Analyzing protein-protein interactions and constructing realistic molecular models are everyday tasks for molecular modelers. However, reconstructing biological assemblies or understanding how proteins interact in their natural symmetric forms can be a daunting task without the right tools. The Symmetry Mate Editor in SAMSON provides an elegant solution to this challenge by letting you visualize and generate symmetry mates directly from PDB files, helping you interpret protein arrangements with ease.
Why focus on symmetry mates?
Symmetry mates provide critical insights for both structural biology and molecular design workflows:
- Visualizing biological assemblies: Biological assemblies, such as oligomers or protein complexes, are often stored in fractional components within crystallographic units. Symmetry mates allow you to reconstruct these full assemblies quickly.
- Exploring protein-protein interactions: Understanding the interfaces between proteins helps in predicting binding sites or analyzing docking outcomes.
- Symmetric design: Scientists frequently need tools to construct symmetric protein complexes, scaffolds, or nanostructures for protein engineering and drug delivery applications.
Activating and working with the Symmetry Mate Editor
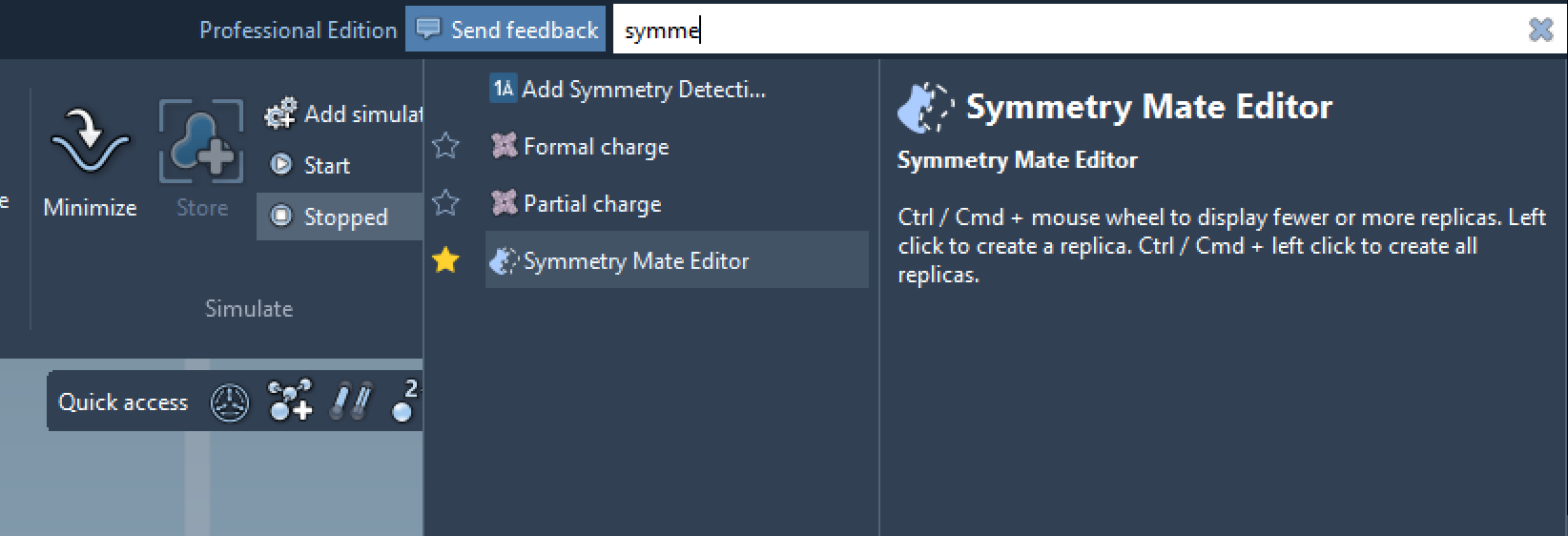
The Symmetry Mate Editor can be activated in SAMSON through the Find everything feature (press Shift + E and search for “Symmetry Mate Editor”) or via the viewport editors menu. Once activated, control nodes representing symmetry transformations become visible in the viewport. These nodes act as references to explore and generate replicas.
With interactive tools like Ctrl/Cmd + mouse wheel, you can scale the number of visible control nodes. Hovering over these nodes previews the replicas in real-time—an excellent feature for seeing the full protein assembly and symmetry transformations without clutter.
