When working in molecular modeling, one common task is exploring how changes to specific molecular fragments impact properties like binding affinity or interactions with a target protein. This is where the SMILES Manager in SAMSON comes in handy. In this blog post, we’ll focus on a key feature: how to identify and replace patterns in molecular structures to generate analogs quickly and efficiently.
What’s the challenge?
Imagine you have a molecule, and you suspect that swapping out a specific fragment — say, aromatic carbons — with another atom or group might reveal something interesting about the molecule’s behavior or interactions. Doing this manually for multiple patterns or analogs can be tedious and error-prone. The SMILES Manager simplifies this process with automated tools and a user-friendly interface.
Highlighting patterns: the first step
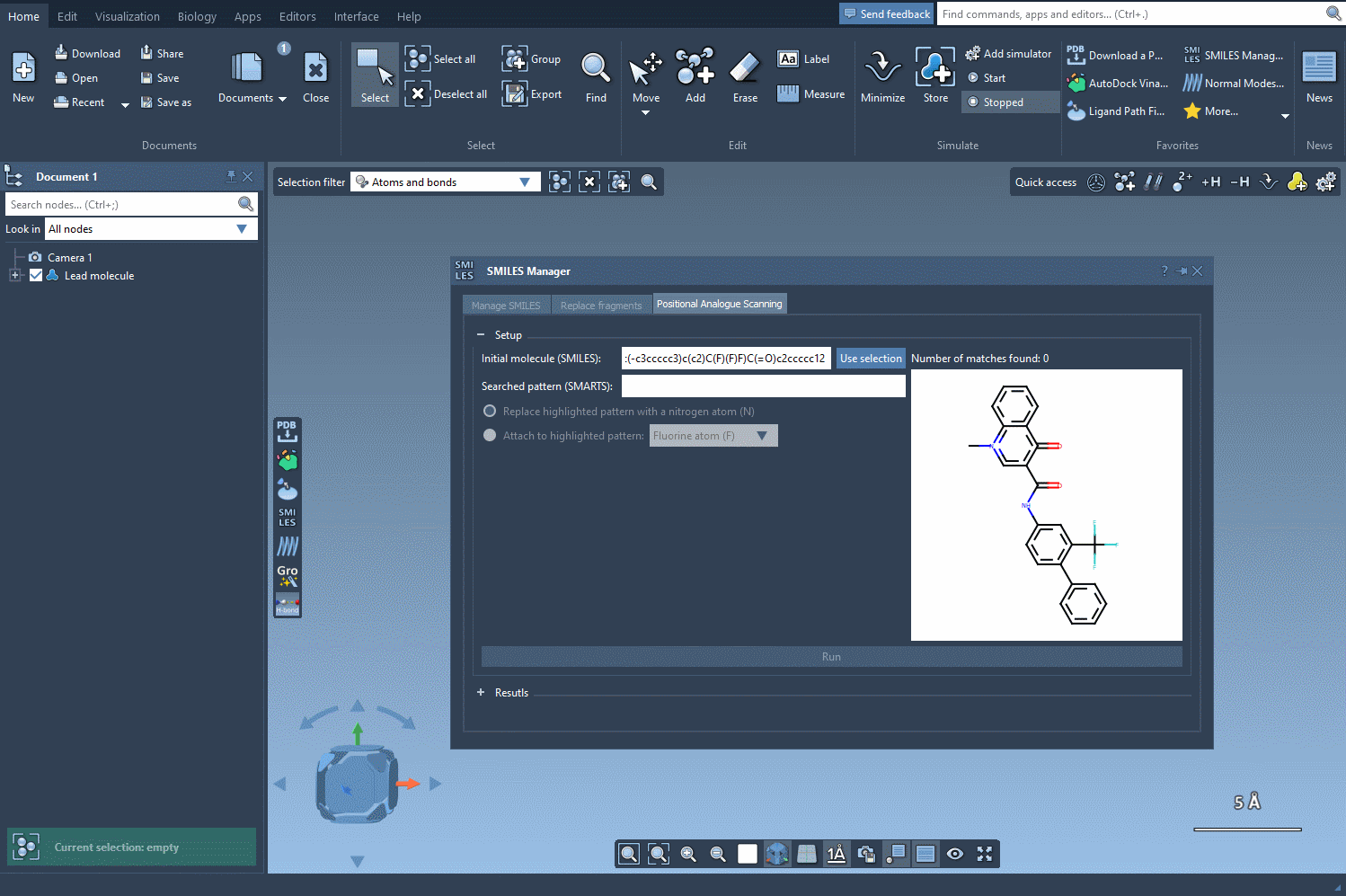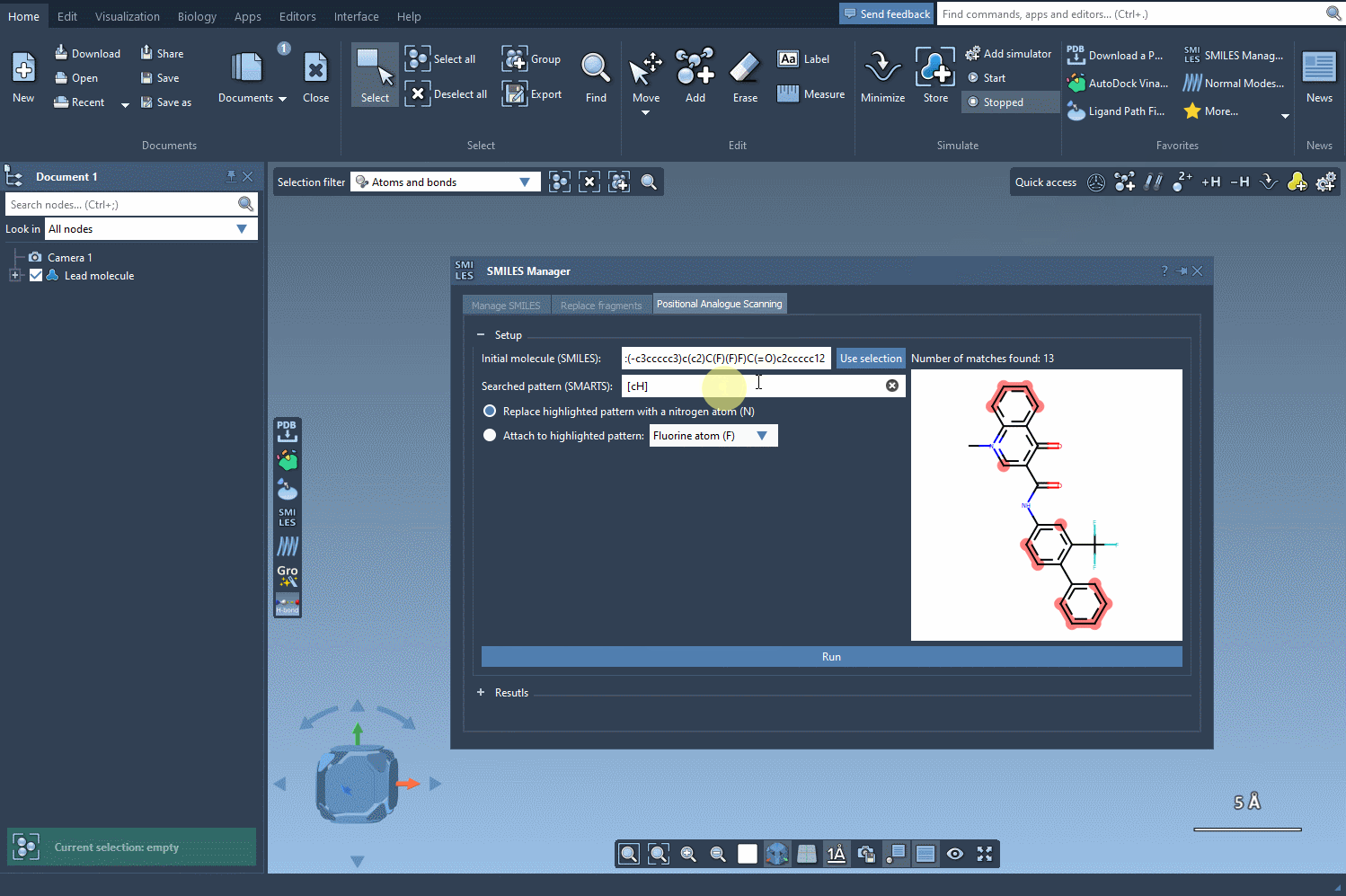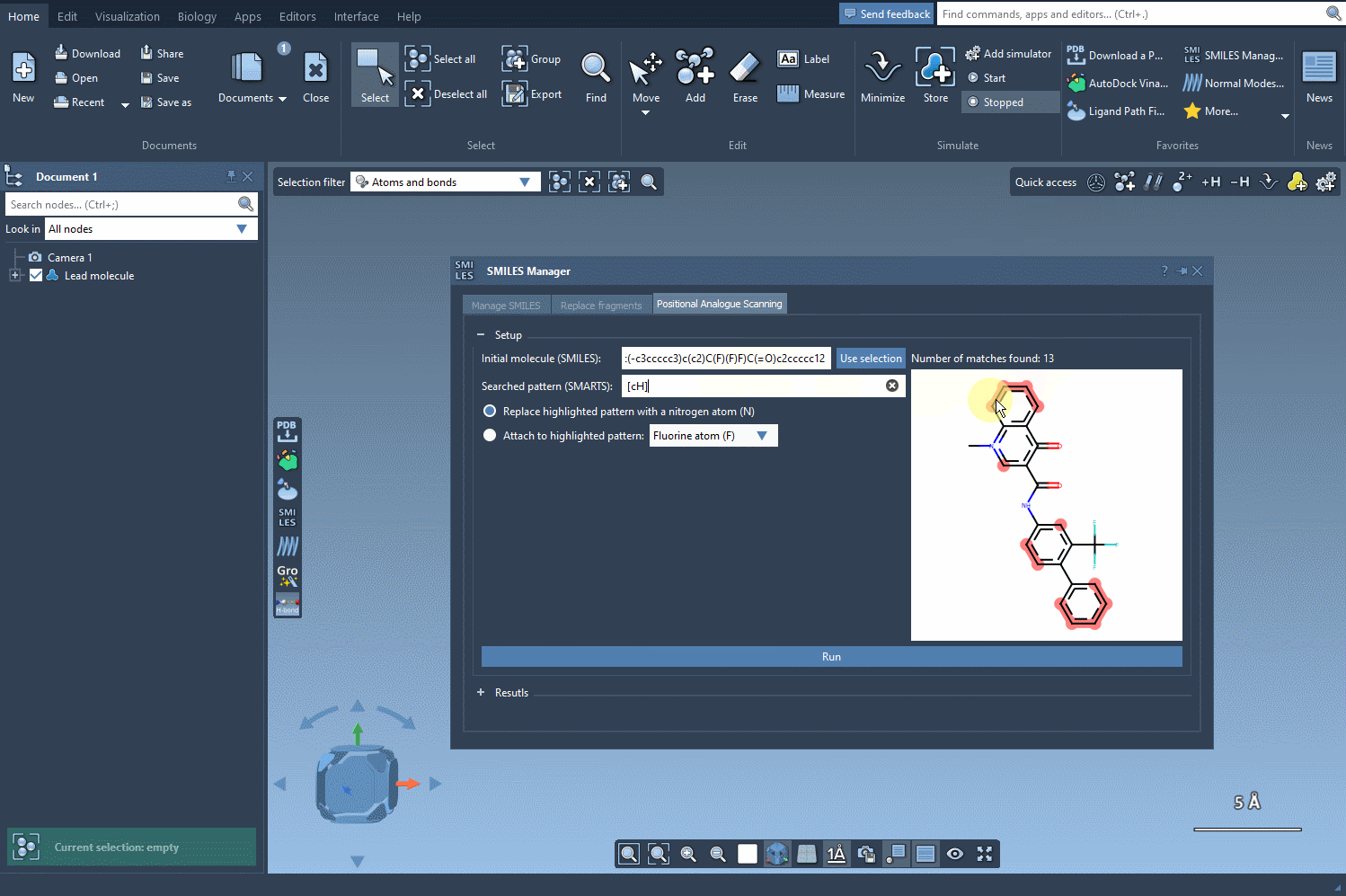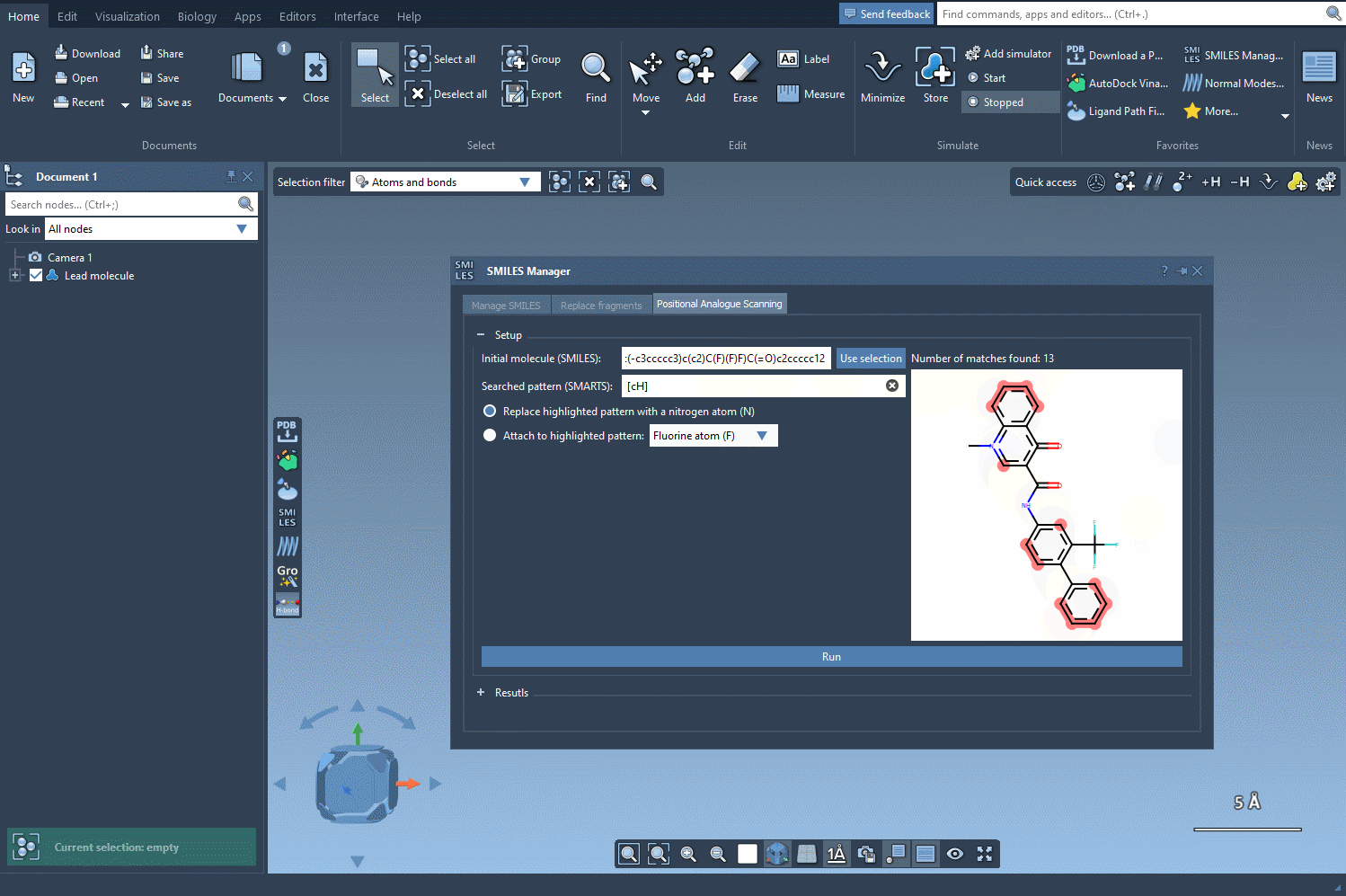
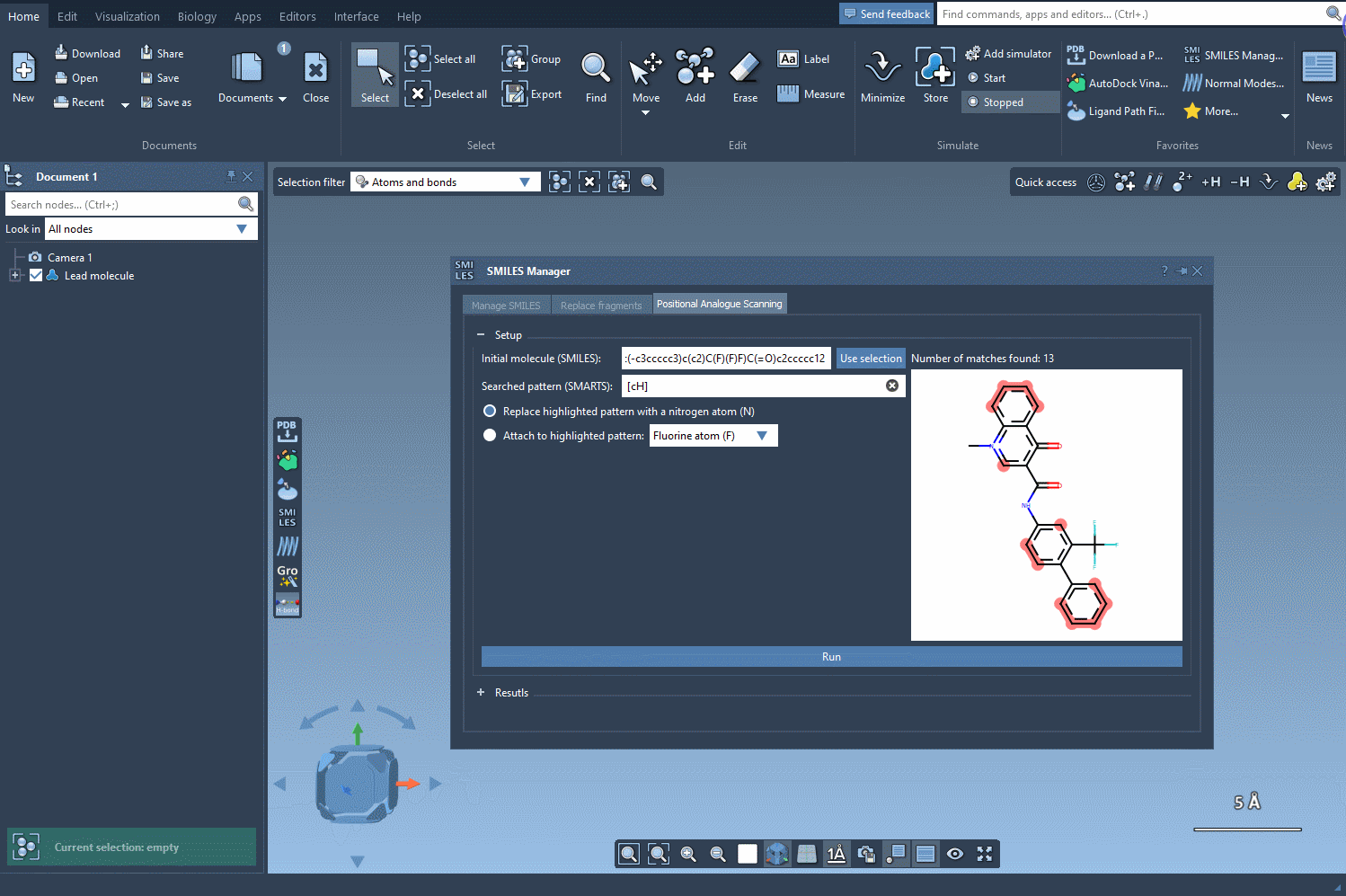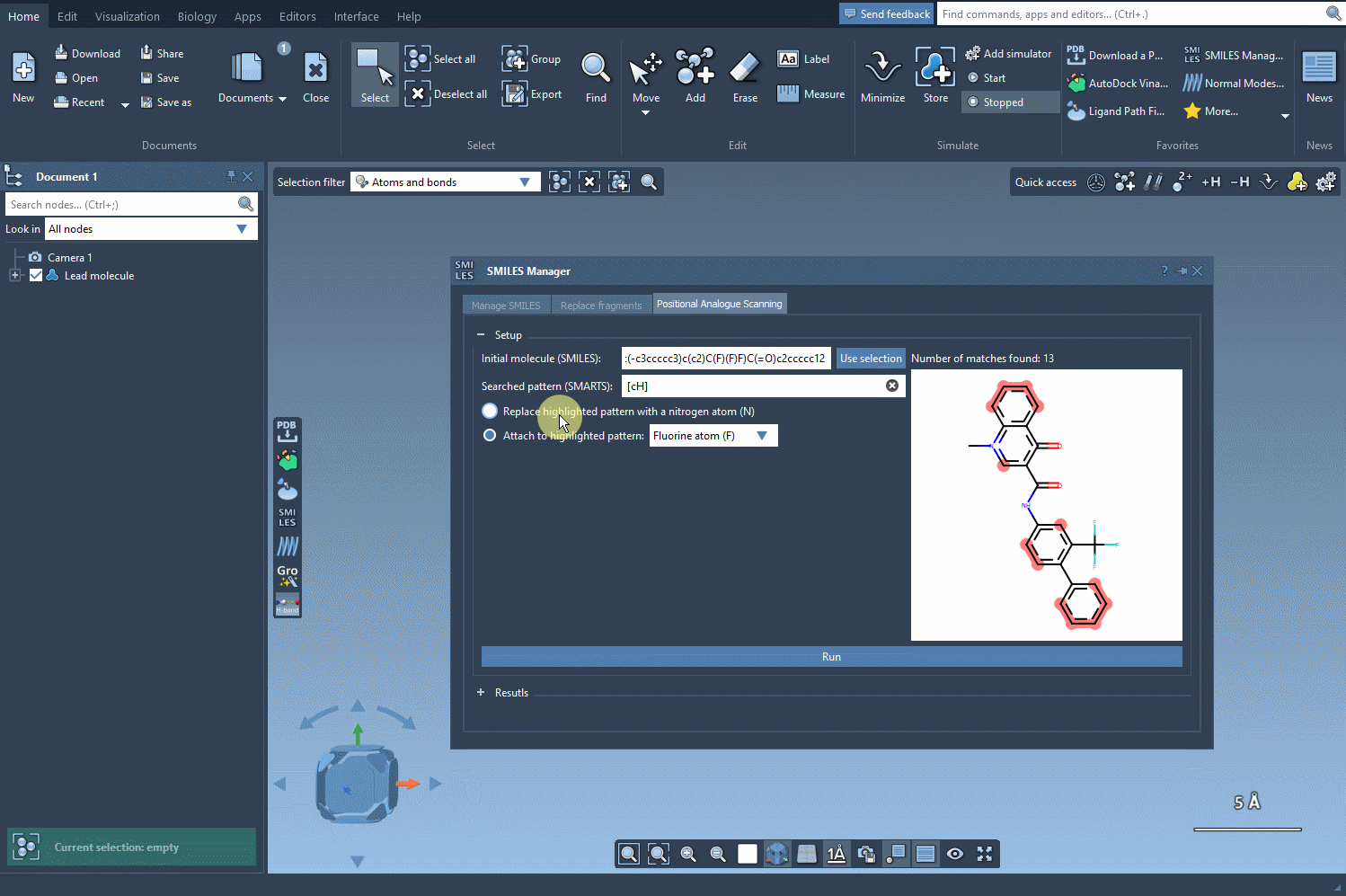