For molecular modelers working on reaction coordinate studies, free energy calculations, or tracking ligand movements, exporting precise atomic trajectories is a recurring challenge. Without the right tools, this task can become time-consuming and error-prone. If you’ve been facing this issue, SAMSON’s Export Along Paths extension offers a straightforward, efficient solution.
Why Export Atomic Trajectories?
Atomic trajectories are pivotal for understanding molecular interactions, ligand pathways, and free energy profiles. When studying ligand unbinding processes or reaction mechanisms, generating a reliable reaction coordinate or tracking specific atom groups is critical. SAMSON’s Export Along Paths app simplifies this by allowing you to export atomic positions along predefined paths in different formats.
Getting Started
Before diving in, ensure that you have the Export Along Paths extension installed. If it’s not part of your setup yet, head over to its extension page, click Add, and restart SAMSON to activate the feature.
Step 1: Load a Sample System
To try out the app, you can load a sample system provided in the documentation. Navigate to Home > Download in SAMSON and paste this link: https://www.samson-connect.net/documents/046f1acd-c799-40f6-8185-cb4847eff795. Click Download to load a structural model of Lactose permease (1PV7) and its ligand Thiodigalactosid (TDG) along with precomputed unbinding paths generated by the Ligand Path Finder. You can visualize the protein, ligand, and paths in the Document view under the respective names such as Protein_chain_A and TDG. Here’s a visual reference:

Step 2: Open the Export Along Paths App
To access the Export Along Paths app, go to Home > Apps > All > Export Along Paths. Alternatively, press Shift + E to find it quickly using the search feature. Here’s what the app interface looks like:
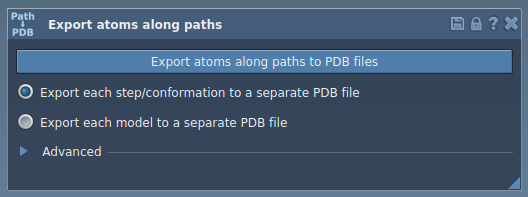
Exporting Atomic Trajectories
SAMSON’s app offers two main options for trajectory export:
Option 1: Export All Atoms
- In the Document view, select one or more paths for trajectory export.
- Choose your export mode:
- All frames in a single PDB file
- Each frame as a separate PDB file
- Click Export atoms along paths to PDB files. You’ll be prompted to choose a destination folder and file prefix. Optionally, you can adjust the frame export interval in the Advanced section.
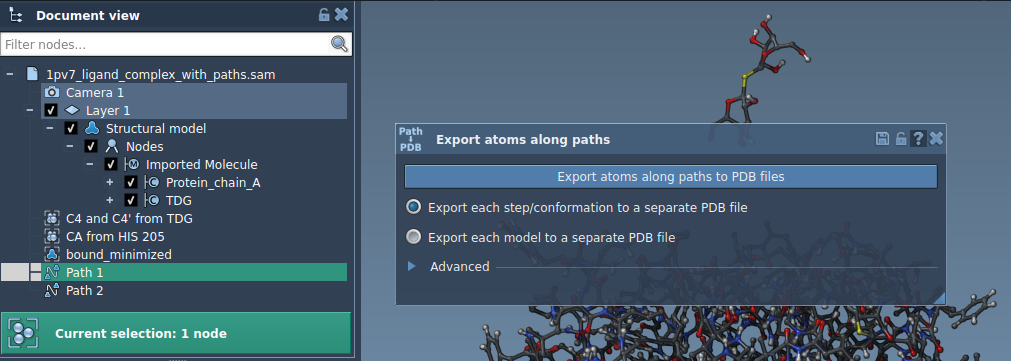
Option 2: Export a Subset of Atoms
For targeted analysis, you might only want to export trajectories of a specific group of atoms, like the ligand or binding site. Here’s how:
- Expand the Advanced panel in the app interface.
- Select specific atoms in the Document view, for instance,
TDG. You can add this selection to the export list as a named model by clicking Add.
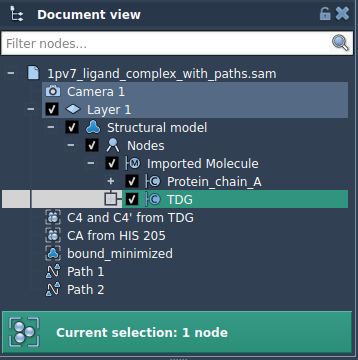
Multiple groups of atoms can be named, managed, and exported simultaneously. When ready, select the paths and export format before initiating the extraction.
Applications in Molecular Modeling
Exporting atomic trajectories is invaluable for tasks such as:
- Generating reaction coordinate files for accurate free energy profiling.
- Visualizing ligand exit or entry pathways to improve molecular designs.
- Focusing on specific atom groups (e.g., ligand, backbone) for detailed analysis.
- Creating intermediate states for simulation checkpoints or presentations.
To delve deeper and understand how you can integrate this tool into your workflow, visit the full documentation.
Note: SAMSON and all SAMSON Extensions are free for non-commercial use. Download SAMSON today at SAMSON Connect.





