Designing new molecular analogues to study or optimize properties like binding affinity is a common challenge in molecular modeling. But what if there were a quick and efficient way to explore analogue possibilities directly on your platform without tedious manual edits? In this post, we’ll show you how you can leverage SAMSON’s SMILES Manager extension to perform Positional Analogue Scanning. This technique allows you to explore chemical modifications systematically and study their impact on molecular properties.
Start from a Single Molecule
The first step in this process is selecting or defining the molecule you want to modify. You can either enter the molecule’s SMILES code directly, or, if you have it already in your SAMSON workspace, just select it and hit the Use selection button. It’s that straightforward!
Here’s an example using SMILES code: CN1C=C(C(=O)Nc2ccc(-c3ccccc3)c(c2)C(F)(F)F)C(=O)c2ccccc12. This molecule will serve as the base for our analogue generation experiment:
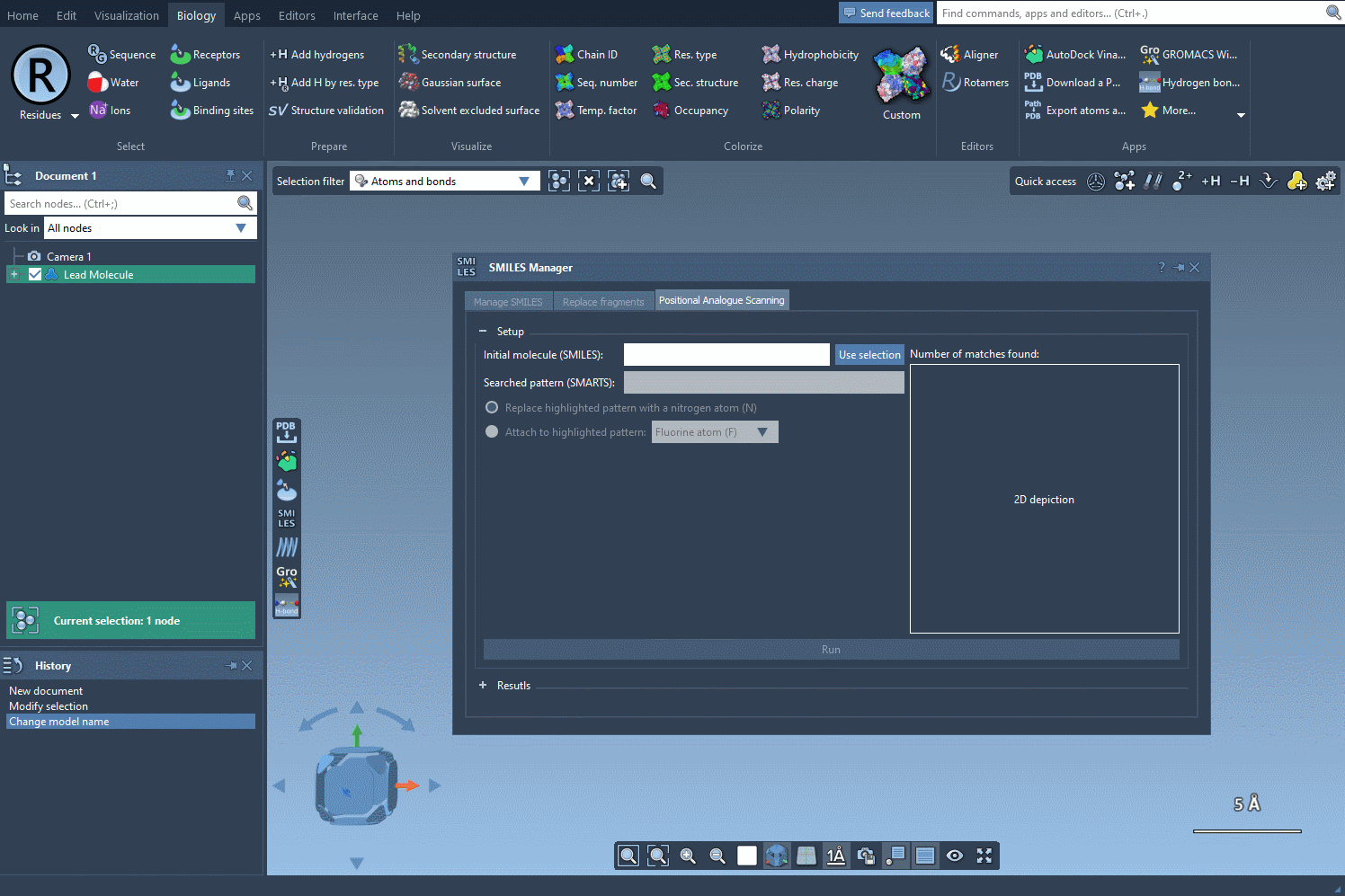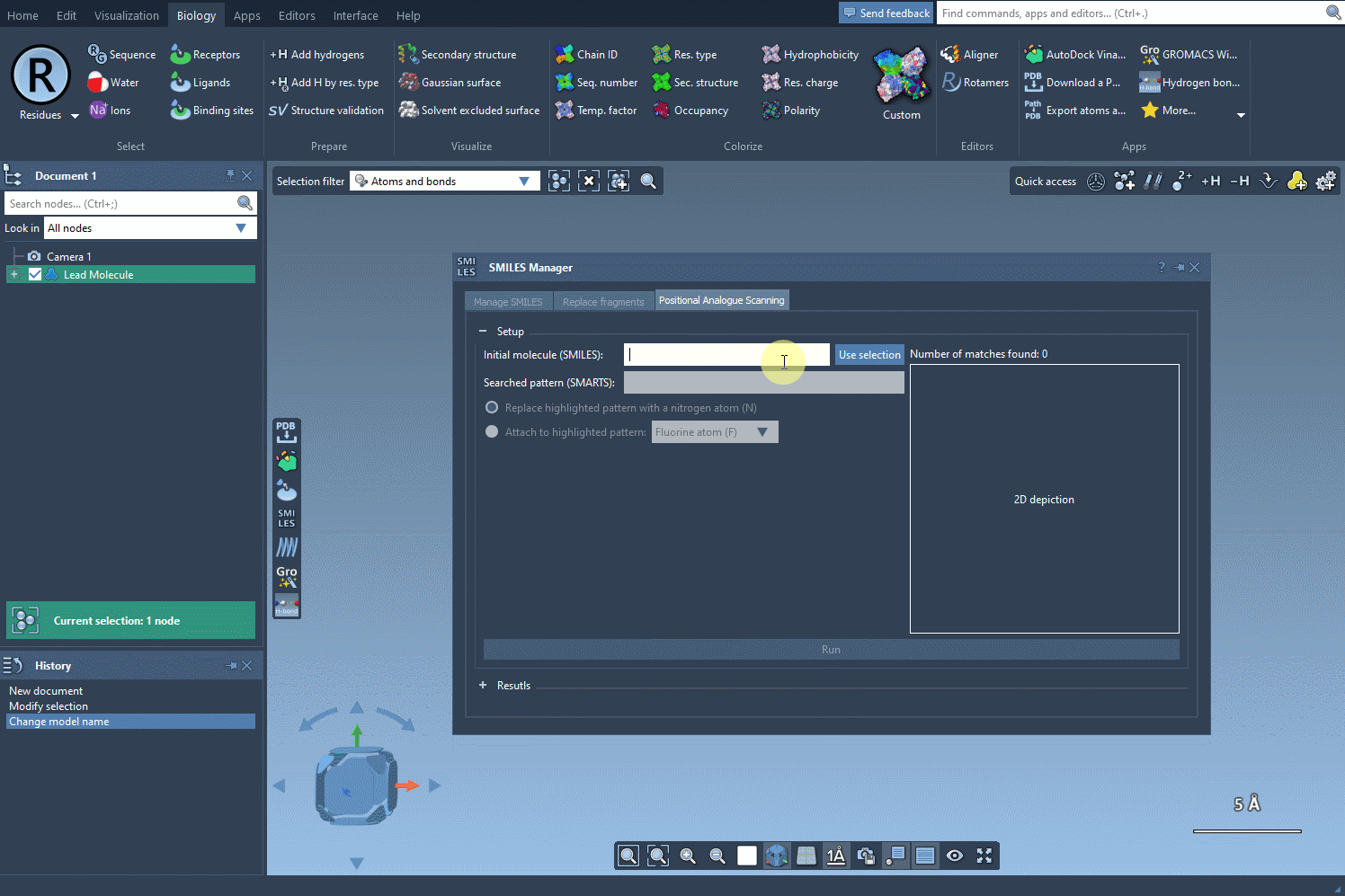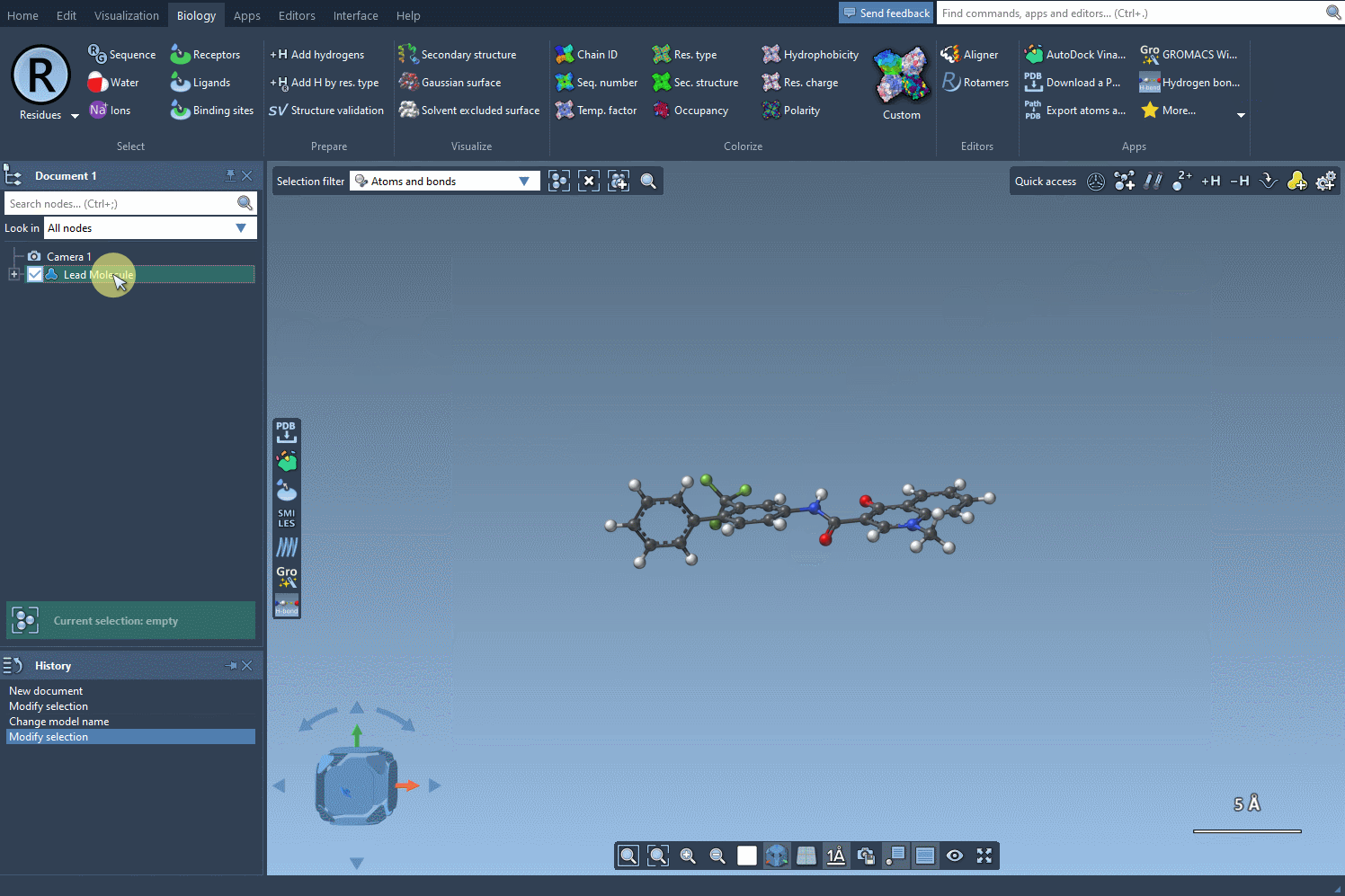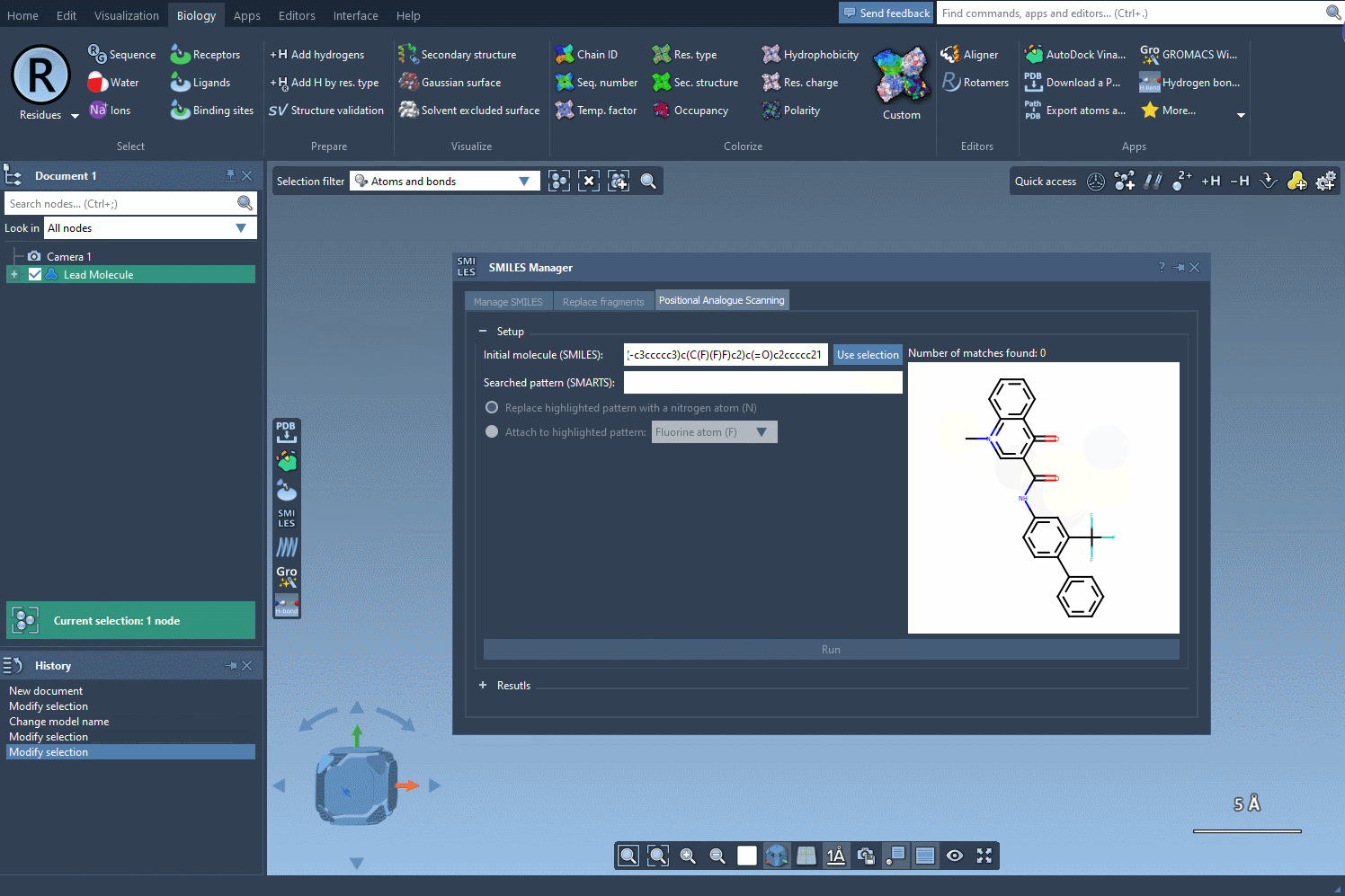
Targeted Pattern Replacement
The key to positional analogue scanning is identifying the pattern you want to replace or modify. In SAMSON, this involves defining your search pattern using SMARTS code. For