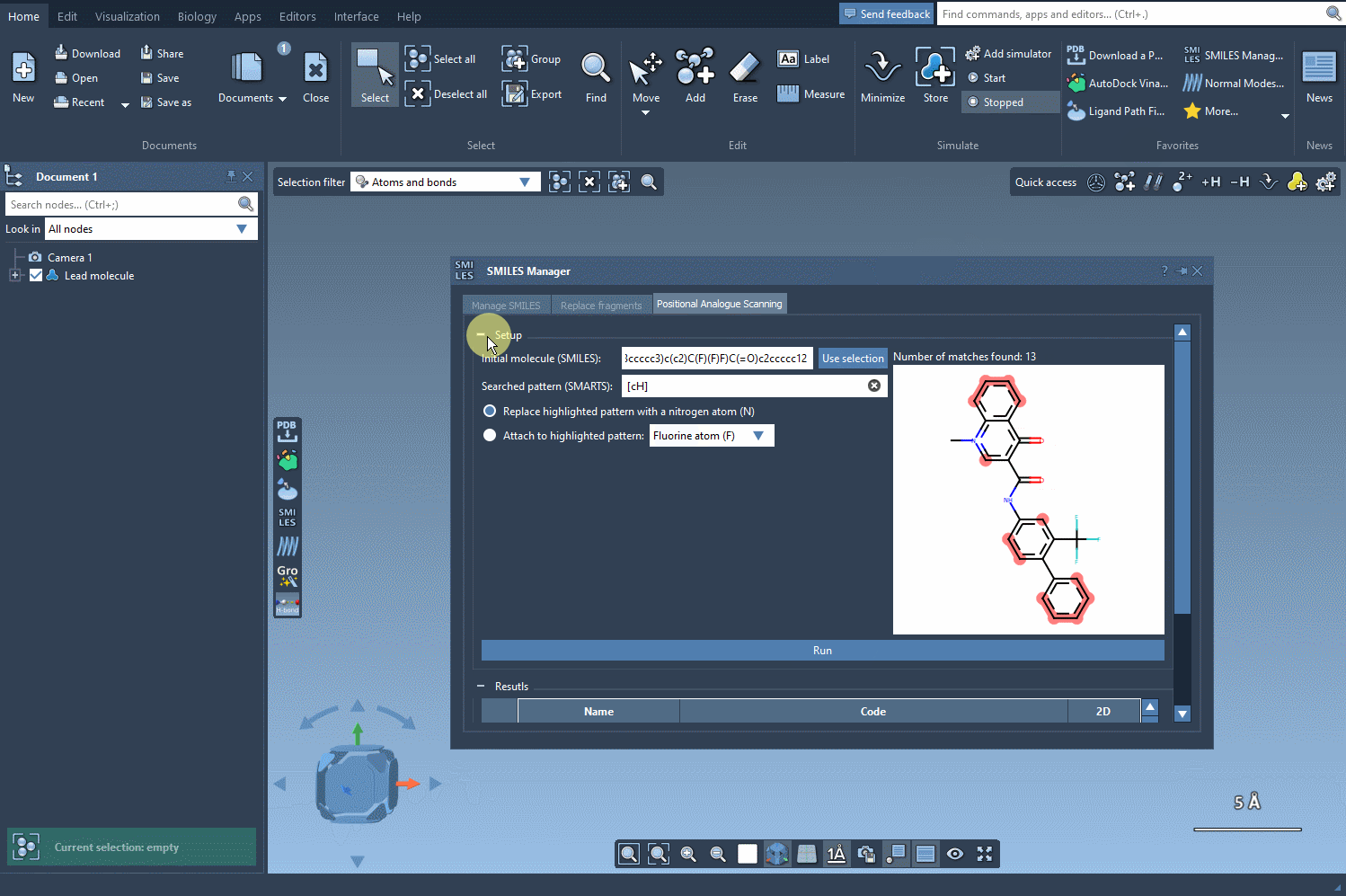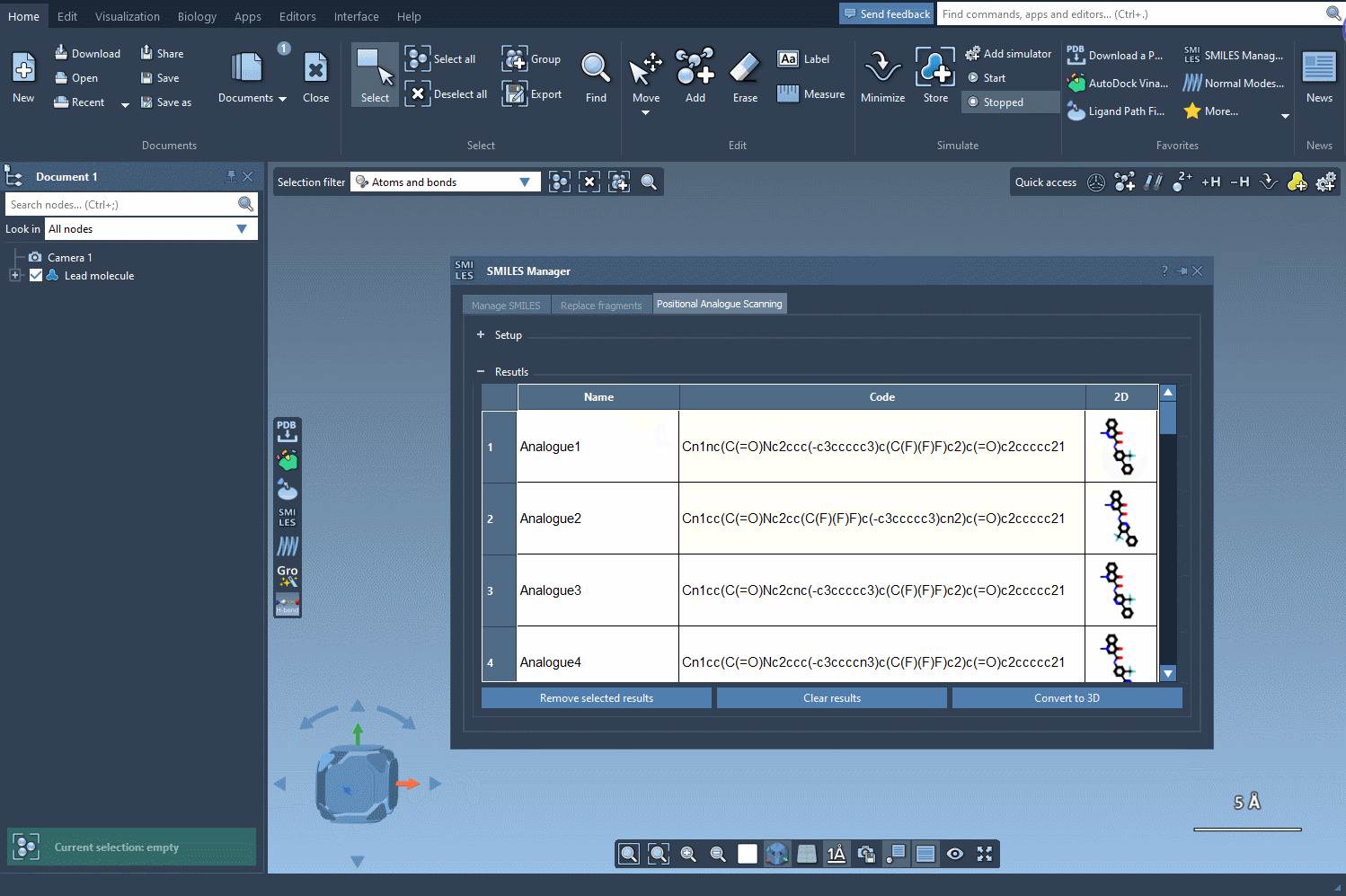
To begin, start with your molecule of interest, which you can define either by entering its SMILES code or by selecting it directly within SAMSON and clicking the Use selection button.
