For molecular modelers, capturing and analyzing atomic movements along defined paths can be invaluable for understanding reaction mechanisms, ligand binding, and free energy calculations. However, extracting accurate trajectory data efficiently is often a complex task. The Export Along Paths extension in SAMSON provides a streamlined solution to help you export atomic coordinates (for all or selected atoms) along pre-defined paths seamlessly. Here, we’ll dive into how this capability empowers your molecular modeling workflows.
Why Export Atomic Trajectories?
Imagine you’ve traced a ligand’s unbinding pathway or calculated a reaction coordinate. The next step is often to export atomic movements along these paths for external analysis, such as generating reaction coordinate files for free energy profiling. This precise data extraction is critical for simulation and visualization tasks. The Export Along Paths extension helps you do just that, whether for all atoms or a subset.
Step-by-Step: Exporting Atomic Trajectories
Step 1 – Open the App
To begin, open the Export Along Paths app in SAMSON through Home > Apps > All > Export Along Paths. Alternatively, use the quick access feature (Shift + E) to locate the app efficiently.
![]()
Step 2 – Export All Atoms or Selected Atoms
The flexibility of the app makes it a powerful tool. You can choose one of two main options:
Option 1: Export All Atoms
- Specify the path(s) you want to export in the Document view.
- Set the export mode:
- Combine all frames in one PDB file.
- Export each frame as a separate PDB file.
- Click Export atoms along paths to PDB files. You’ll be prompted to select the destination folder and a file prefix to complete the operation.
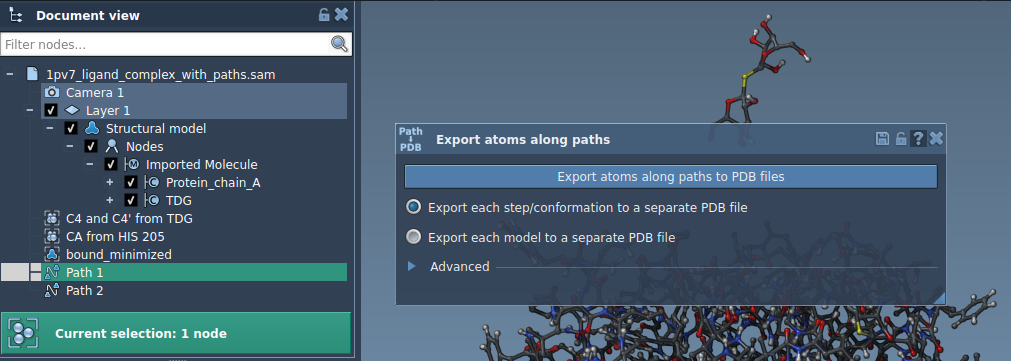
Option 2: Export a Subset of Atoms
If you’re interested in exporting only specific groups of atoms (e.g., a ligand like TDG), follow these steps:
- Open the Advanced panel and select the subset of atoms directly in the Document view.
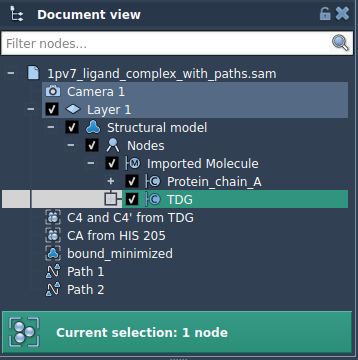
- Click Add to define the selection as a model for export.
Repeat this process as needed to add more atom groups. You can rename each model, manage included atoms through a context menu, and reset models if necessary.
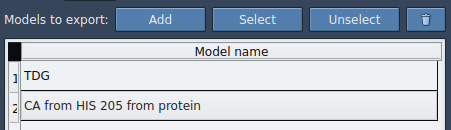
Finally, select your path(s), choose the output mode (single or multiple PDB files), and hit Export atoms along paths to PDB files. The app will prompt you to specify the destination folder and naming format for your files.
Applications for Molecular Modelers
The ability to export atomic trajectories along paths opens the door to multiple use cases, such as:
- Generating reaction coordinate files for free energy profiling.
- Exporting trajectory data for ligand exit/entry pathways.
- Visualizing intermediate states during molecular dynamics.
- Isolating specific subsets of atoms (e.g., a binding site, backbone, or ligand).
Conclusion
The Export Along Paths extension is a practical tool for molecular modelers seeking to extract atomic movements along custom pathways easily. Whether you’re focused on reaction mechanisms or ligand trajectory analysis, this app simplifies data export and empowers your simulations.
To learn more, visit the full documentation page: Export Along Paths.
Note: SAMSON and all SAMSON Extensions are free for non-commercial use. Get SAMSON at SAMSON Connect.