example, if you’d like to target aromatic carbon in your molecule, you would use the SMARTS code [cH]. This highlights matches within your molecule, as shown in the following animation:
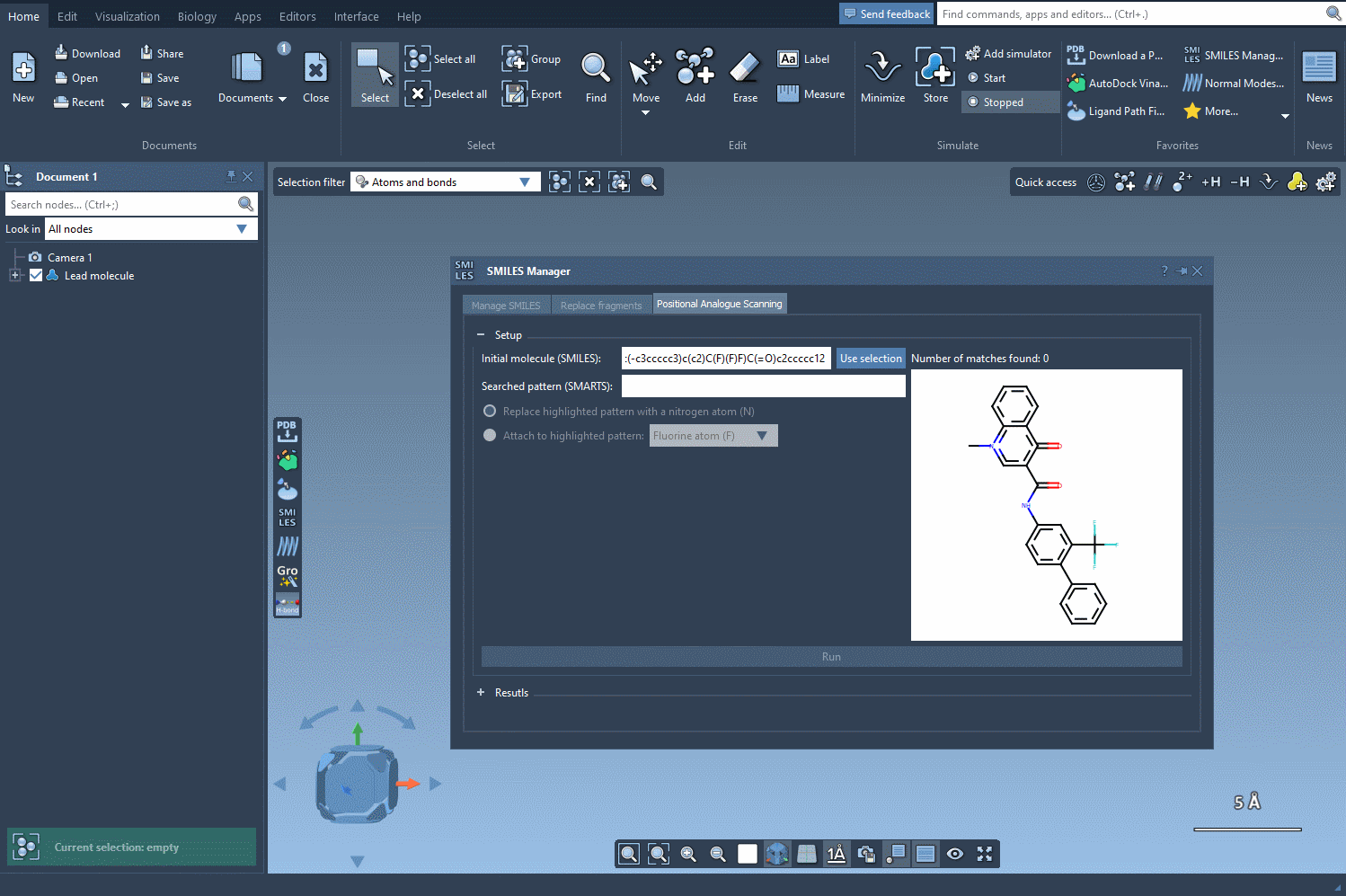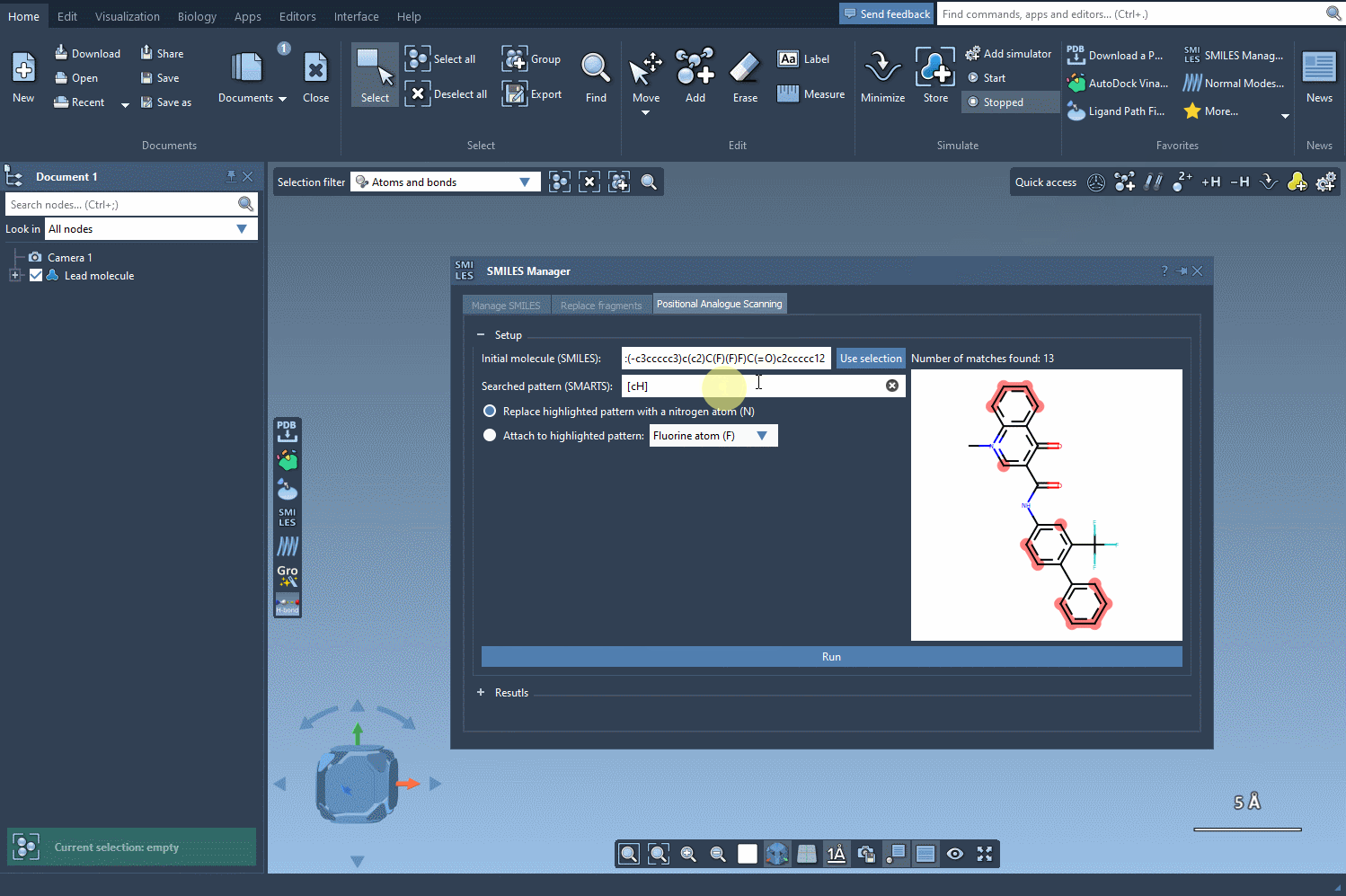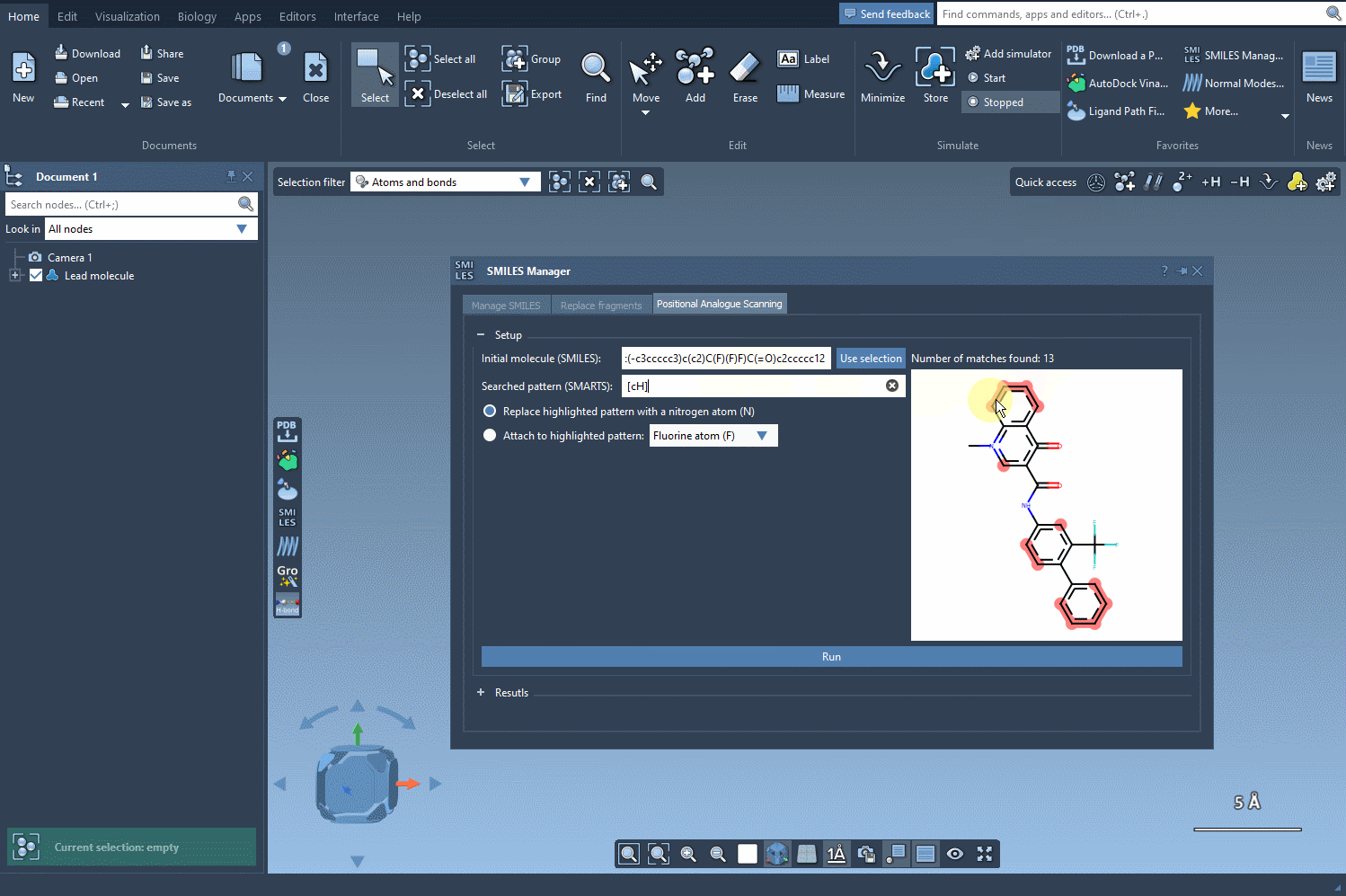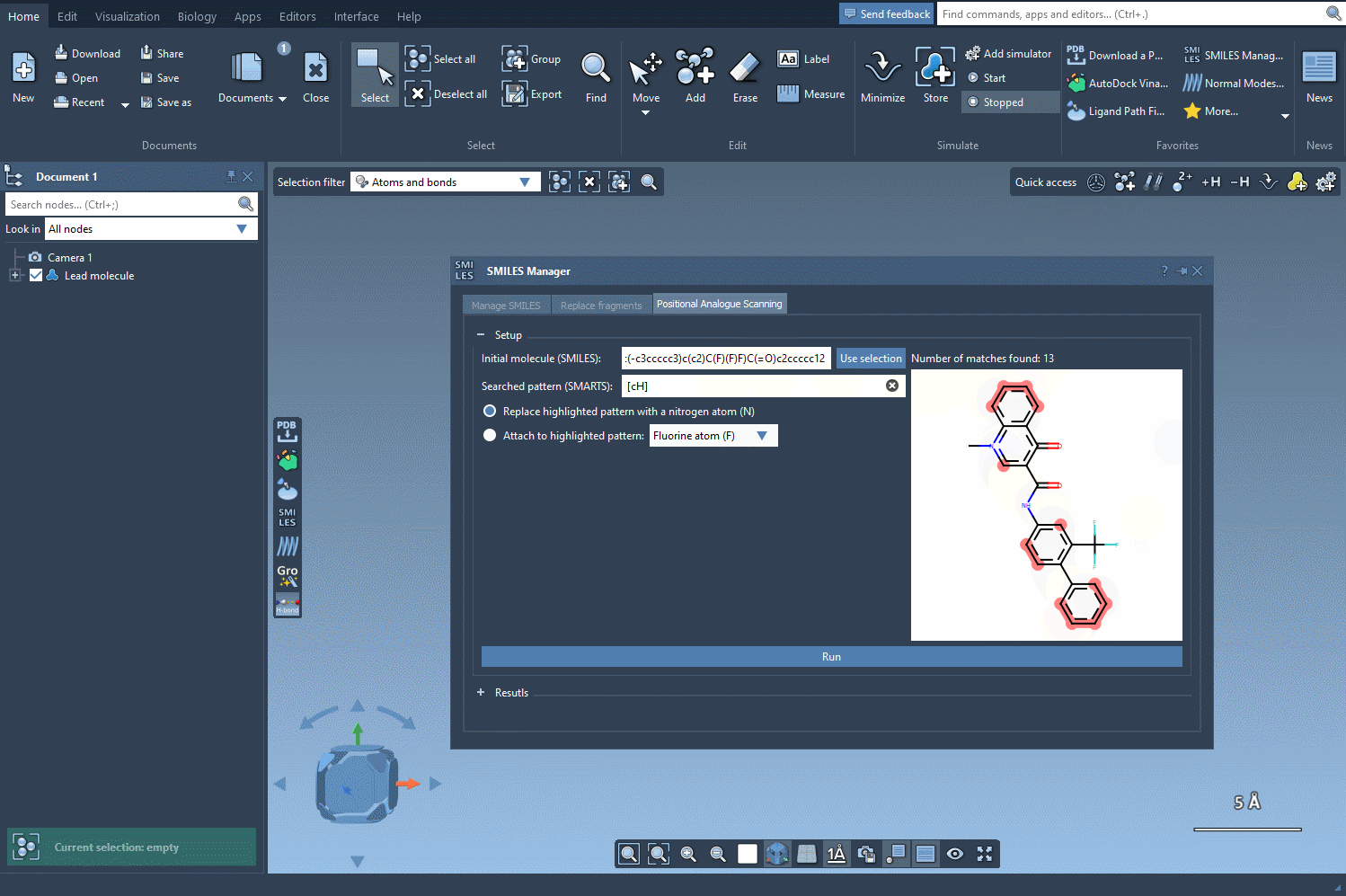
With the targeted areas highlighted, you can now choose whether to replace the pattern or attach something to it.