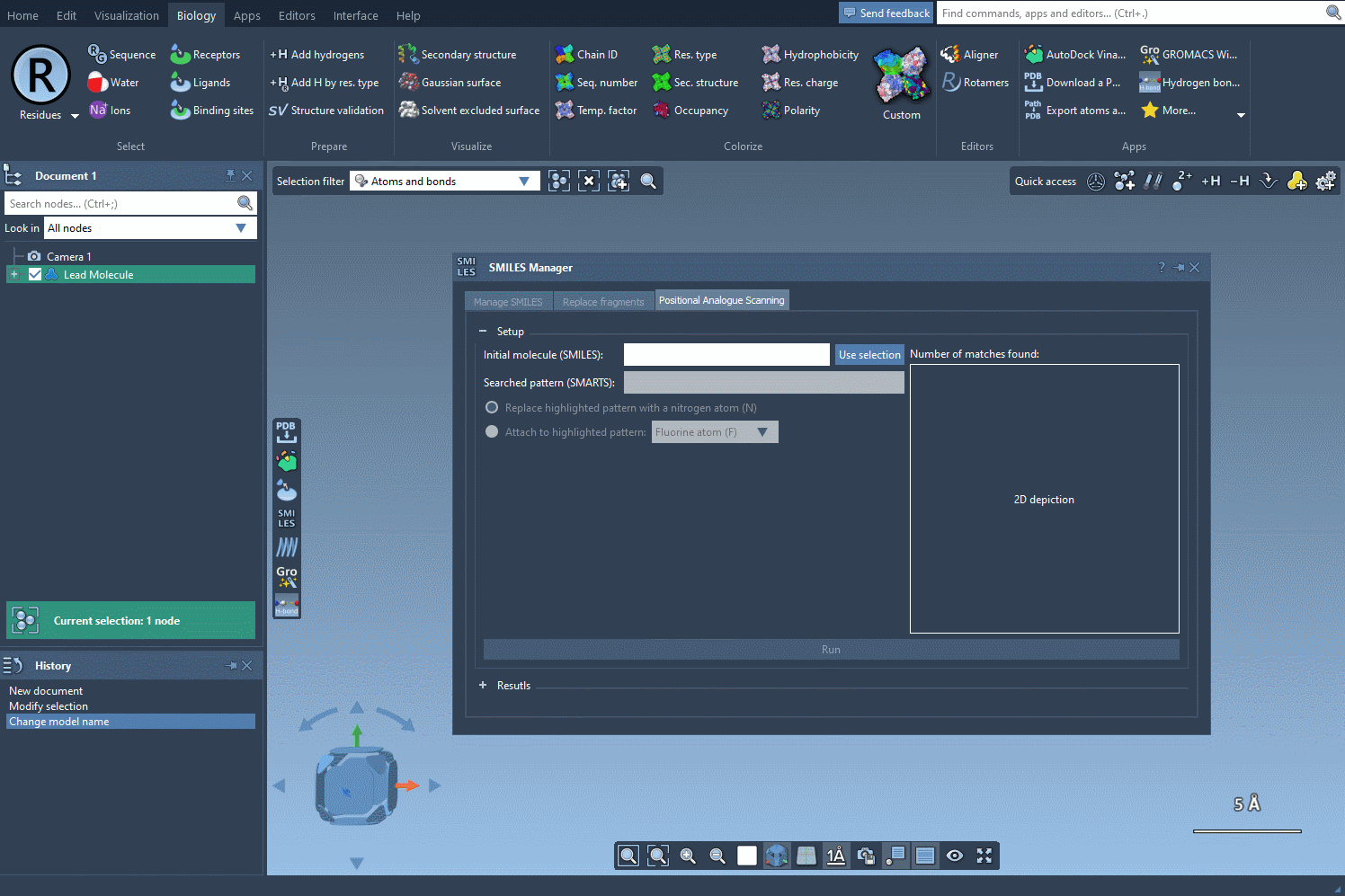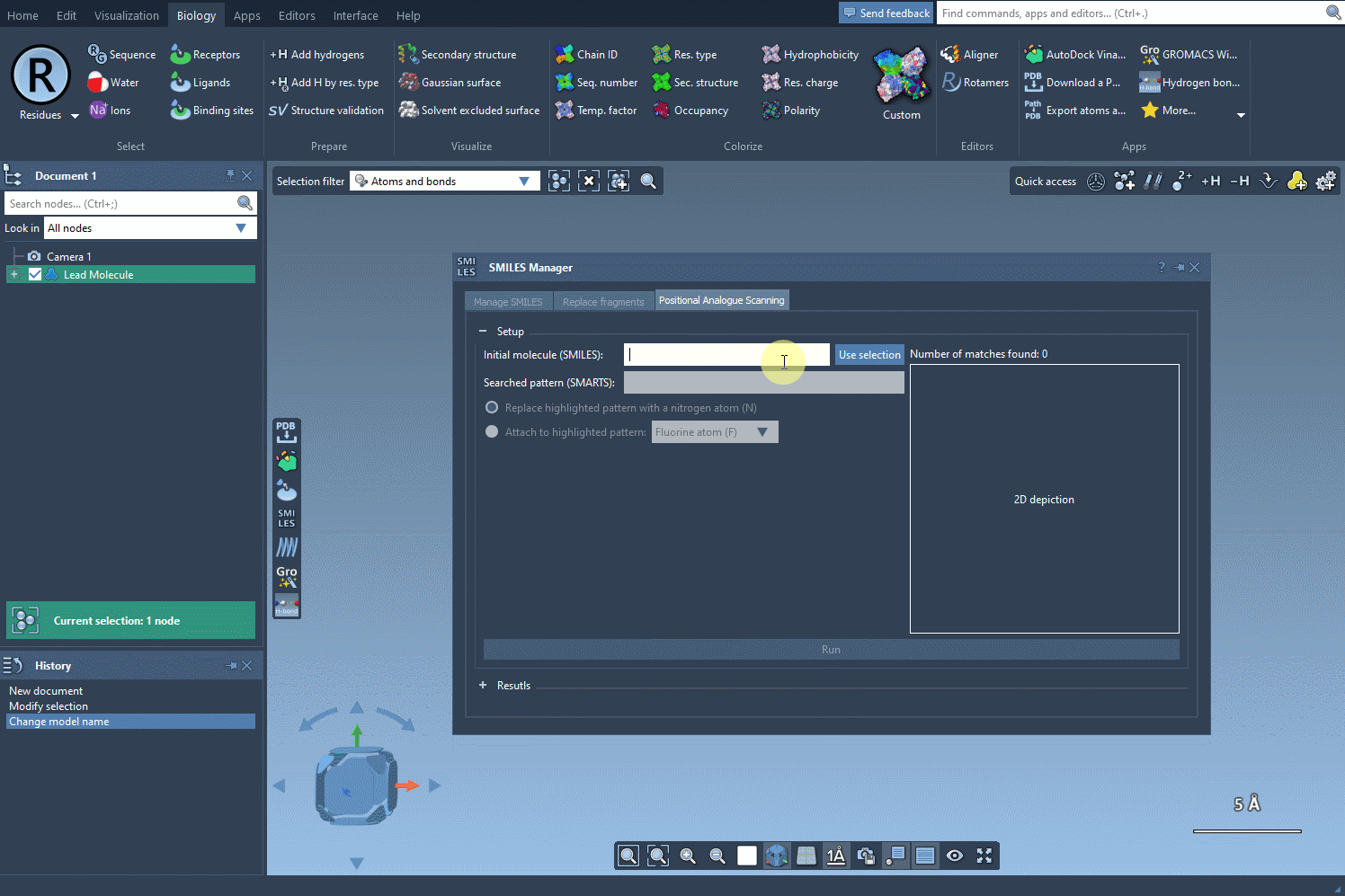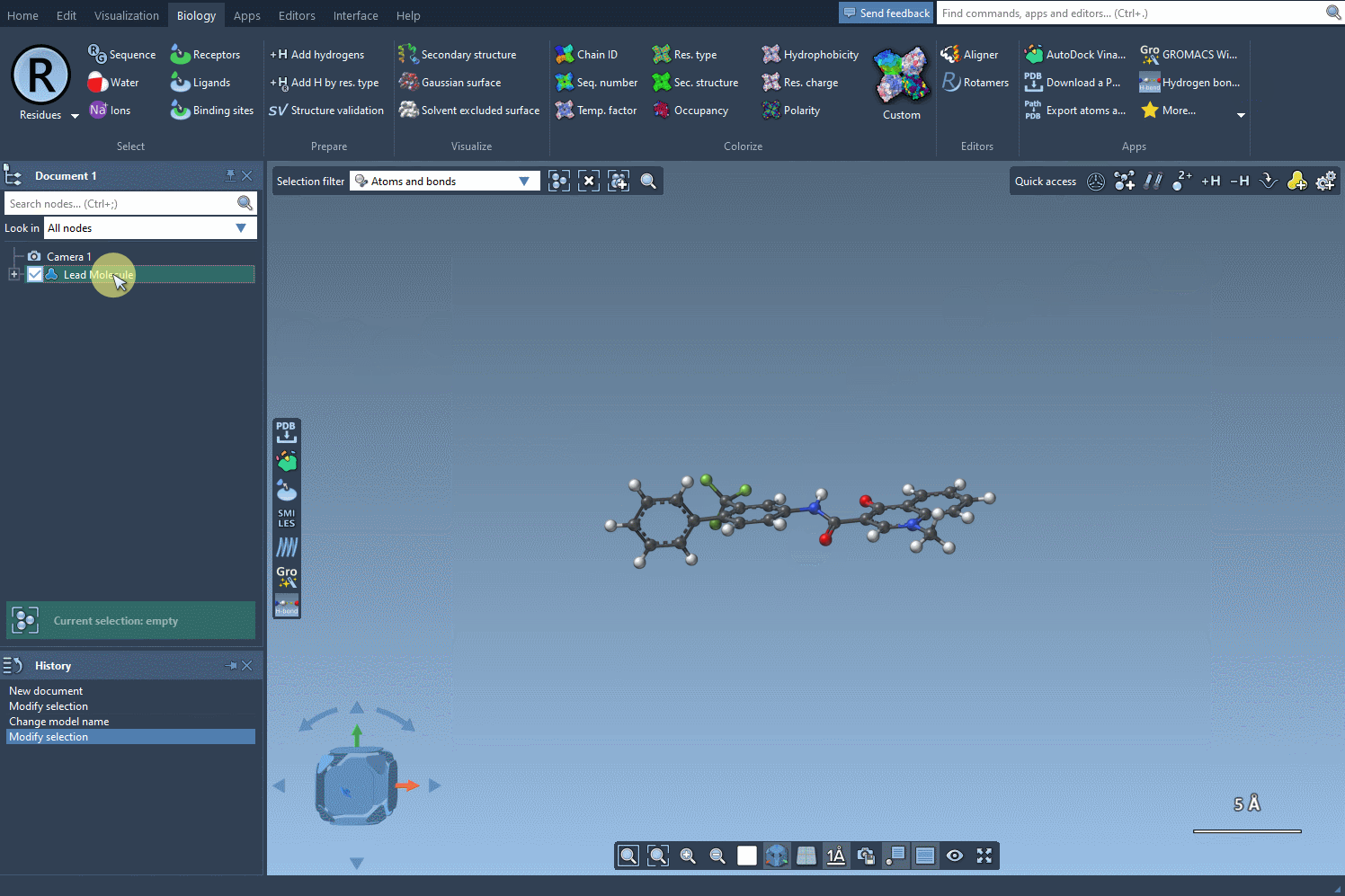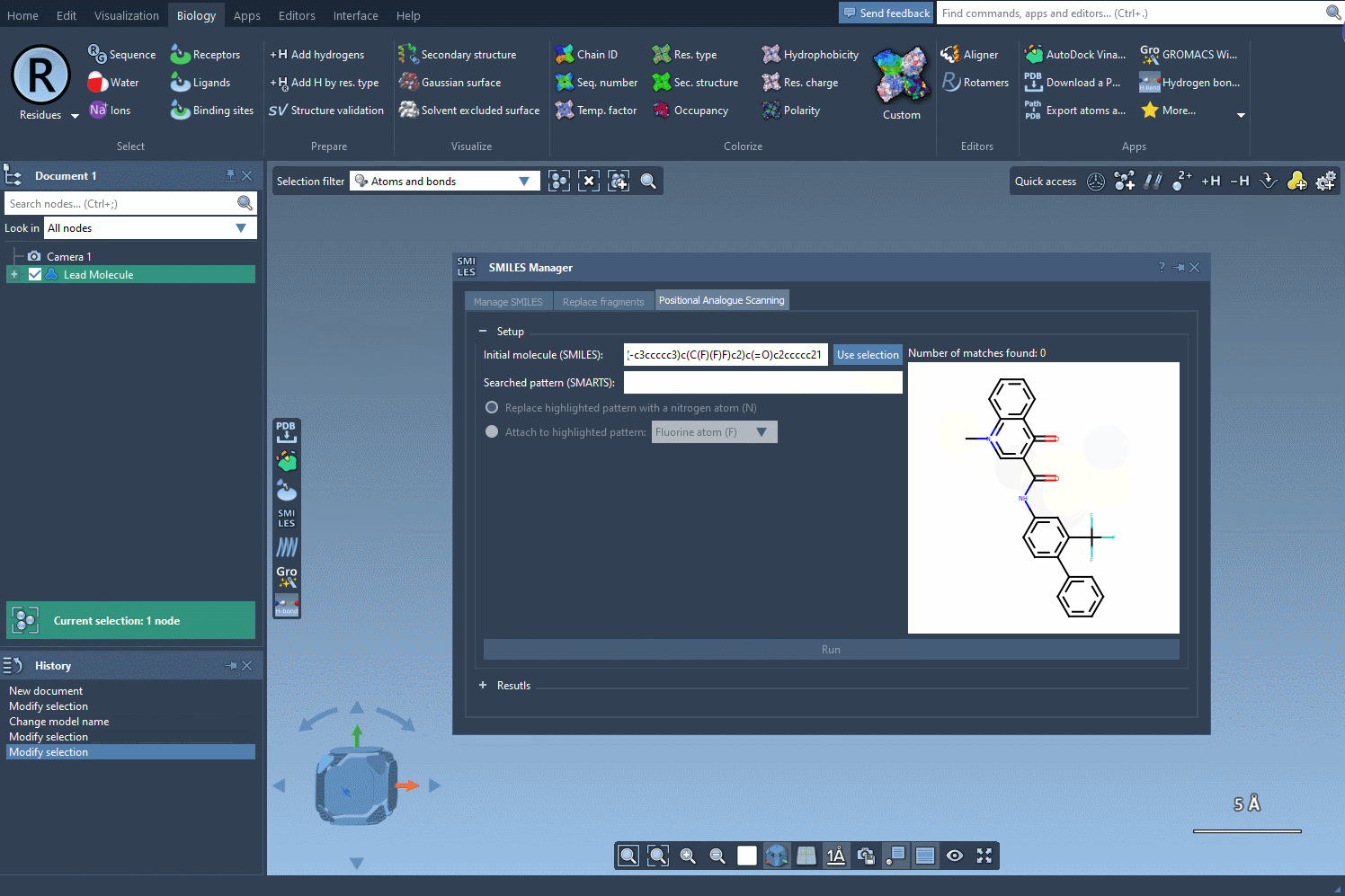
Next, define the specific pattern you want to search for in the molecule. This step allows you to focus on particular parts of the structure that you’re interested in modifying. For example, in one common use case, aromatic carbons are defined as the pattern using the SMARTS code [cH]. Once you input this pattern, the SMILES Manager highlights all instances of it in the molecule.
Replacing patterns
After identifying the pattern, choose how you’d like to modify it. You can replace it with alternative atoms like nitrogen (N) or attach groups such as fluorine (F) or methyl (CH3). With just a click on the Run button, the SMILES Manager generates a set of analogs with the specified substitutions. This process is not only swift but also ensures consistent and accurate results.
Beyond generating SMILES codes, the SMILES Manager also provides depictions of the resulting analogs, which can be further analyzed or edited.
What’s next?
Once you have the generated analogs, you can move on to studying them in greater detail. For instance, you could convert the analogs into 3D structures and perform docking studies using extensions like Autodock Vina Extended. This lets you investigate how the substitutions influence molecular interactions with a target protein. Such insights are invaluable in domains like drug discovery and material research.
This streamlined approach saves time and allows molecular modelers to focus on analysis rather than repetitive manual edits.
Curious to explore more? Check out the full tutorial on positional analogue scanning using the SMILES Manager here: https://documentation.samson-connect.net/tutorials/smiles-manager/perform-positional-analogue-scanning-using-the-smiles-manager-element/.
Note: SAMSON and all SAMSON Extensions are free for non-commercial use. You can download SAMSON at https://www.samson-connect.net.





