For molecular modelers, especially those engaged in interactive molecular editing, adapting molecular structures while maintaining physical realism can be a challenging task. This is where the Interactive Modeling Universal Force Field (IM-UFF) in the SAMSON platform comes into play. This extension enhances the popular Universal Force Field (UFF) model by enabling topological updates in real-time during molecular editing. Below, we’ll explore how IM-UFF can make molecular modeling more flexible and interactive, addressing a key pain point for researchers and developers.
Why IM-UFF Is a Game-Changer
IM-UFF is designed to handle topological changes with ease. It allows users to create and break covalent bonds, modify bond orders, and change atom typizations while maintaining realistic simulations. This ability lets you explore different structural configurations without losing physical accuracy. With this tool, you can interactively edit molecular systems, guided by physically-based inter-atomic forces.
One of IM-UFF’s standout features is its capacity for automatic topology updates. When atoms are displaced significantly during a simulation, bonds break, and new topologies can form. Similarly, bringing atoms closer together results in bond formation. This eliminates the need to manually redefine system topology after edits, streamlining workflows.
How to Get Started with IM-UFF
Here’s a quick guide to setting up and using IM-UFF:
- Add the IM-UFF Extension to your SAMSON interface.
- To start a simulation, open a document with your desired molecular system and add a simulator via Edit > Simulate > Add simulator (shortcut:
Ctrl + Shift + Mon Windows /Cmd + Shift + Mon macOS). - Select Interactive Modeling Universal Force Field from the list of interaction models.
- Choose a state updater to manage the simulation dynamics, such as the Fast Inertial Relaxation Engine (FIRE).
- Click the OK button to launch.
