One of the prominent challenges molecular modelers often face is understanding how proteins interact and organize in their environments. Whether it’s deciphering crystal structures or biological assemblies, the arrangement of proteins plays a crucial role in modeling workflows, simulations, and molecular design. This is where SAMSON’s Symmetry Mate Editor can be invaluable.
Why Understanding Protein Symmetry Matters
Protein assemblies often function as multimeric units in nature, with repeating subunits forming complex structures. Analyzing symmetry is not just about visualizing these arrangements but also about:
- Exploring biologically relevant protein-protein interactions.
- Reconstructing quaternary structures needed for accurate simulations.
- Identifying potential binding sites at symmetrical interfaces.
- Designing symmetric nanostructures or protein cages for bioengineering and drug development.
Manually reconstructing these assemblies can be time-consuming and prone to errors. That’s why using metadata like PDB symmetry records (CRYST1 and BIOMT) to automatically generate these structures is a huge time-saver.
How the Symmetry Mate Editor Simplifies the Process
The Symmetry Mate Editor in SAMSON provides a streamlined interface to generate and explore protein replicas based on symmetry data. Here’s a quick overview of its features:
- Real-Time Node-Based Editing: Once activated, control nodes appear, representing symmetry transformations. Hovering over these nodes previews the replicas in real time.
- CRYST1 vs. BIOMT Records: CRYST1 describes symmetry in crystal lattice, useful for reconstructing crystal packings, while BIOMT maps biological assembly annotations, helping create biologically relevant replicas.
- Undo and Redo: Mistakes happen, but the editor allows you to undo and redo changes effortlessly.