Real-world example: 1BRE
For example, when working with the 1BRE protein structure, scaling control nodes or hovering over them reveals real-time symmetry previews:
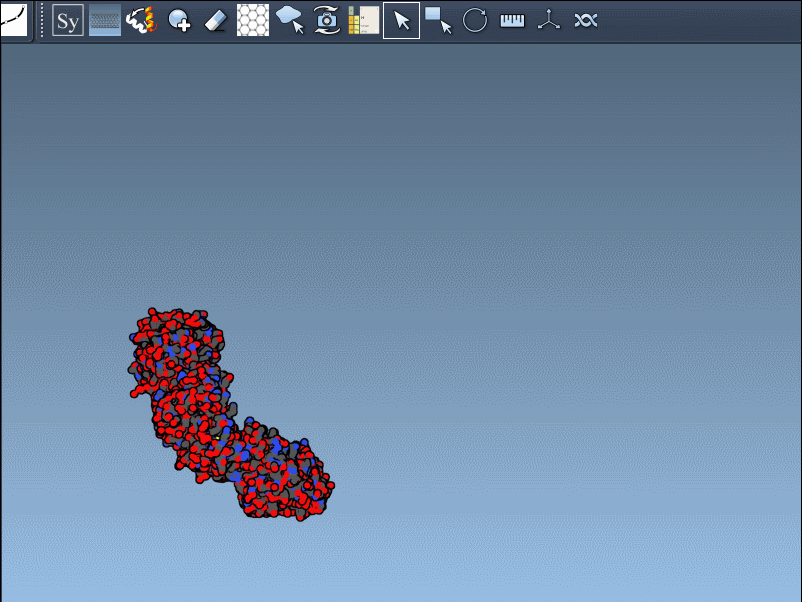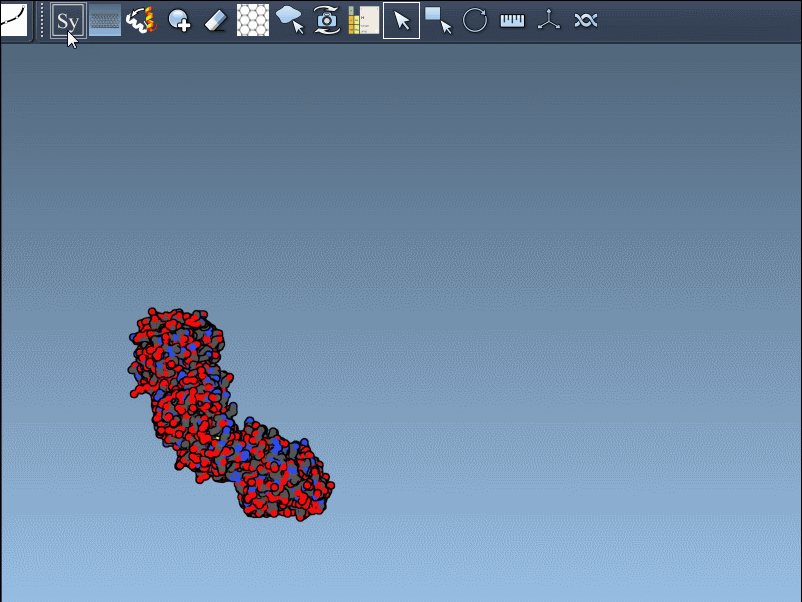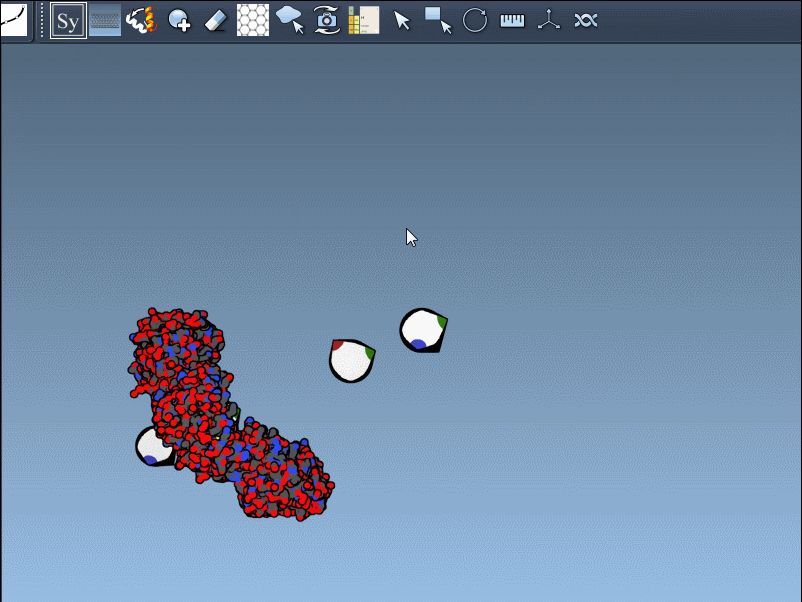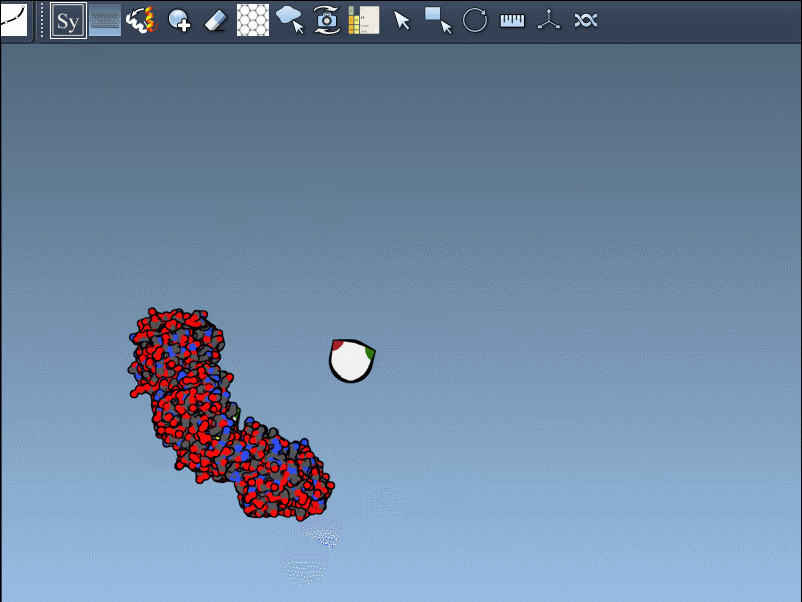
Clicking on the control node permanently generates the replica, creating visually accurate symmetric assemblies in just a few clicks.
CRYST1 vs. BIOMT records
The Symmetry Mate Editor supports two primary types of PDB annotations for symmetry data:
- CRYST1: Captures symmetry from the crystal lattice, useful for visualizing periodic packing.
