Understanding the motion of molecules is a critical challenge for molecular modelers. Whether you’re analyzing ligand unbinding, conformation changes in large proteins, or tracking atomic displacement, visualization tools can offer invaluable insights. One such tool in the SAMSON molecular design platform is the Pathlines visual model. Here, we’ll dive into how to use Pathlines to visualize the motion of a molecule’s center-of-mass (COM) along a path, making it easier to analyze complex molecular motion.
What Are Pathlines and Why Use Them?
Pathlines allow you to display the trajectory of a molecule’s COM as it moves across a simulation or pathway. This visualization is particularly helpful in understanding:
- Ligand unbinding and rebinding mechanisms.
- Conformational changes in macromolecules.
- Reaction coordinate workflows or diffusion processes.
Instead of wading through numerical data, you can watch a clean, animated representation of the motion and gain insights more intuitively.
Step-by-Step: Visualizing Molecular Motion with Pathlines
Here’s how you can start visualizing molecular motion using Pathlines in SAMSON.
1. Load the Sample System
If you’re exploring Pathlines for the first time, SAMSON makes it easy to experiment with a prepared example. Download the tutorial sample system by following these steps:
- Click Home > Download within SAMSON.
- Paste this URL into the provided field: https://www.samson-connect.net/documents/046f1acd-c799-40f6-8185-cb4847eff795, then click Download.
The sample system includes the structural model of Lactose permease (1PV7), its ligand Thiodigalactosid (TDG), and precomputed unbinding paths using the Ligand Path Finder.

2. Select Atoms and Paths
To create a Pathline, you need to select the atoms for which you’d like to visualize the motion and one or more paths representing their trajectory. Here’s how:
- Open the Document view. Locate the group of atoms you’d like to analyze, such as the ligand or a specific protein domain.
- Select the desired atoms along with one or more paths. For multiple selections, hold the Ctrl (Windows) or Cmd (Mac) key while selecting.
If nothing is selected, SAMSON will use the entire system and all paths by default.

3. Create a Pathline Visual Model
Once you’ve selected your atoms and paths, create a Pathline visual model to bring their motion to life:
- Navigate to Visualization > Visual model > More… (shortcut: Ctrl/Cmd + Shift + V).
- In the dialog that appears, select Pathline of the center of mass and click OK.
SAMSON will generate a visual pathline representing the motion of the selected atoms’ COM along the specified paths. This visual representation gives you clarity that traditional data analysis methods might lack.
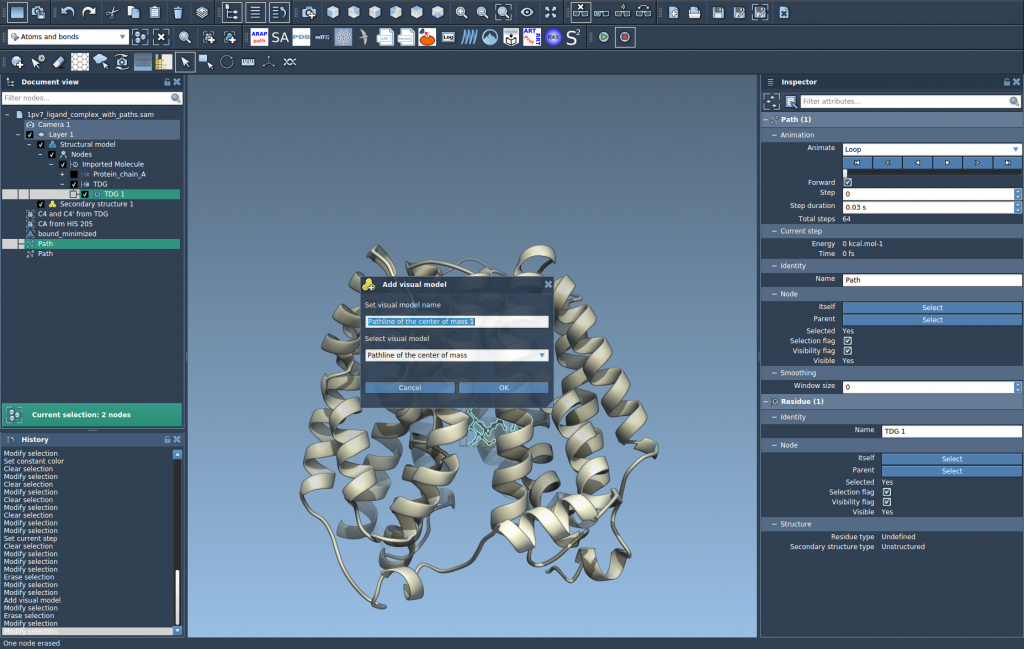
4. Customize and Explore
Pathlines are not just static visuals—they’re interactive and customizable:
- Double-click a path in the Document view to start or stop its animation.
