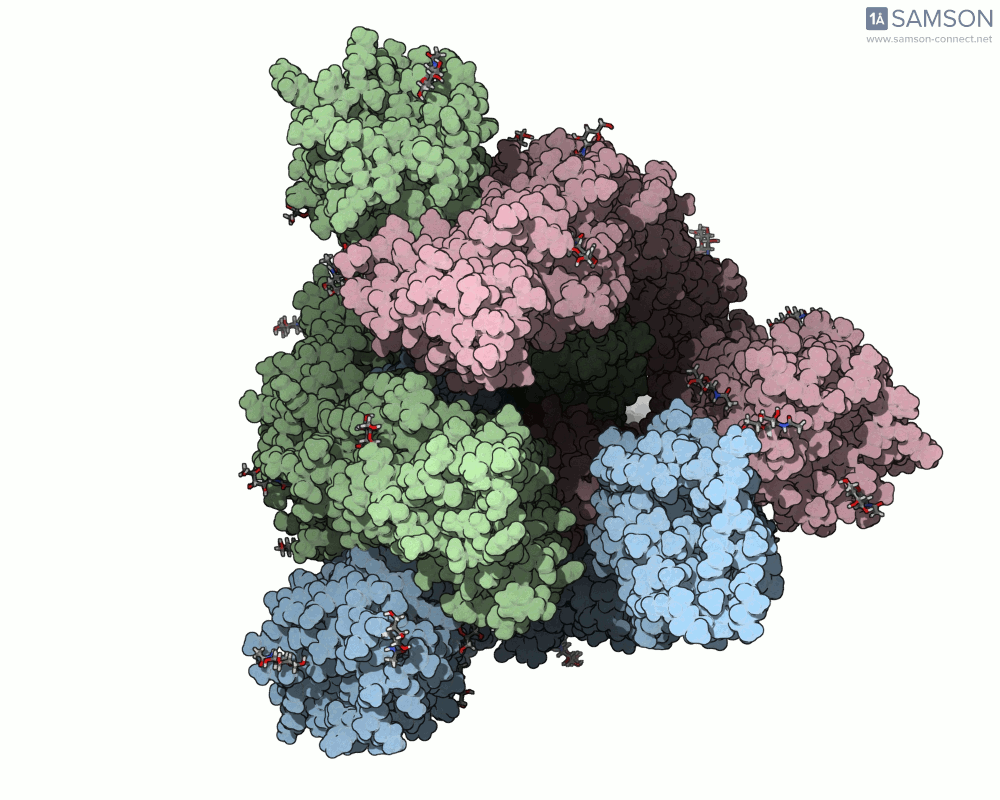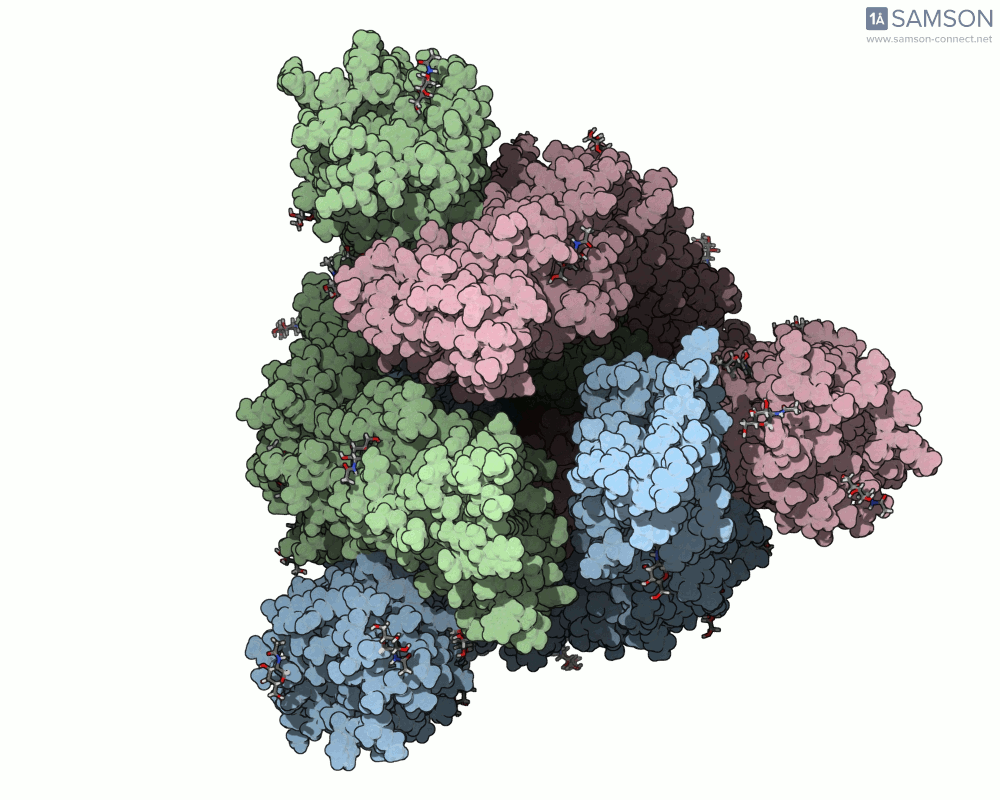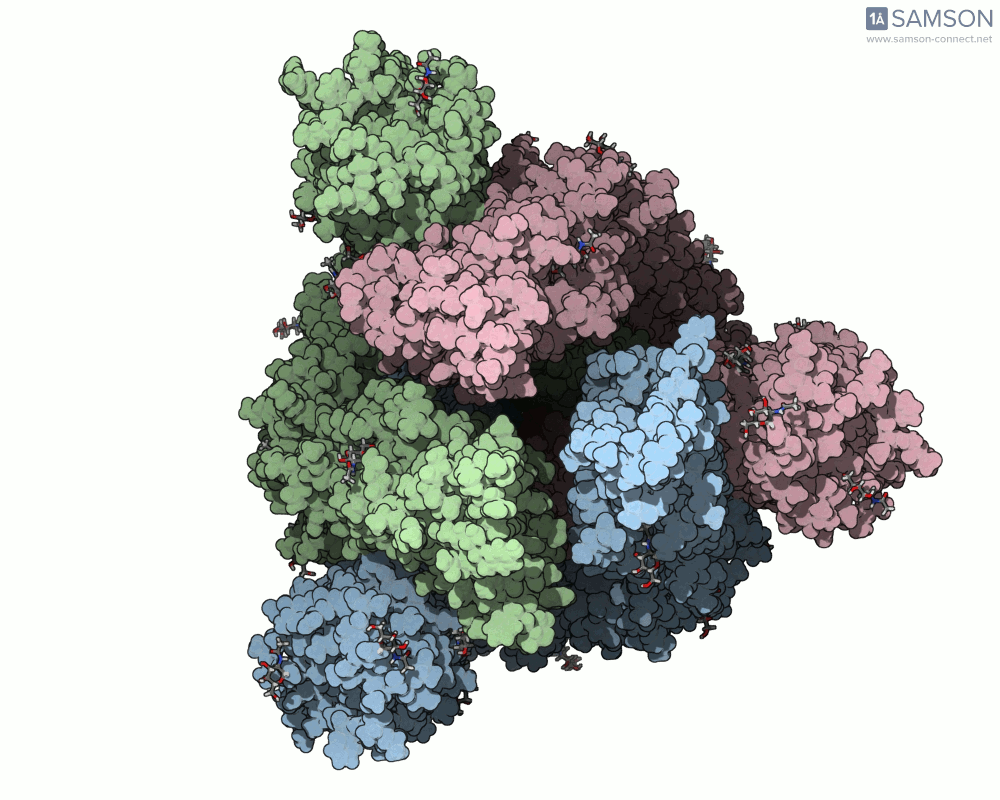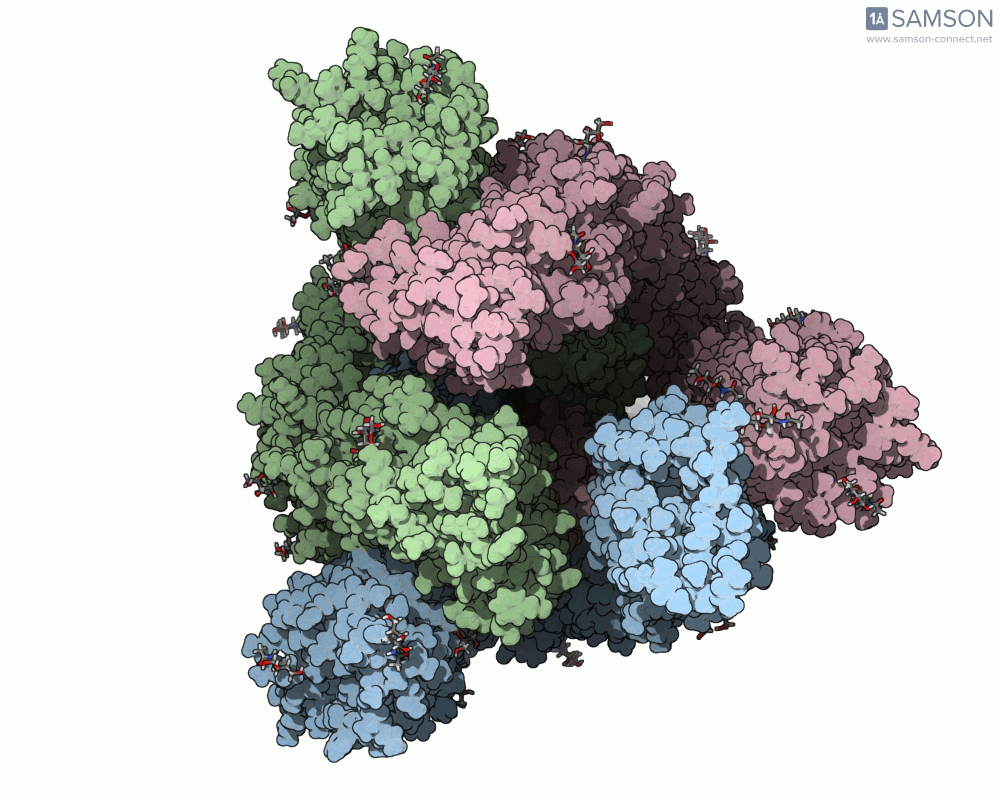
These animations allow researchers to observe structural changes and better hypothesize about functional implications within SARS-CoV-2’s interaction with human cells.
How Was This Computed?
Generating this motion required a careful computational pipeline:
- Set Bond Orders: Adjusted bond orders for sugars in the spike to ensure proper hydrogen addition.
- Added Hydrogens & Minimization: Hydrogens were added to both closed (PDB 6VXX) and open (PDB 6VYB) states, followed by structural minimizations.
- Initial Path Generation: Interpolated a motion path between closed and open states using the ARAP Interpolation Path module.
- Path Refinement with NEB: Improved the generated path using the P-NEB module, enhancing the accuracy of structural changes along the trajectory.
This hybrid approach—combining rigidity-based interpolation with energy-based refinement—enabled a detailed representation of the transition.
The Power of Open Data
All trajectories, images, and animations shared in this project are released under an open CC BY 4.0 license. This ensures researchers worldwide can freely access and use these resources to advance their own studies.
Learn more about the SARS-CoV-2 spike opening motion and computational workflow here.
Note: SAMSON and all SAMSON Extensions are free for non-commercial use. You can download SAMSON at https://www.samson-connect.net.