- Scaling Control: Adjust the number of visible symmetry nodes with Ctrl (Cmd on Mac) + mouse wheel, making navigation and visualization more manageable.
Step-by-Step Guide to Generating Symmetry Mates
- Add the Symmetry Mate Editor extension if it’s not already installed in SAMSON.
- Open a PDB file containing symmetry data.
- Activate the editor from the viewport (Shift + E and search for “Symmetry Mate Editor”) or the Editors menu (General > Symmetry Mate Editor).
- Hover over control nodes to explore transformations. Left-click to generate permanent replicas.
- Want the entire assembly at once? Hold Ctrl (or Cmd on Mac) while clicking any node to apply all transformations simultaneously.
An example animation demonstrates how smooth this process can be:
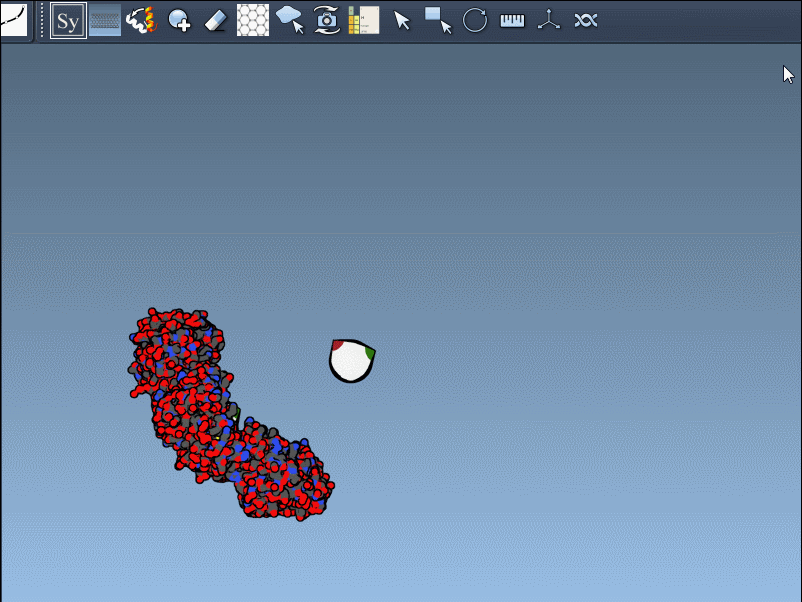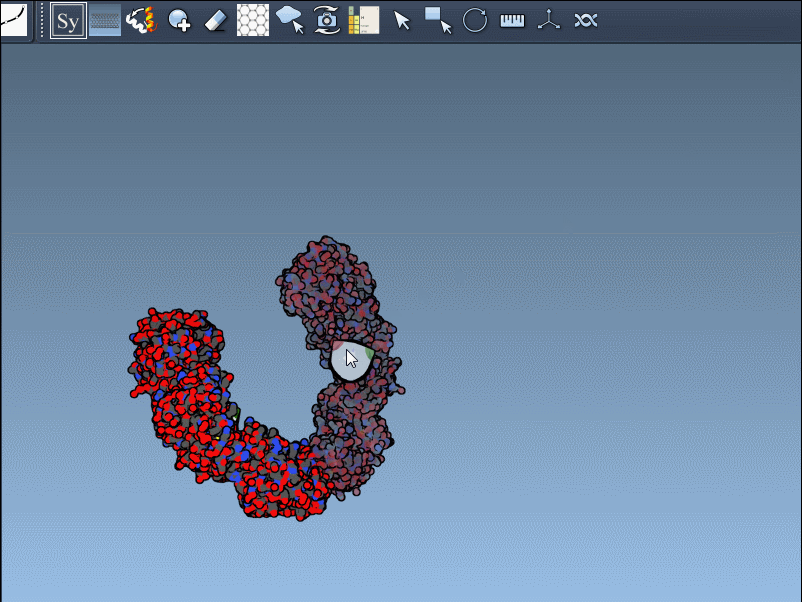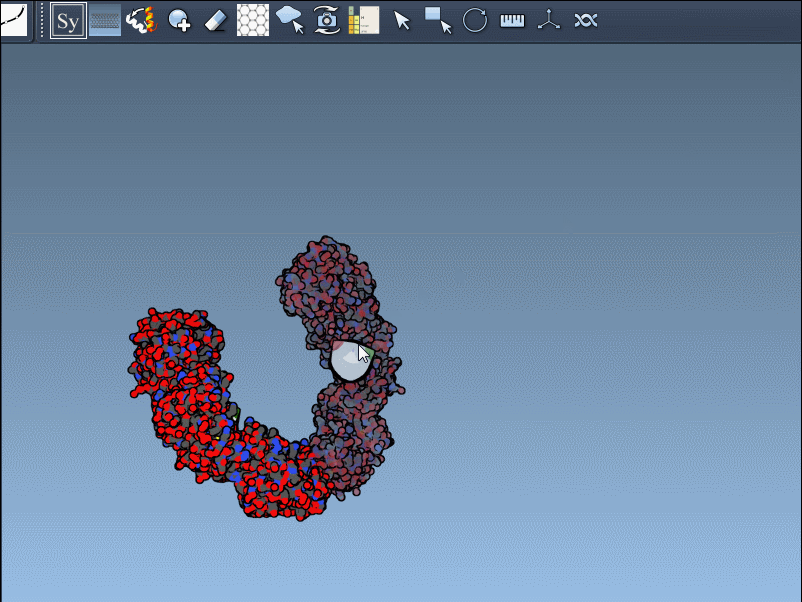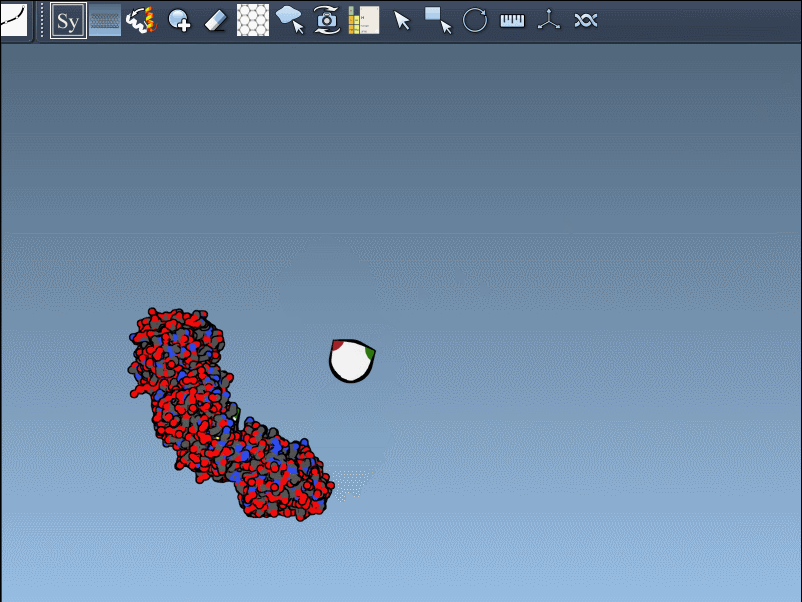
Enhancing Visualization
To further improve clarity, combine the Symmetry Mate Editor with other tools available in SAMSON:
- Ribbons Model: Use this to colorize chains for better visual distinction. It’s particularly useful in complex structures.
- Symmetry Detection: Analyze axes and RMSD for deeper insights into structural symmetries.
Broad Applications
The possibilities with the Symmetry Mate Editor are immense:
- Model complete oligomeric assemblies accurately.
- Design symmetric protein cages or nanostructures, paving the way for novel drug designs.
- Prepare systems for detailed molecular dynamics (MD) simulations, particularly when studying symmetric chains.
Key Takeaway
By leveraging symmetry metadata stored in PDB files, SAMSON’s Symmetry Mate Editor reduces complexity and makes the study of protein interactions and assemblies considerably easier. To dive deeper and access detailed guidelines, visit the original documentation page.
SAMSON and all SAMSON Extensions are free for non-commercial use. Download SAMSON for free at SAMSON Connect.