- Right-click a path to access additional options under Path > ….
- Customize the display properties of the Pathline (such as color and thickness) using the Inspector.
This flexibility makes it easy to adapt the visualization to your specific needs.
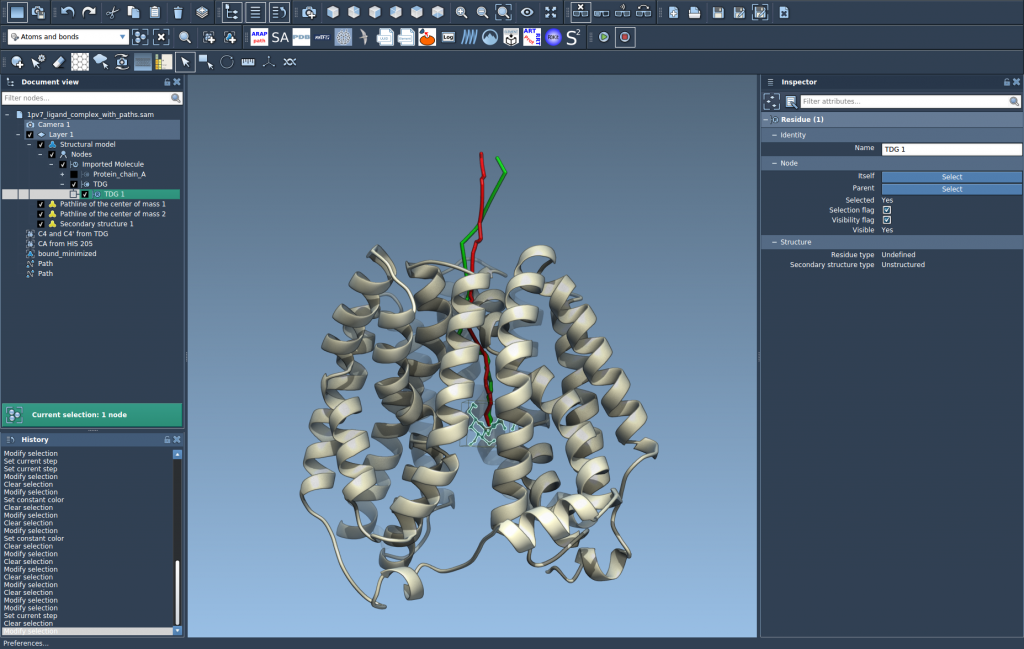
Applications of Pathlines
Pathlines unlock numerous possibilities for molecular modelers, including:
- Visualizing ligand unbinding and rebinding pathways.
- Studying collective domain movements in protein complexes.
- Tracking center-of-mass motion in reaction coordinate workflows.
Whether you’re exploring diffusion, conformational changes, or other molecular phenomena, Pathlines help you better understand your system.
To learn more about using Pathlines in SAMSON, visit the official documentation page: https://documentation.samson-connect.net/tutorials/pathlines/pathlines/.
SAMSON and all SAMSON Extensions are free for non-commercial use. Get SAMSON today at https://www.samson-connect.net.





