Molecular modelers frequently face a daunting challenge: how to efficiently dock and analyze an entire library of ligands against protein receptors while still gaining valuable insights from visualizations and results. This pain point resonates deeply with researchers handling vast datasets and navigating complex workflows. Fortunately, the AutoDock Vina Extended SAMSON Extension offers a solution tailored to streamline this process.
The Power of Ligand Library Docking
The AutoDock Vina Extended SAMSON Extension unlocks the ability to dock not just single ligands but an entire ligand library. Whether working with a small number of compounds or a vast library retrieved from resources like the ZINC library, this tool allows modelers to:
- Set up a receptor or receptor library with optional flexible side chains.
- Apply customizable minimization to ligands, ensuring optimal conditions for docking.
- Lock specific bond types or adjust rotatable bonds for a tailored computational approach.
- Visualize docking results with structure and conformation clarity.
From one interface, users can quickly prepare the docking environment, run simulations that factor in receptor flexibility, and dive headfirst into result analysis and visualization, no matter the scale of their project.
Step-by-Step: Setting Up a Ligand Library for Docking
Using this SAMSON Extension, you can easily dock an entire ligand library. Here’s how to get started:
1. Upload Your Ligand Library
Prepare your ligands in formats like MOL2 or SDF, ensuring they contain the essential structural and chemical data. From the Set ligand section, choose Ligand library and upload the folder containing your library. The extension allows flexibility when dealing with multiple ligands per file.
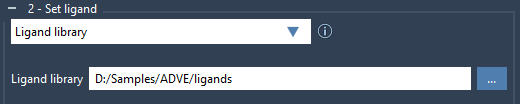
2. Enhance with Ligand Minimization
Ligands from libraries might not be in ready-to-dock condition. Check the Minimize option to enhance ligand geometries by minimizing energy states or adding missing hydrogens. Presets allow you to customize minimization steps and stopping criteria. The minimization process ensures ligands are primed for precise docking.

3. Configure Docking Parameters
Customize configurations such as the exhaustiveness of the search or limits on the number of binding modes generated. Use the Options menu to fine-tune these settings based on your system’s needs. SAMSON’s intuitive interface provides clear control over all parameters related to ligand docking.
4. Run the Docking
Once your ligands and receptors are set, click on Dock library to begin computations. The AutoDock Vina Extended Extension processes your library efficiently, providing accurate results even for large datasets. Results are automatically saved, ensuring no data is lost and computations can be revisited if needed.
5. Analyze Like a Pro
When the docking is complete, visualize results directly in the SAMSON interface. The results folder includes top ligand poses with ranked affinities, molecular configurations, and additional docking parameters.
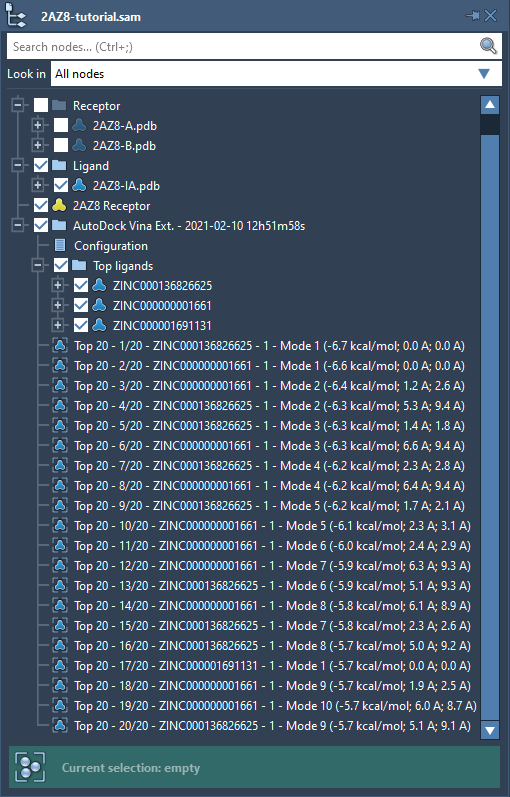
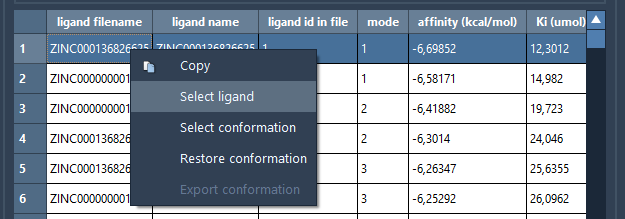
You can thoroughly inspect affinity plots and tables, filter data, or export selected ligands for molecular dynamics simulations or further refinement. Interactive plots provide an intuitive way to identify pivotal docked conformations, letting you restore and examine specific ligand-receptor interactions in detail.
Conclusion
The AutoDock Vina Extended SAMSON Extension simplifies the process of docking ligand libraries and empowers molecular modelers with powerful tools for analysis. By reducing complexity and offering extensive visualization capabilities, it enables researchers to focus on interpreting results rather than managing computational workflows.
For detailed step-by-step instructions on setting up ligand libraries or learning advanced tips, visit the documentation page.
Note: SAMSON and all SAMSON Extensions are free for non-commercial use.





