Molecular modelers working on the SARS-CoV-2 virus face an intricate yet essential challenge: accurately capturing the dynamics of the spike protein’s conformational changes. The opening motion of the spike, transitioning from a closed to an open state to interact with human ACE2 receptors, holds immense significance in understanding virus-cell binding and designing potential interventions. But how exactly can this dynamic be visualized and studied effectively? Let’s explore an insightful approach and the tools used in SAMSON for modeling this pivotal motion.
The Role of Spike Conformational Dynamics
The SARS-CoV-2 spike protein facilitates viral entry by binding to human ACE2 receptors. This binding relies on the spike’s receptor-binding domain (RBD), which flips from a closed state (downward) to an open state (upward) to expose itself for interaction. Capturing this transition is crucial for multiple applications: designing antiviral strategies, visualizing antibody binding, or even simulating mutation effects.
Visualizing the Spike’s Opening
SAMSON offers a unique perspective on the spike’s opening motion through curated visualizations, computational trajectories, and animations. For instance, the following animations demonstrate this transition from different angles:
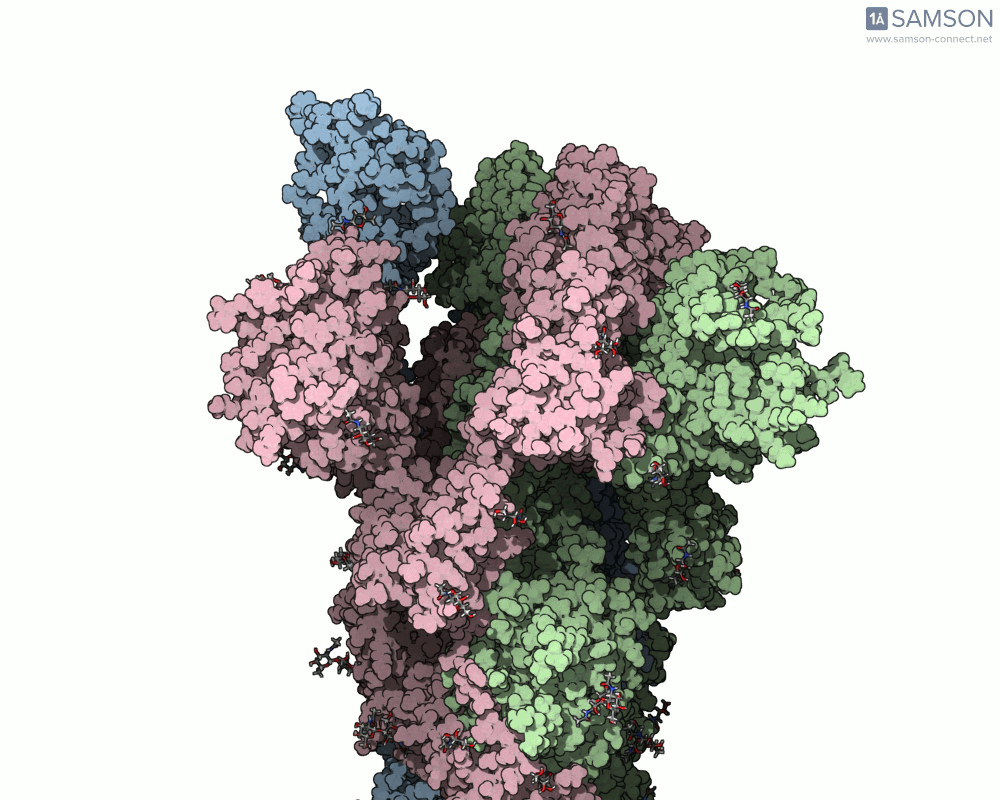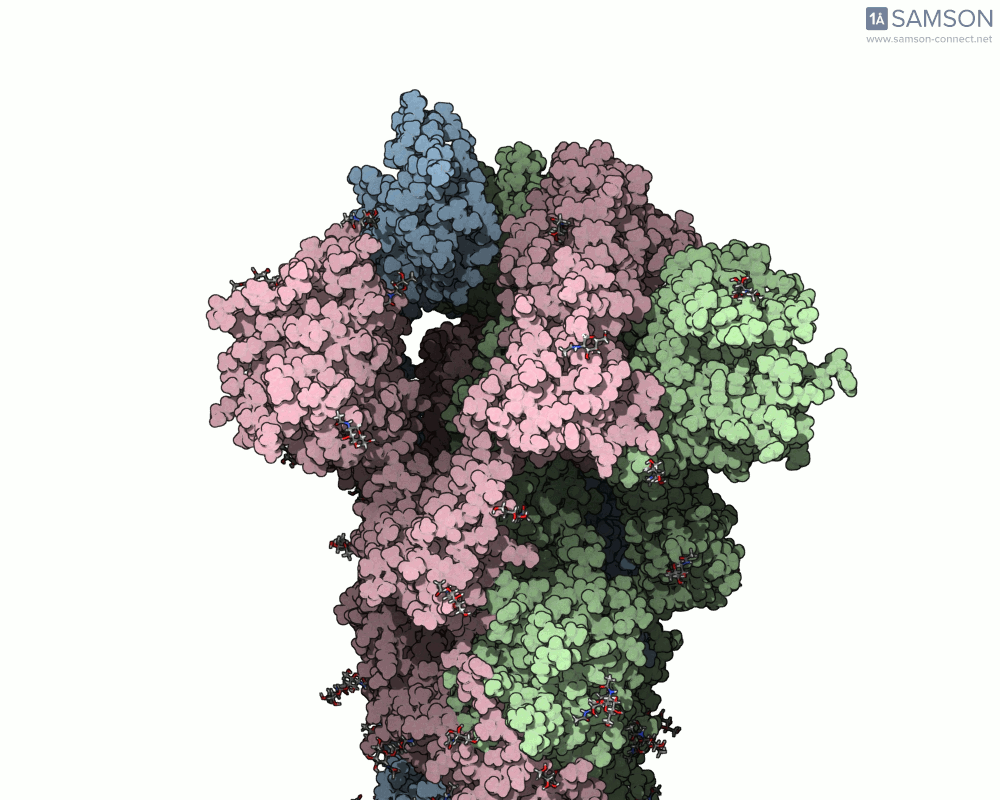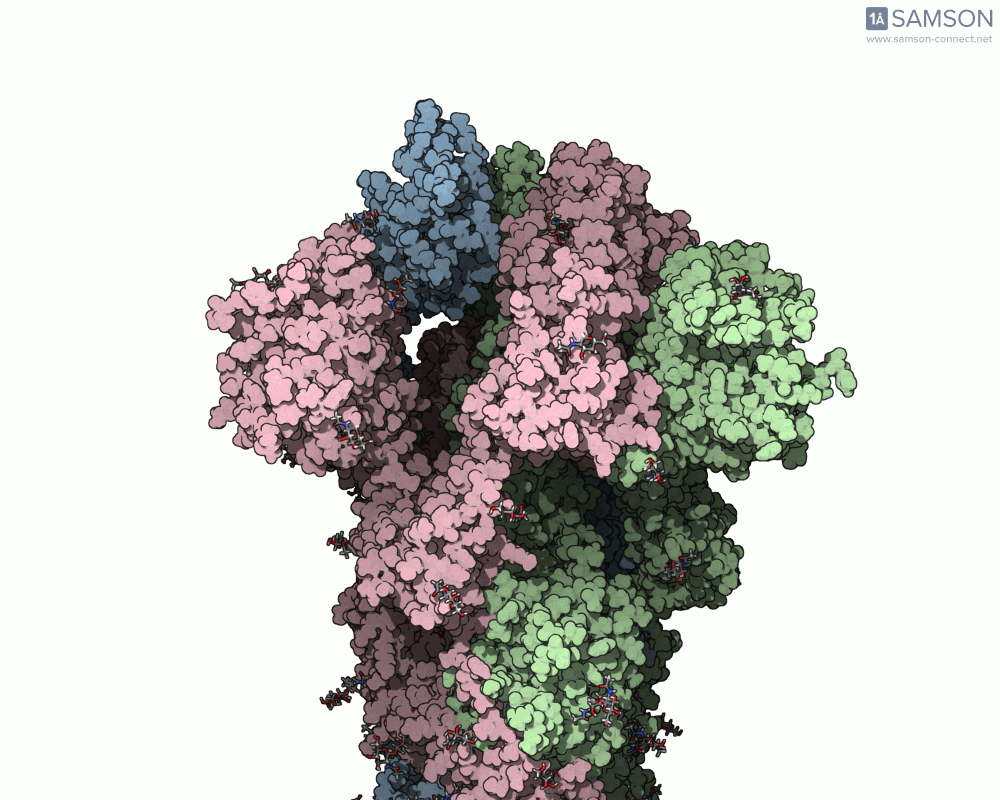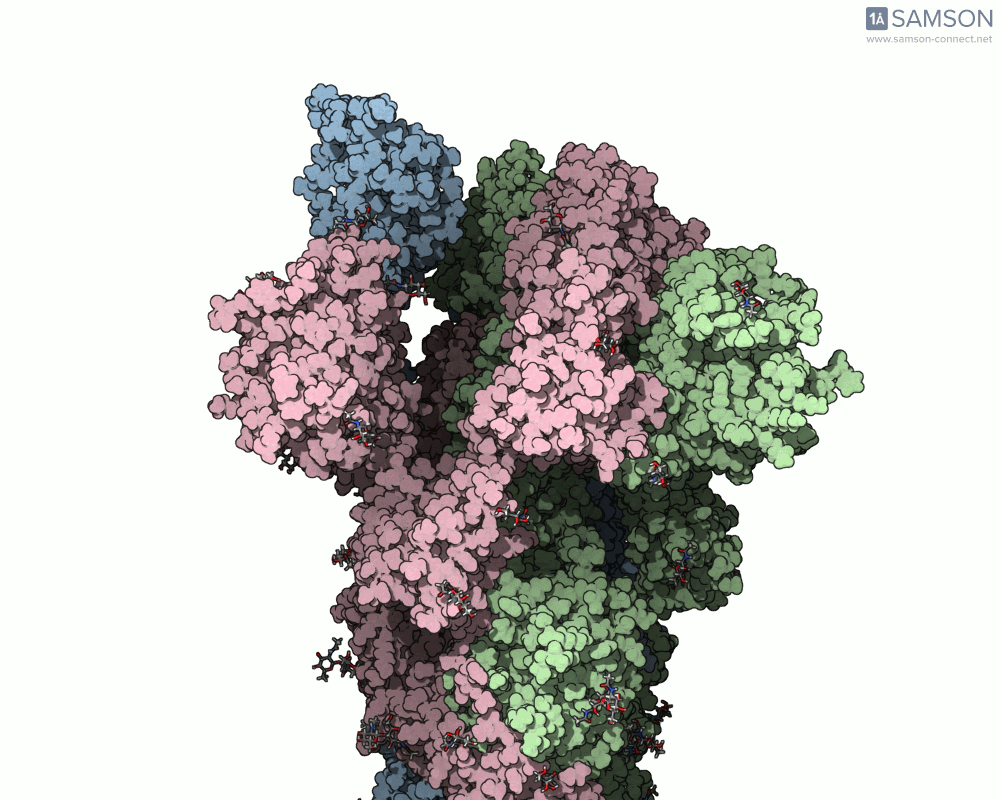