- BIOMT: Defines symmetry from biological assembly annotations, instrumental in reconstructing functional complexes.
The editor makes it effortless to toggle between these modes. For instance, when using 1B5S, you can view both CRYST1 and BIOMT symmetries interactively:
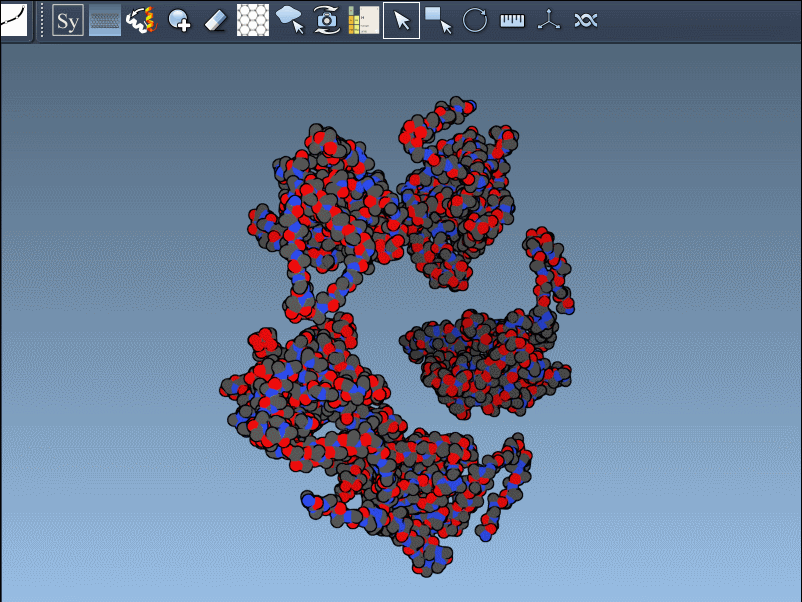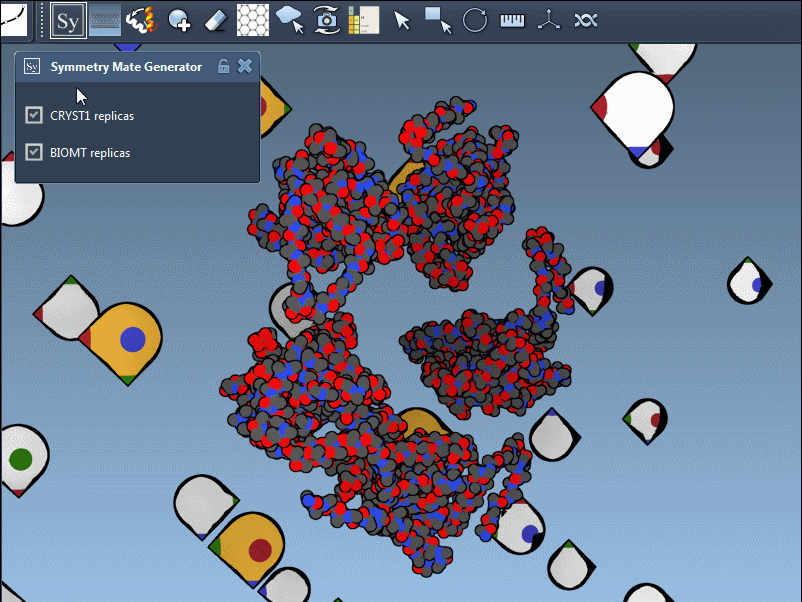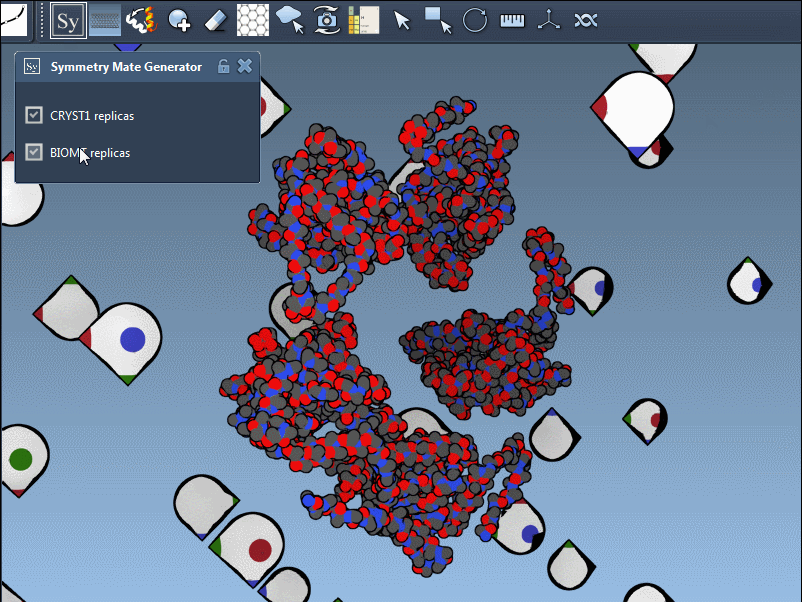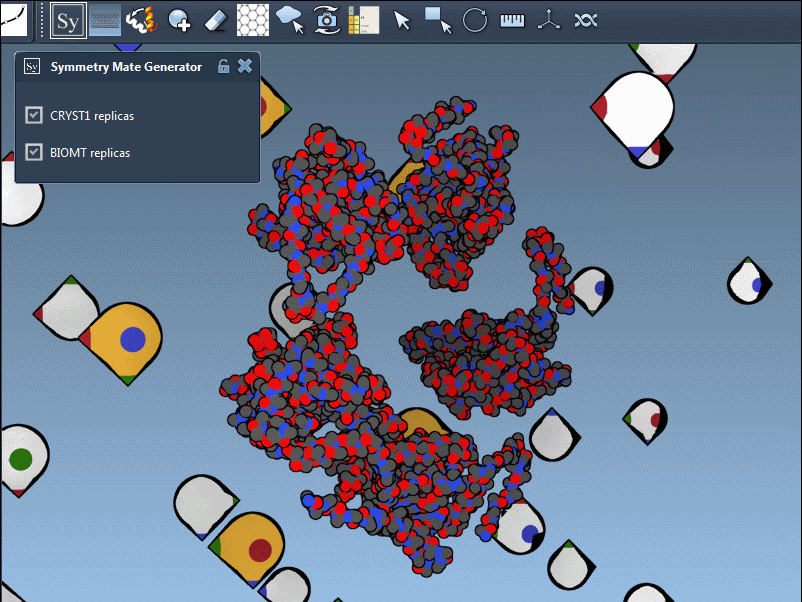
Pro tips for better results
- Use the Ribbons model to emphasize chain differences, and colorize visual data to improve clarity.
- Combine the Symmetry Mate Editor with Symmetry Detection for analyzing axes and RMSD.
- Make use of the Undo option (Ctrl/Cmd + Z) to revert unwanted replicas smoothly.
The Symmetry Mate Editor provides a powerful way to investigate and manage symmetric assemblies effortlessly. Whether you are studying protein-protein interactions or designing molecular scaffolds, this tool streamlines symmetry-based workflows precisely and interactively.
To dive into all the details, check out the full documentation page here.
Note: SAMSON and all SAMSON Extensions are free for non-commercial use. Get started with SAMSON at https://www.samson-connect.net.