Once initialized, you’ll have access to the IM-UFF parameter window, where you can configure simulation settings. Two important options are:
- Static topology (UFF only): Enables switching between standard UFF and IM-UFF modes.
- Keep vdW for manipulated: Controls whether van der Waals forces are considered during manual manipulation of atoms. Disabling it makes atom manipulations simpler by removing obstructive repulsive forces.
Key Highlights in Simulation Mode
When the IM-UFF simulation is running, you can observe the molecular system’s behavior as atoms are moved. Bonds will adapt or break depending on the atomic displacements. This enables intuitive editing of molecular structures while the system maintains physical consistency. For example, slight movements preserve bond topology, while larger displacements update it dynamically.
Below is an example of an IM-UFF simulation:
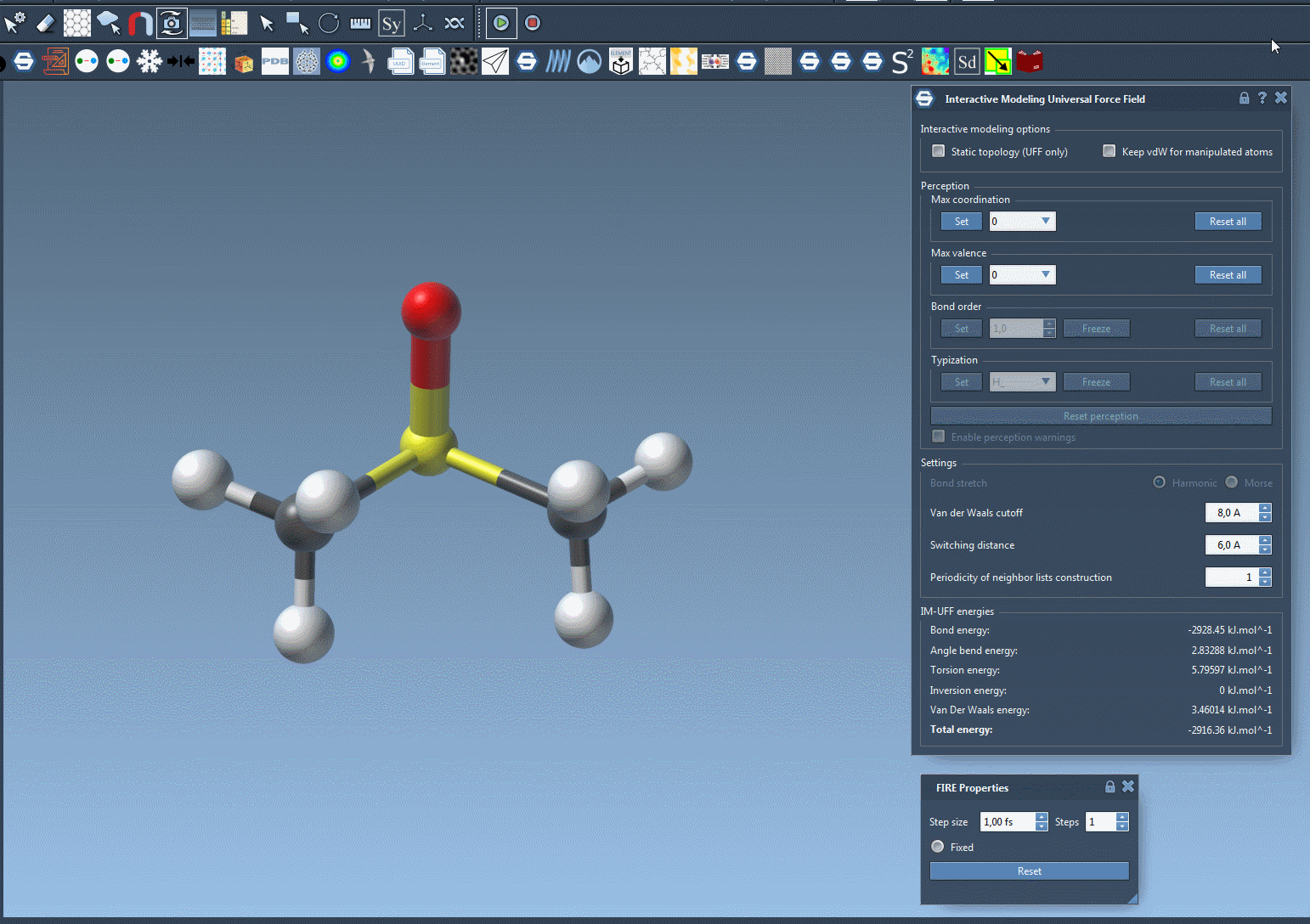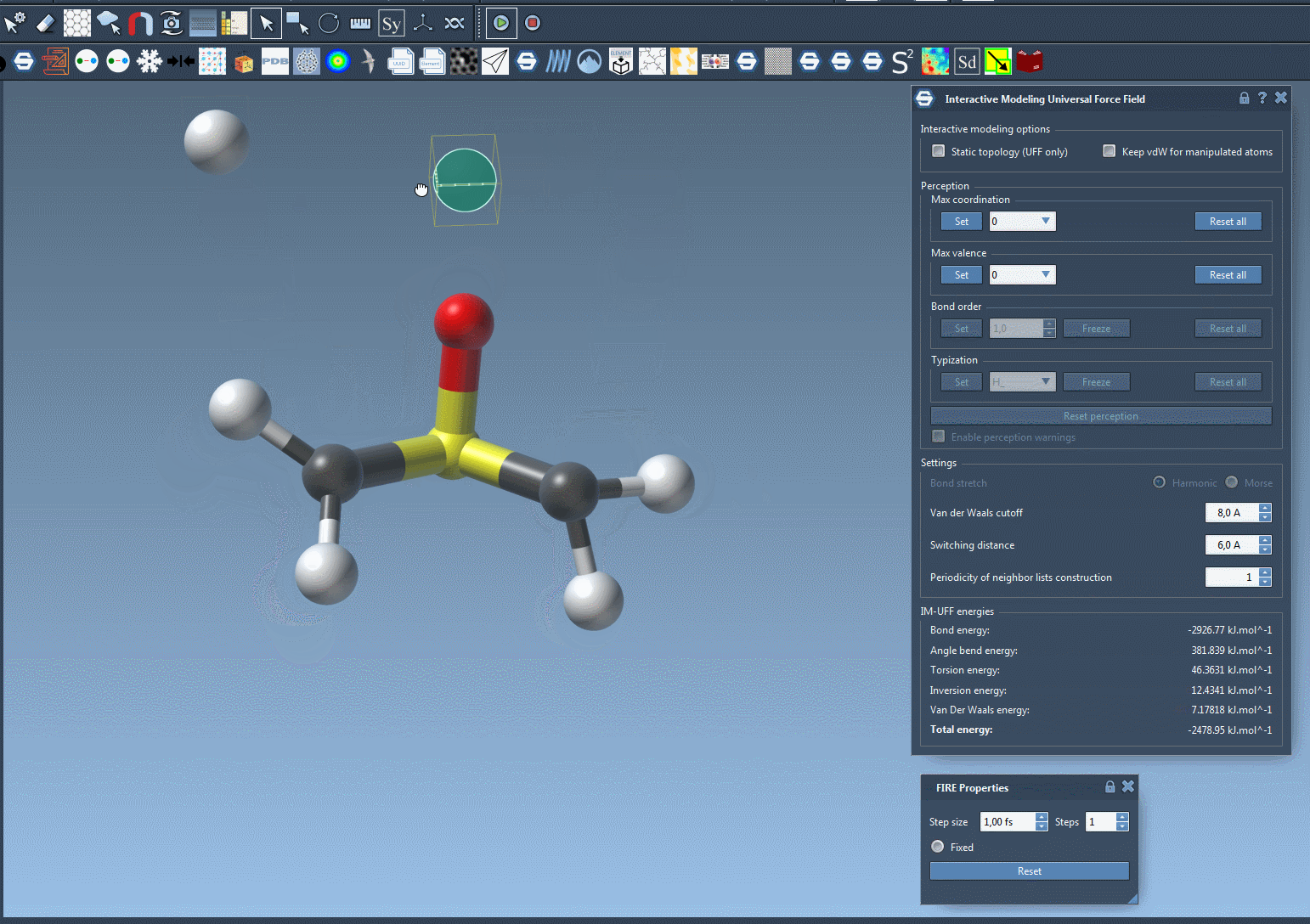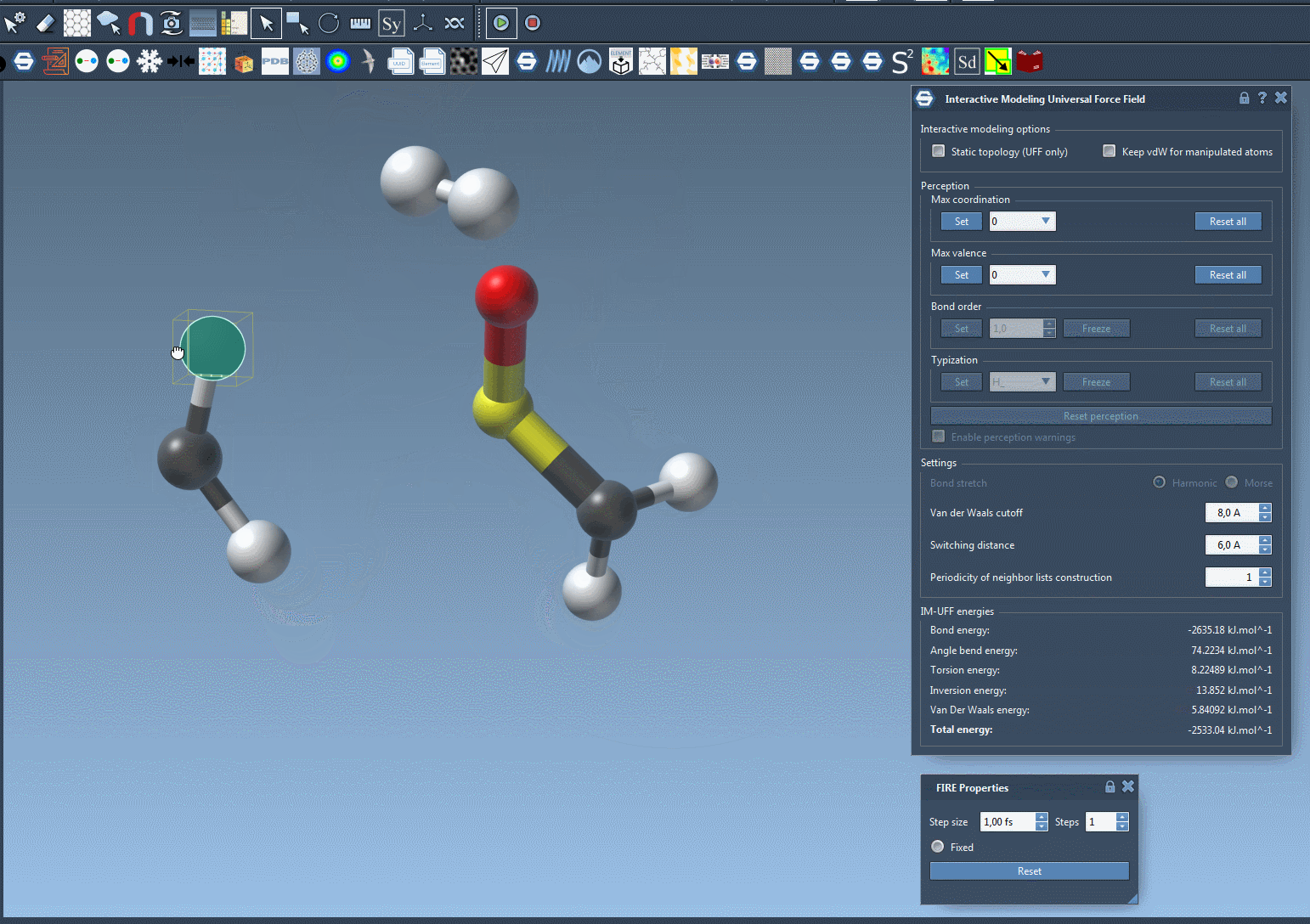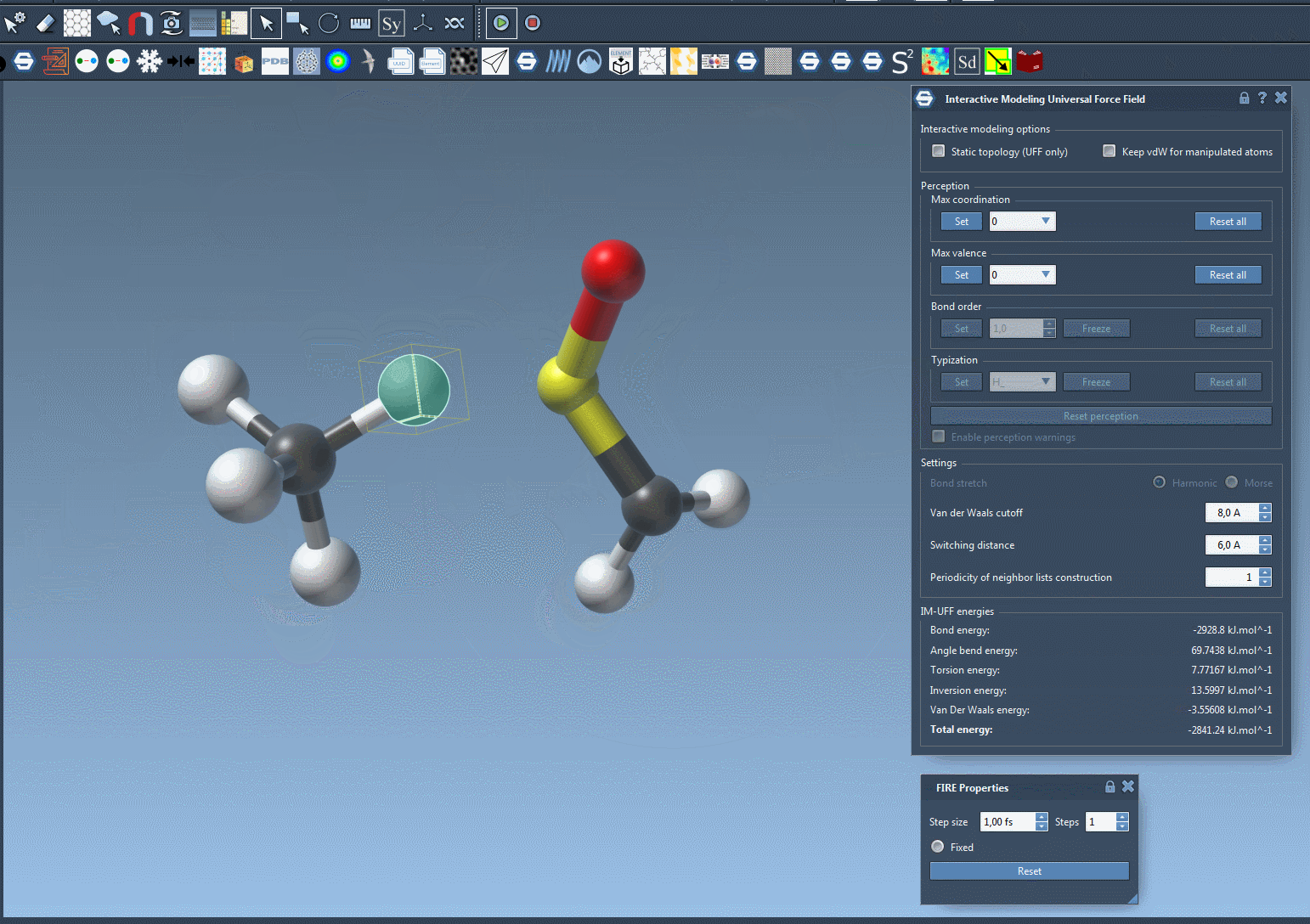
Additional Customization for Advanced Users
IM-UFF offers several customization options to make the tool fit specific needs. Users can set parameters such as the van der Waals cutoff, switching distance, and periodicity of the neighbor list construction. Notably, during IM-UFF’s dynamic topology updates, some settings like maximum coordination and valence can be adjusted to fine-tune the atomic typization process. For more information on customization, you can refer to the UFF tutorial.
Conclusion
IM-UFF is a powerful tool for molecular modelers seeking interactive and intuitive workflows. By enabling seamless topology changes and providing extensive customization options, it significantly reduces the complexity of molecular editing tasks. To explore the full capabilities of IM-UFF, visit the official documentation page.
SAMSON and all SAMSON Extensions are free for non-commercial use. To start using SAMSON, visit SAMSON-Connect.