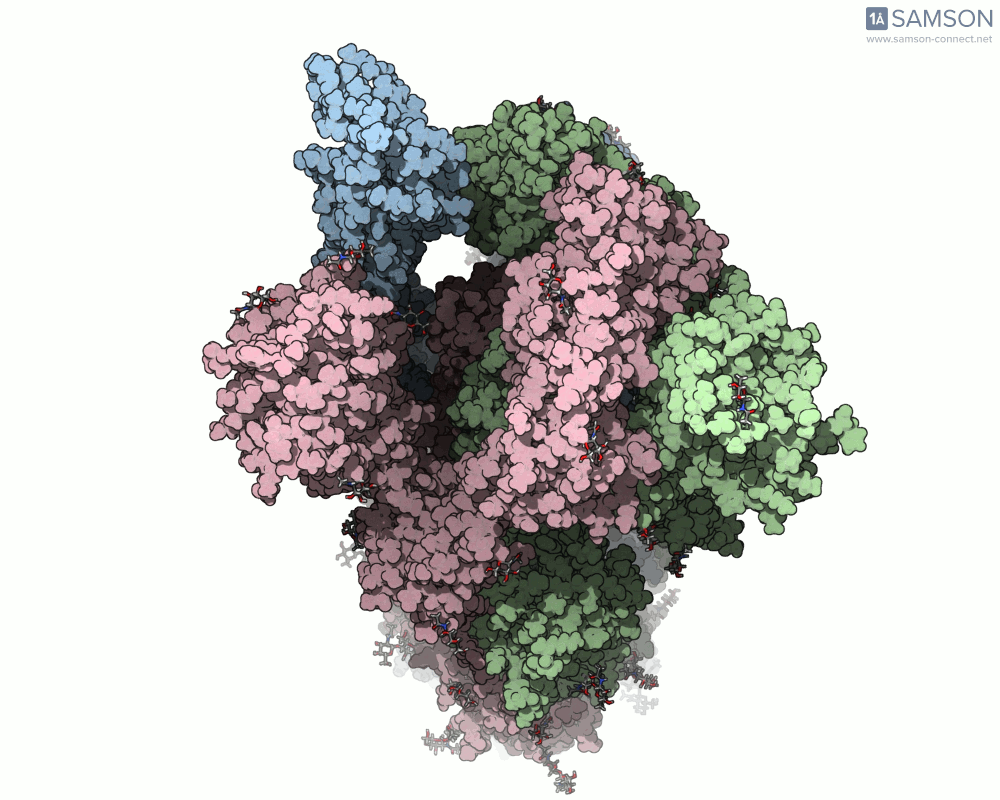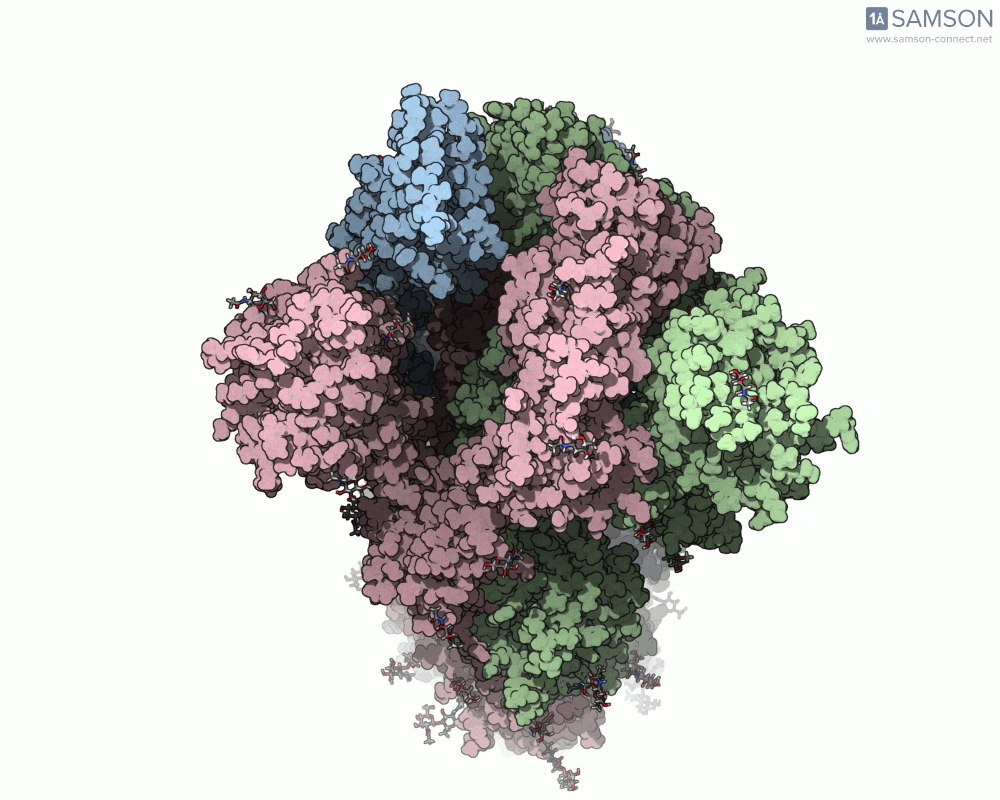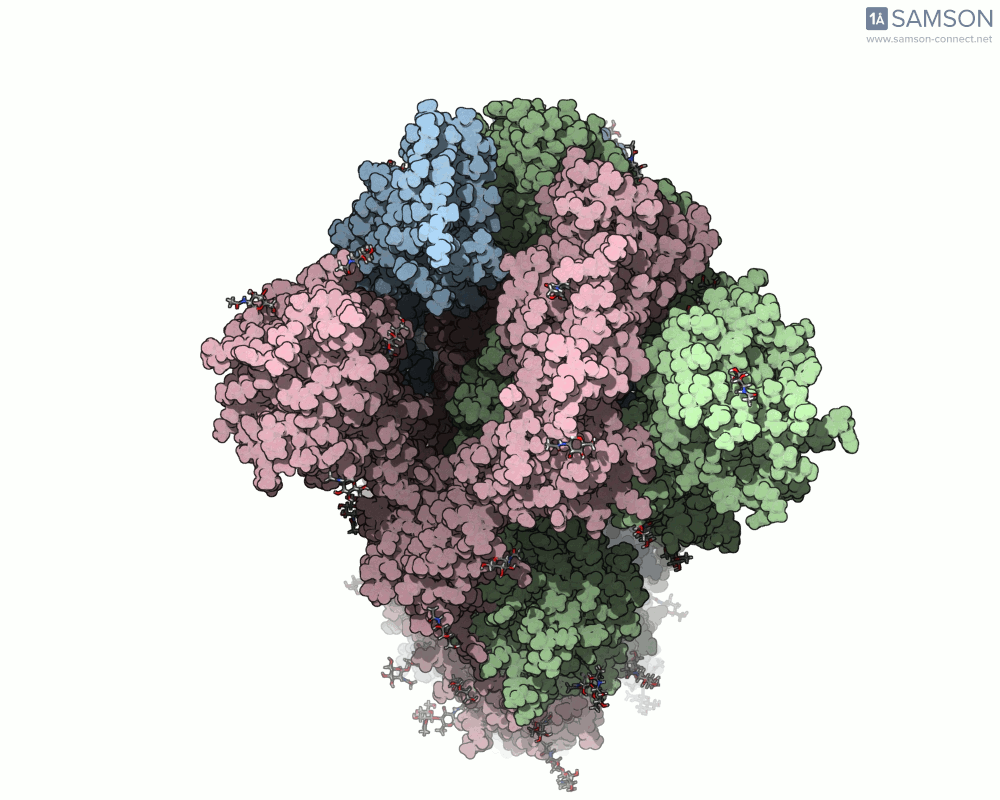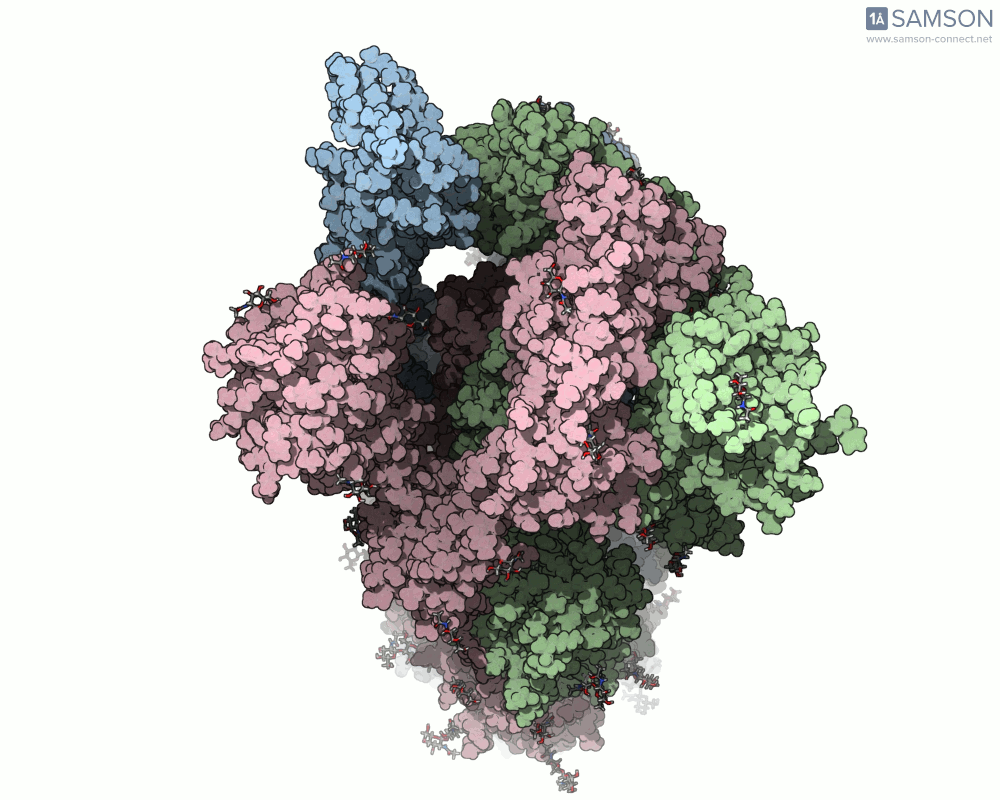
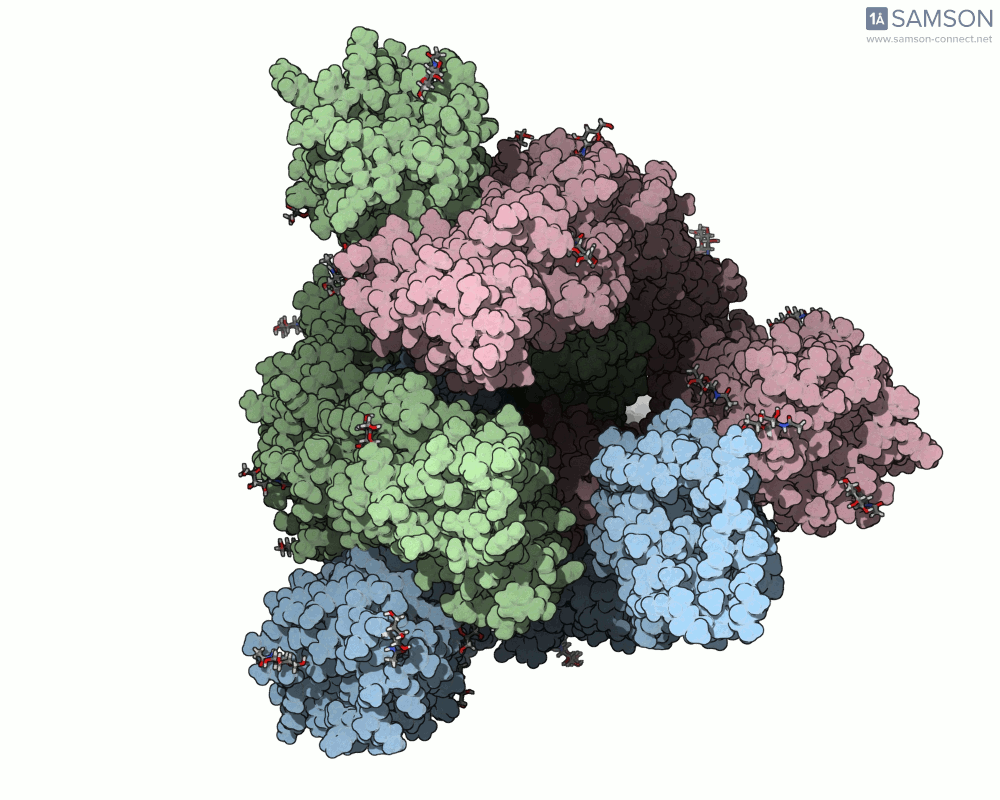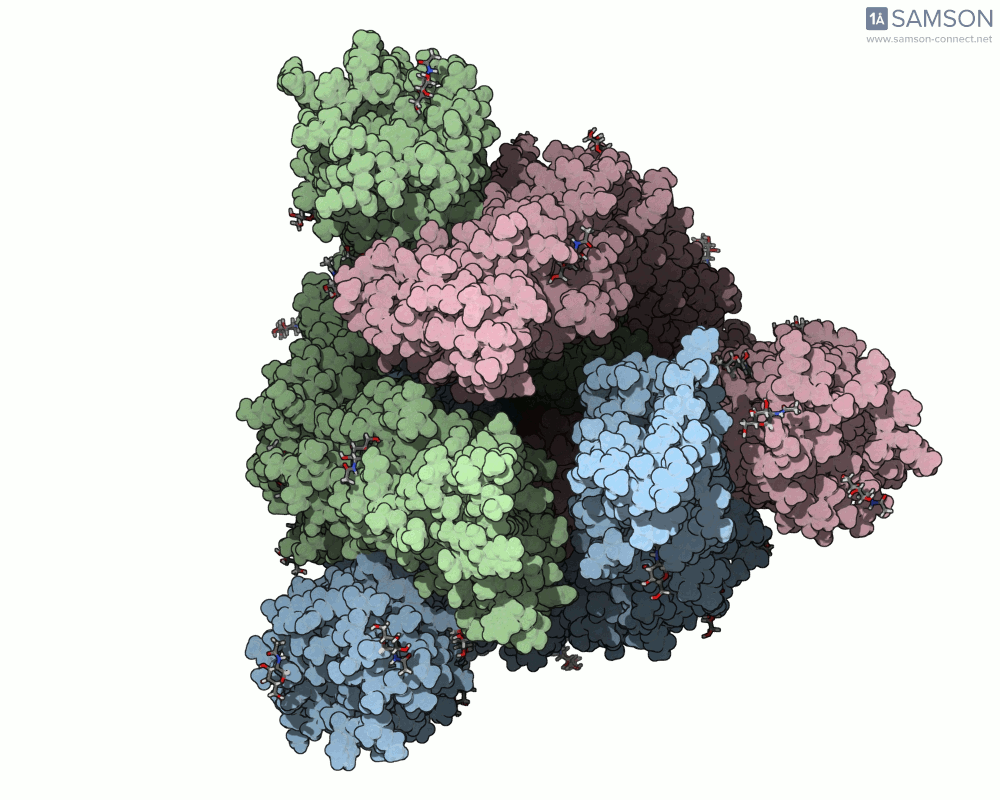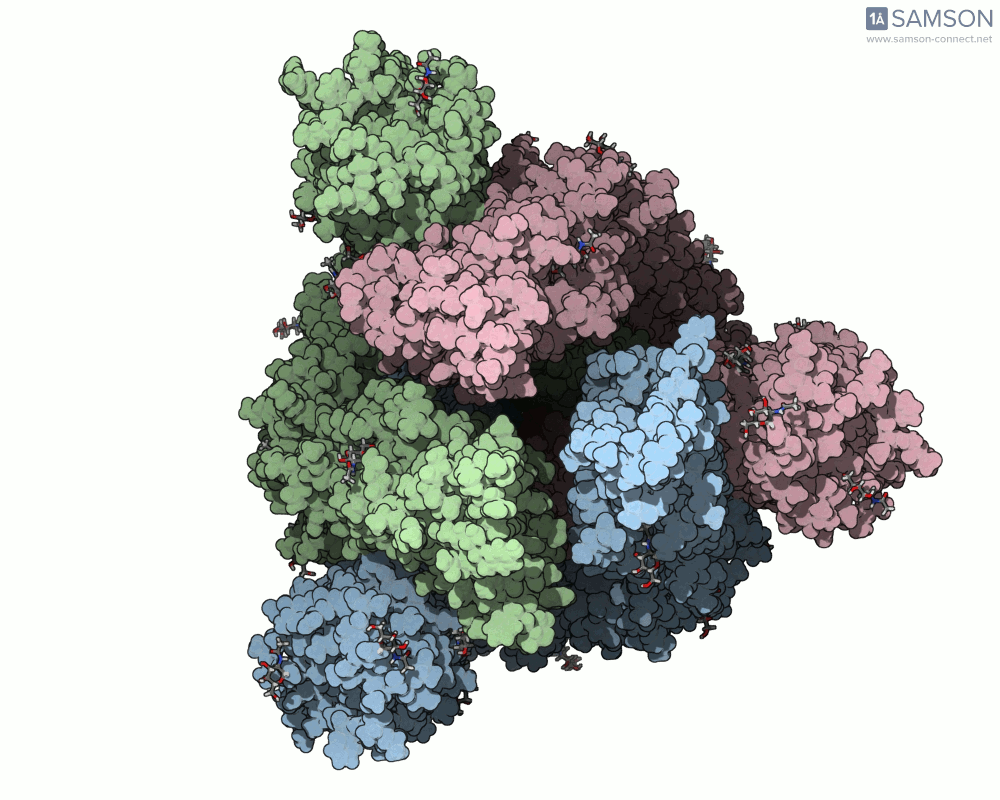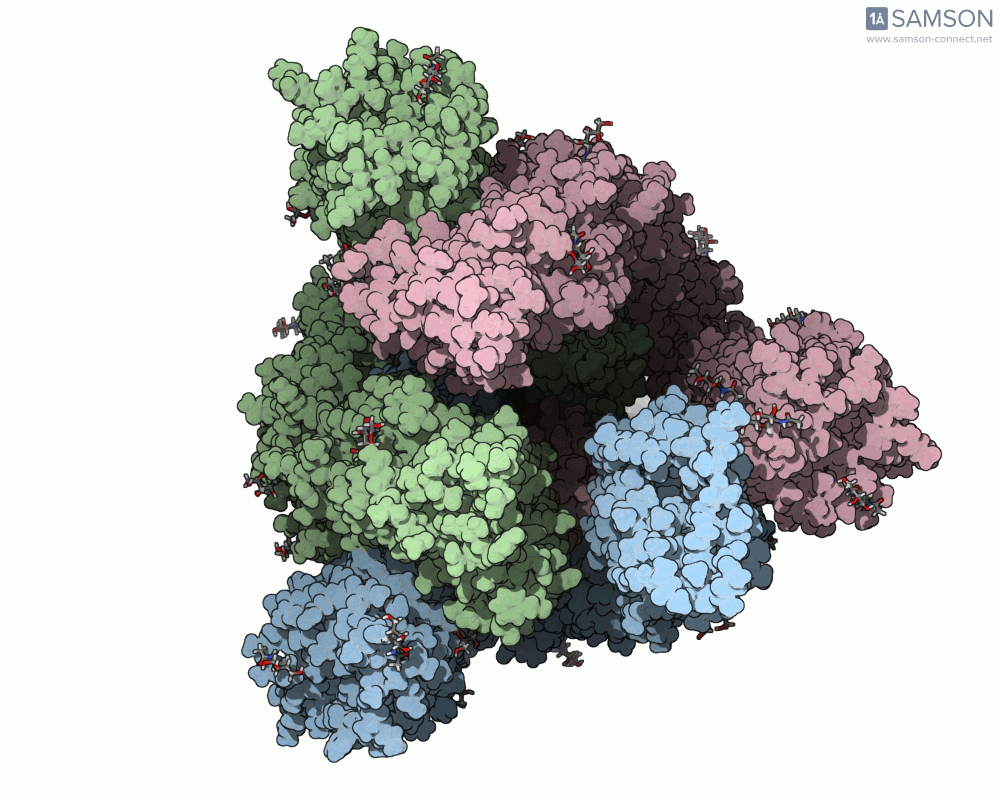
These engaging visuals not only make it easier to comprehend the spike’s behavior but also serve as