Easily Generate Analogues
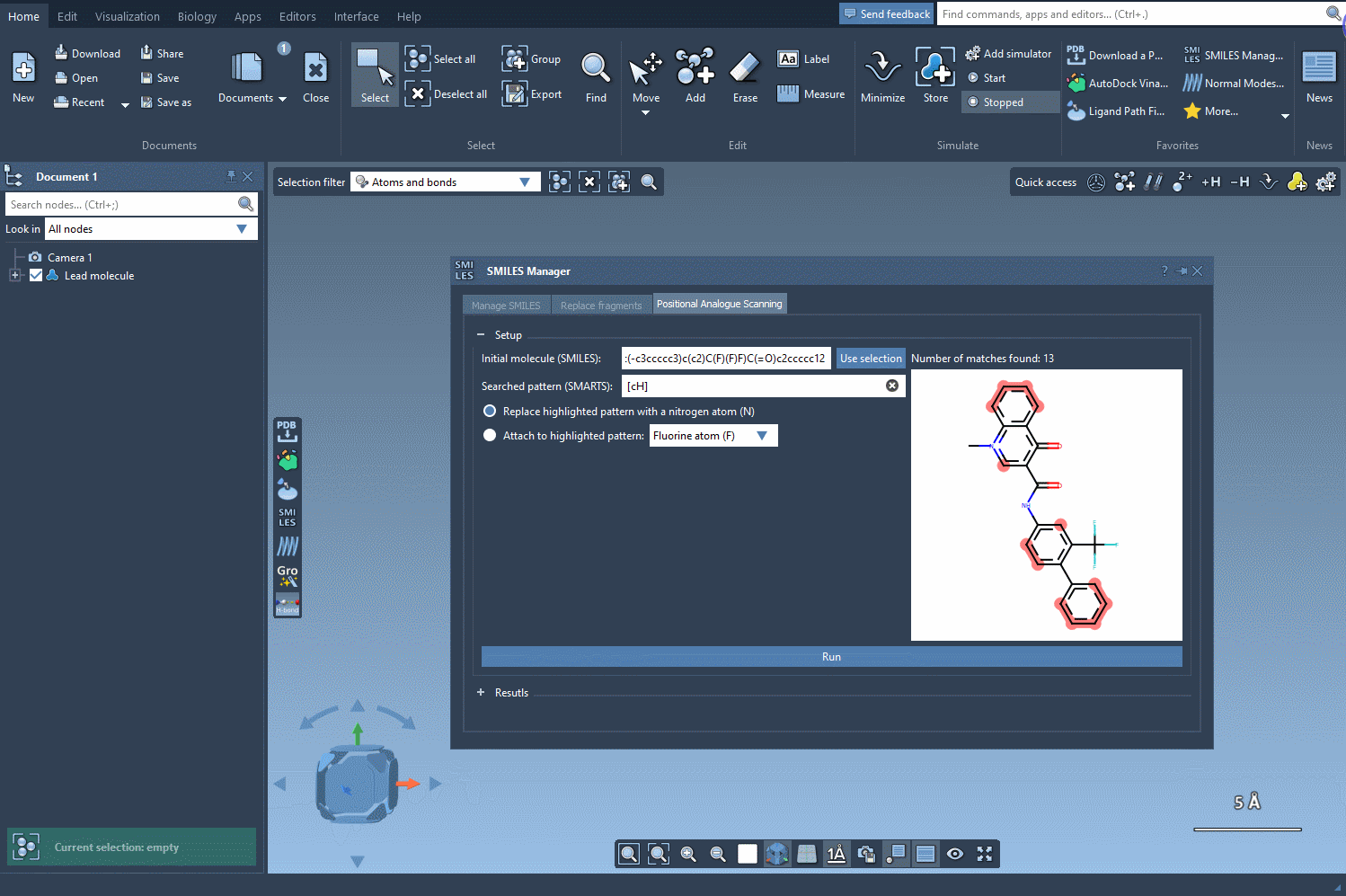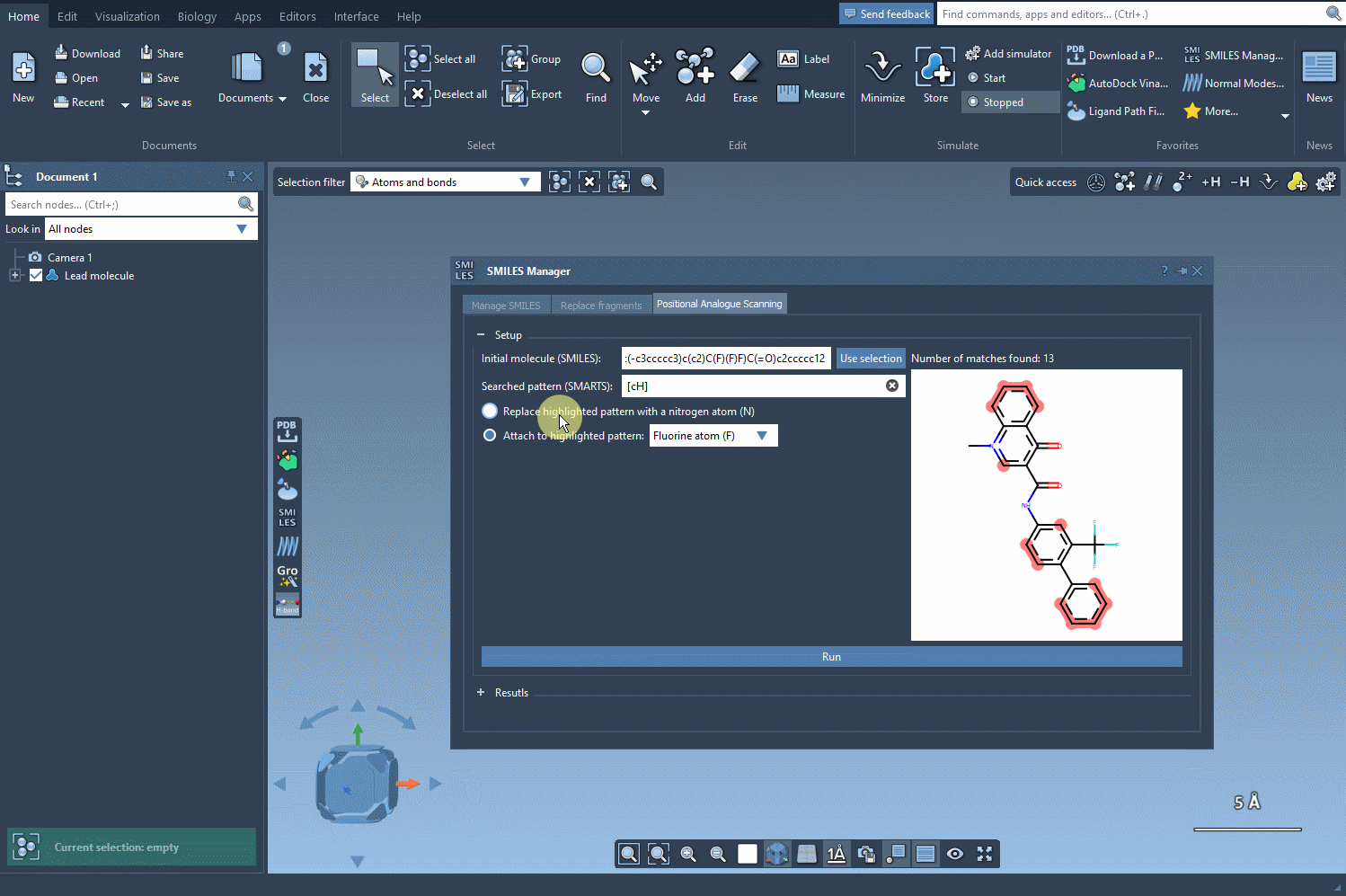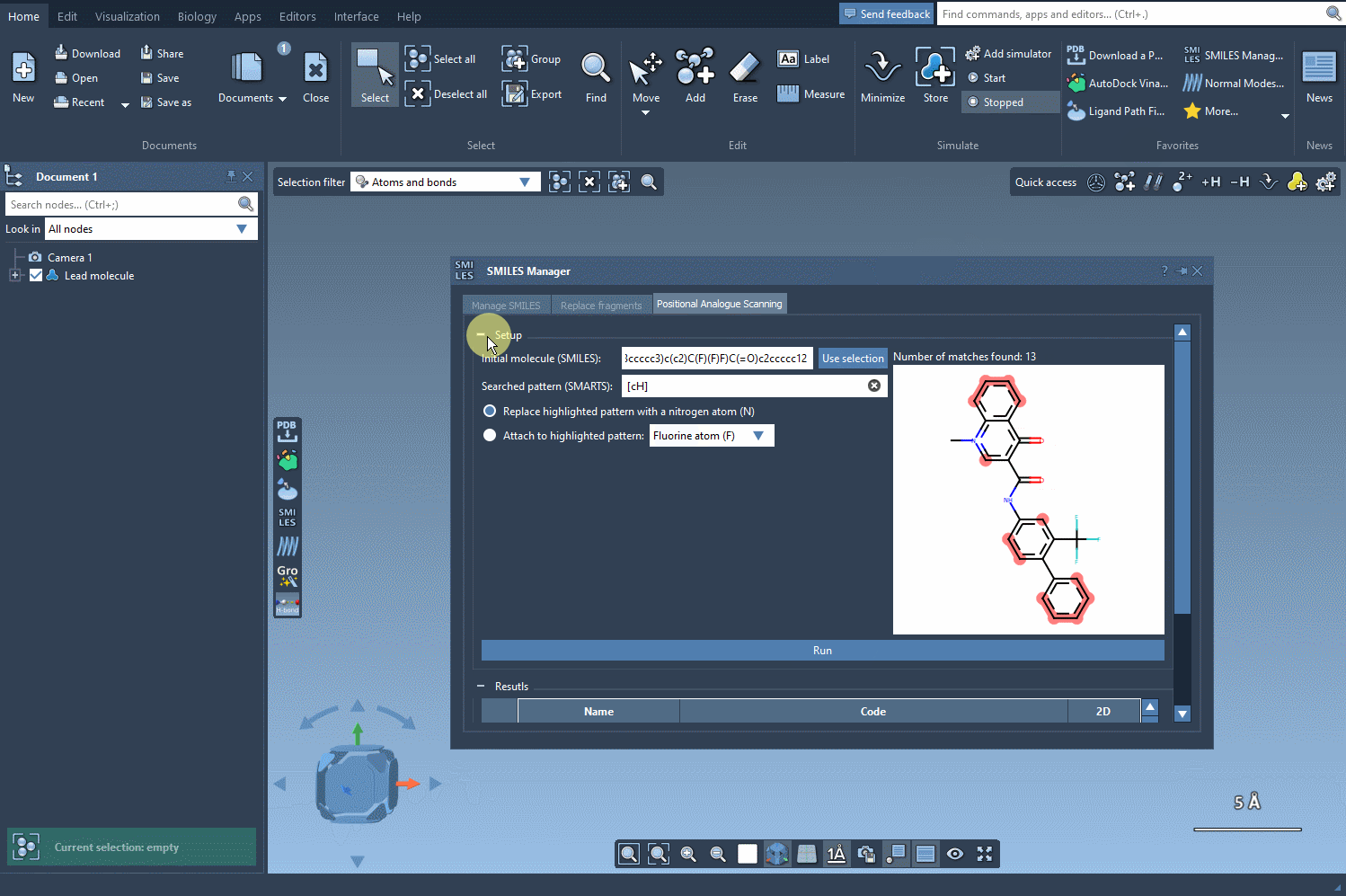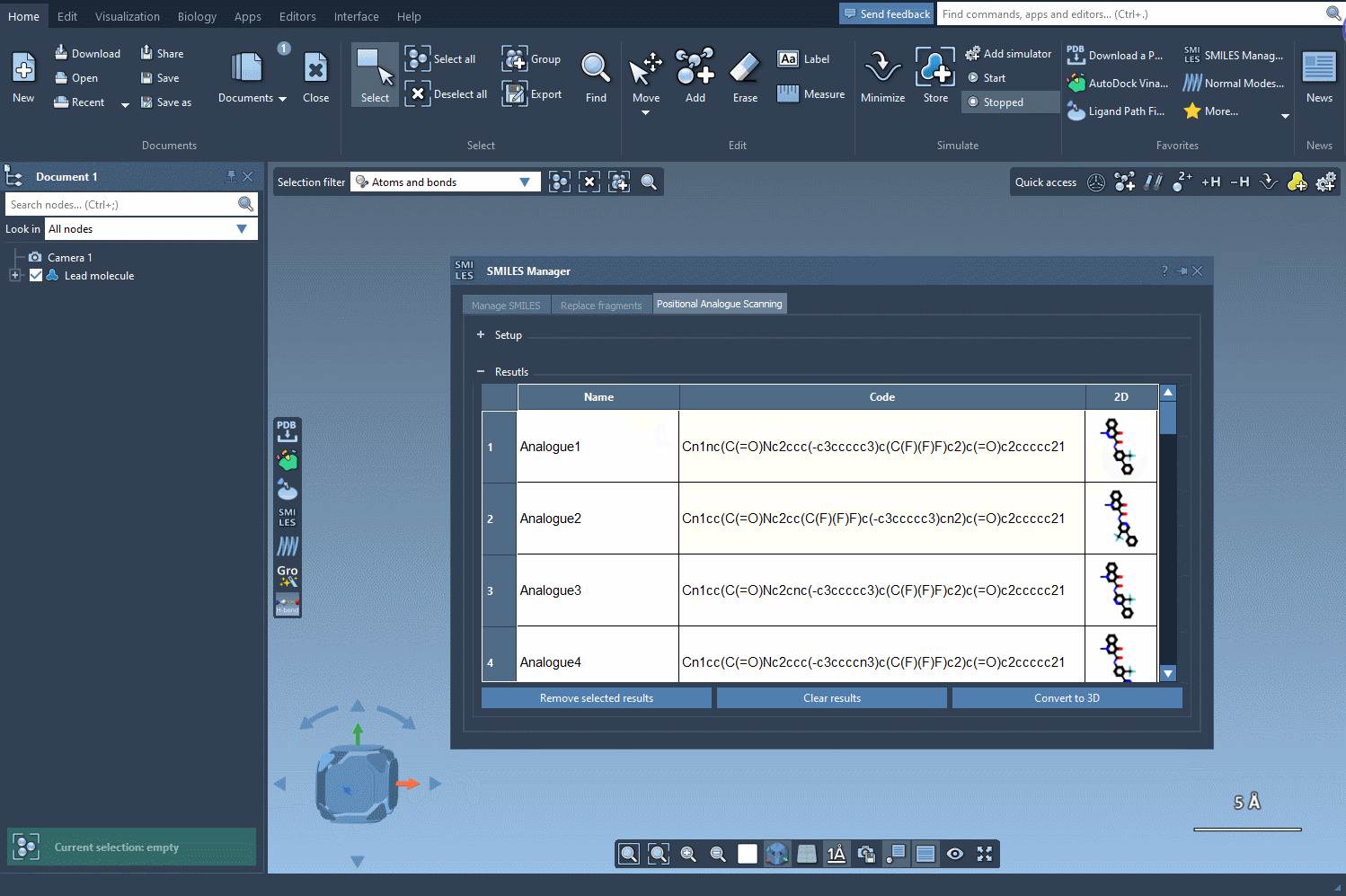
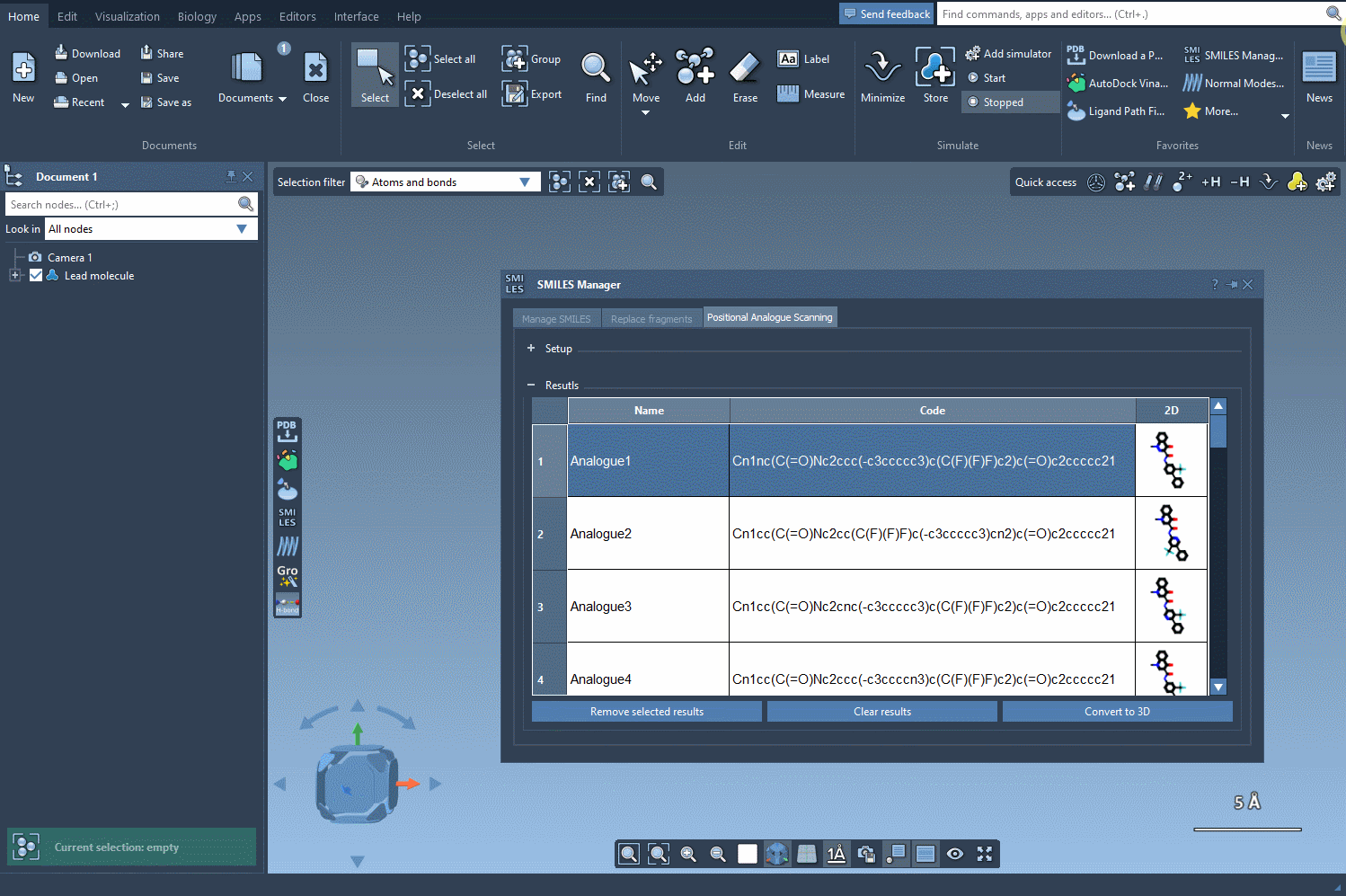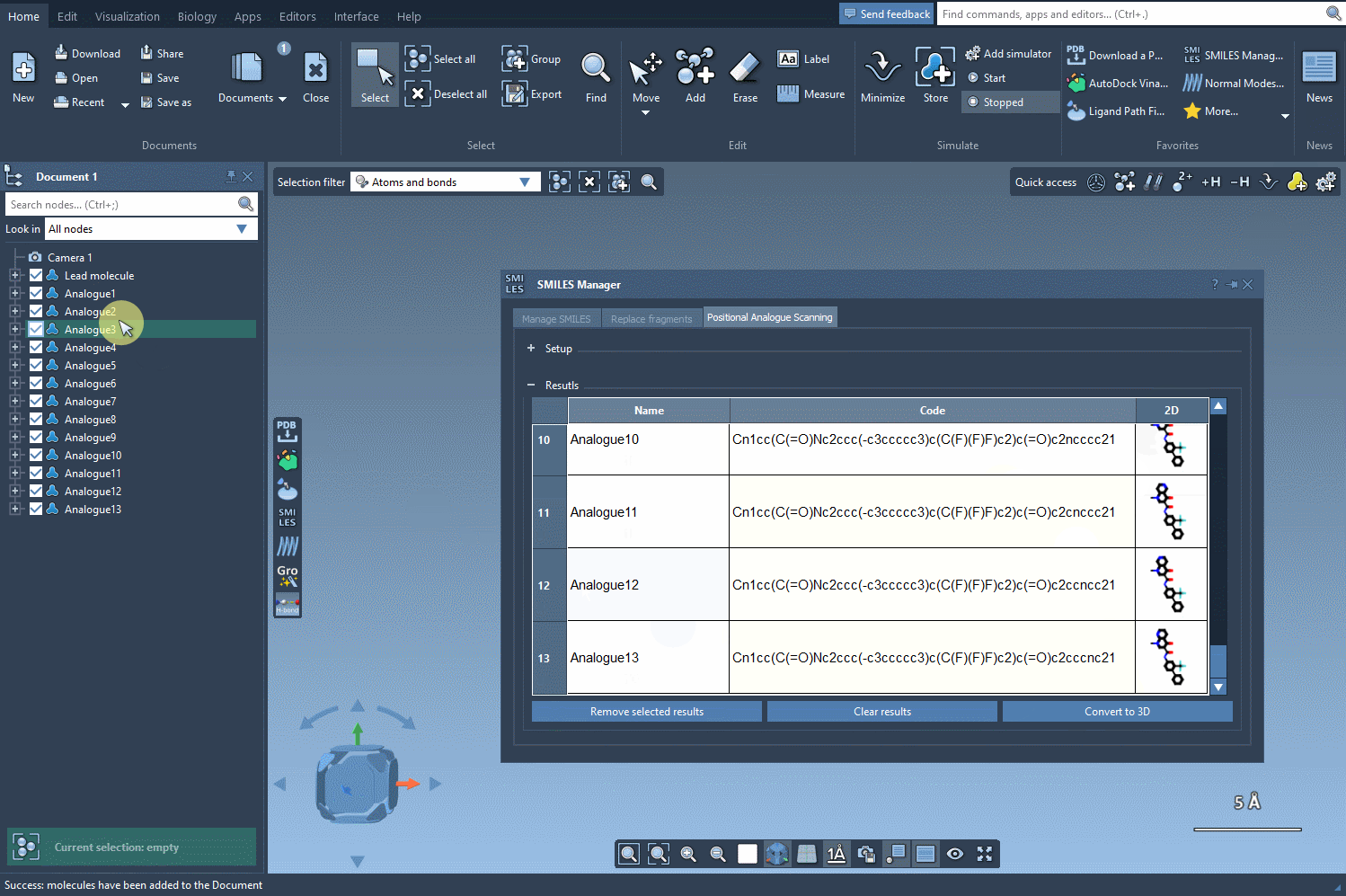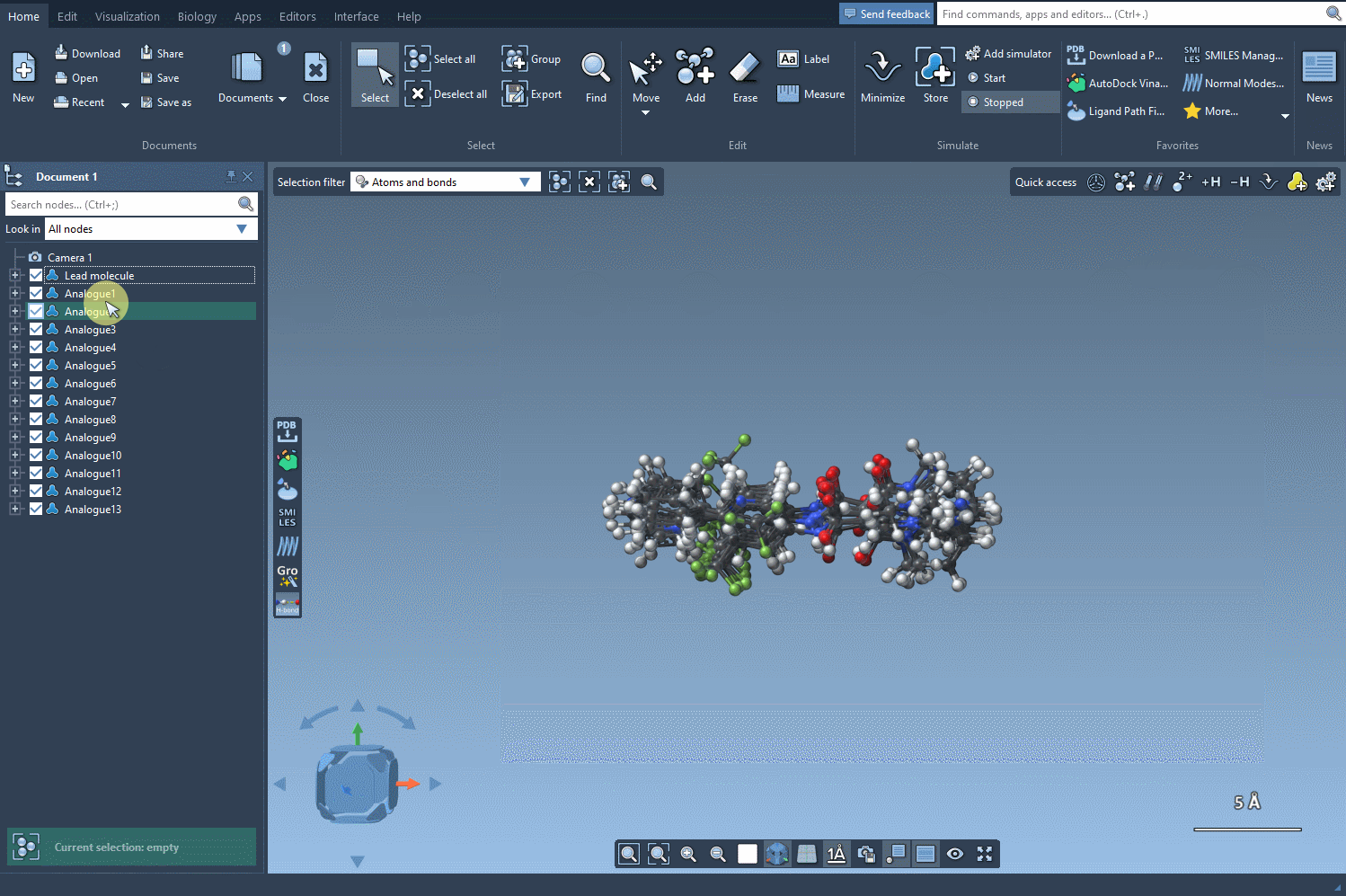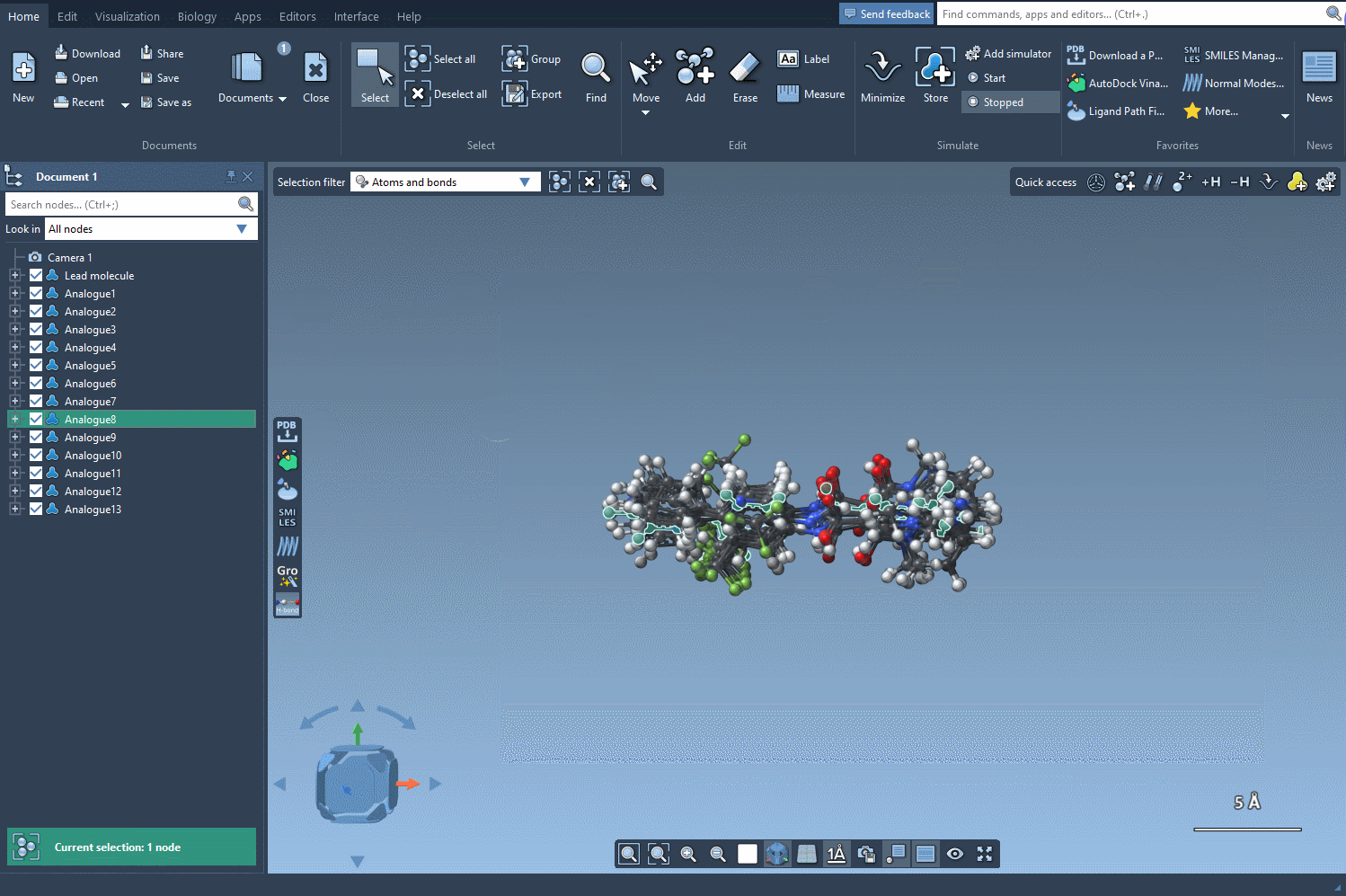
In our example, let’s replace the pattern with a nitrogen atom (N). Alternatively, you could attach a fluorine atom (F) or a methyl group (CH3). Once you’ve specified your choice, simply click the Run button to generate the analogues. SAMSON will quickly provide not only the updated SMILES codes but also 2D depictions of your analogues, making it easy to evaluate the results:
Modify and Fine-Tune Results
Your generated analogues will appear in a results table where you can perform additional modifications or analyses. Here’s what you can do within this table:
- Modify individual analogues by changing their names or SMILES codes directly in the table.
- View 2D depictions in more detail by double-clicking or opening them in a larger window.
- Toggle the visibility of analogue depictions in the results table for cleaner organization.
- Generate 3D structures for further study, such as docking or interaction analysis.
- Remove specific results or clear the entire table for a fresh start.
These tools are designed to give you flexibility and control over your analogue design process.
Bring It into 3D
If you’re ready to investigate your analogues in 3D, the Convert to 3D button makes it simple. The 3D structures of the analogues can then be utilized for more advanced studies, like docking experiments using SAMSON’s Autodock Vina Extended extension. This lets you see how changes in structure impact interactions with target proteins.
Whether you’re optimizing binding affinity, studying molecular properties, or exploring chemical possibilities, positional analogue scanning in SAMSON provides a streamlined and practical solution. Want to learn more? Head over to the official documentation page: here.
Note: SAMSON and all SAMSON Extensions are free for non-commercial use. You can get SAMSON today at SAMSON Connect.