starting points for further computational analyses, such as docking studies or pathway optimizations.
Ready-to-Use Data for Molecular Modeling
For modelers who want a hands-on approach, SAMSON provides downloadable trajectory files:
These files capture the defined transition pathway from the closed to the open state and are compatible with further analyses in SAMSON or other molecular modeling tools. However, it’s worth noting that these calculated trajectories are provided “as is” and may require additional validation for experimental applications.
SAMSON’s Pipeline for Spike Motion Modeling
The computed trajectories were generated within SAMSON using a combination of accessible modules, specifically the ARAP Interpolation Path module and P-NEB (Parallel Nudged Elastic Band) module. These steps included:
- Defining starting and ending conformations using known PDB structures (6VXX for the closed state, 6VYB for the open state).
- Employing ARAP to interpolate paths between these states, ensuring structural intermediates remain physically plausible.
- Refining generated paths with the P-NEB module for higher accuracy.
This computational pipeline exemplifies how molecular animations can be rigorously constructed and utilized for more detailed investigations or hypothesis testing.
Conclusion
Modeling the opening motion of the SARS-CoV-2 spike is an essential task for understanding viral behavior and designing therapeutic solutions. Thanks to SAMSON’s trajectory visualizations, downloadable data, and integrated modules, molecular modelers can efficiently approach and analyze this key dynamic.
To explore the SARS-CoV-2 spike’s dynamics in greater detail, visit the original documentation page at the SAMSON tutorial on the spike opening motion.
Note: SAMSON and all SAMSON Extensions are free for non-commercial use. Download SAMSON from SAMSON Connect.





