For molecular modelers, a common task is transforming SMILES (Simplified Molecular Input Line Entry System) strings into 3D molecular structures for simulations and analysis. However, doing this manually or using cumbersome workflows can become a time-intensive challenge, especially for large-scale projects. Enter the SMILES Manager Extension in SAMSON—a powerful tool utilizing RDKit to simplify and accelerate the conversion process.
The Challenge: From SMILES to 3D
SMILES strings are a universal representation of molecules, but they are often only the start of a broader modeling or simulation workflow. Converting these representations into 3D structures is key for deeper analyses, such as docking, dynamics simulations, and property predictions. The manual process of transforming SMILES strings into 3D structures can be prone to errors, especially for complex molecules, and can require multiple tools. The SMILES Manager in SAMSON addresses these challenges by streamlining the transformation process directly within its interface.
How to Seamlessly Generate 3D Structures
Using the SMILES Manager, generating high-quality 3D molecular structures is straightforward. Here’s how:
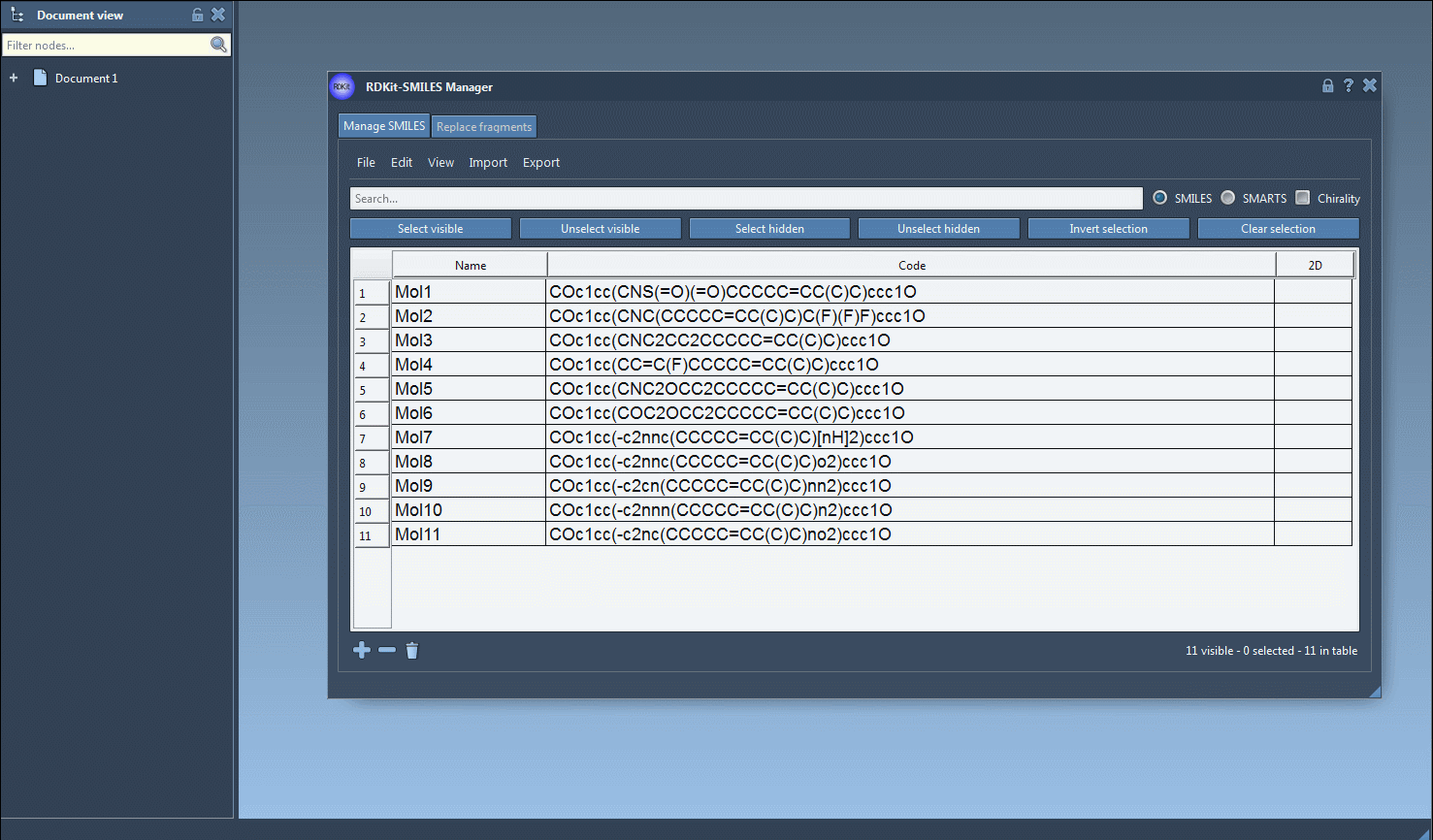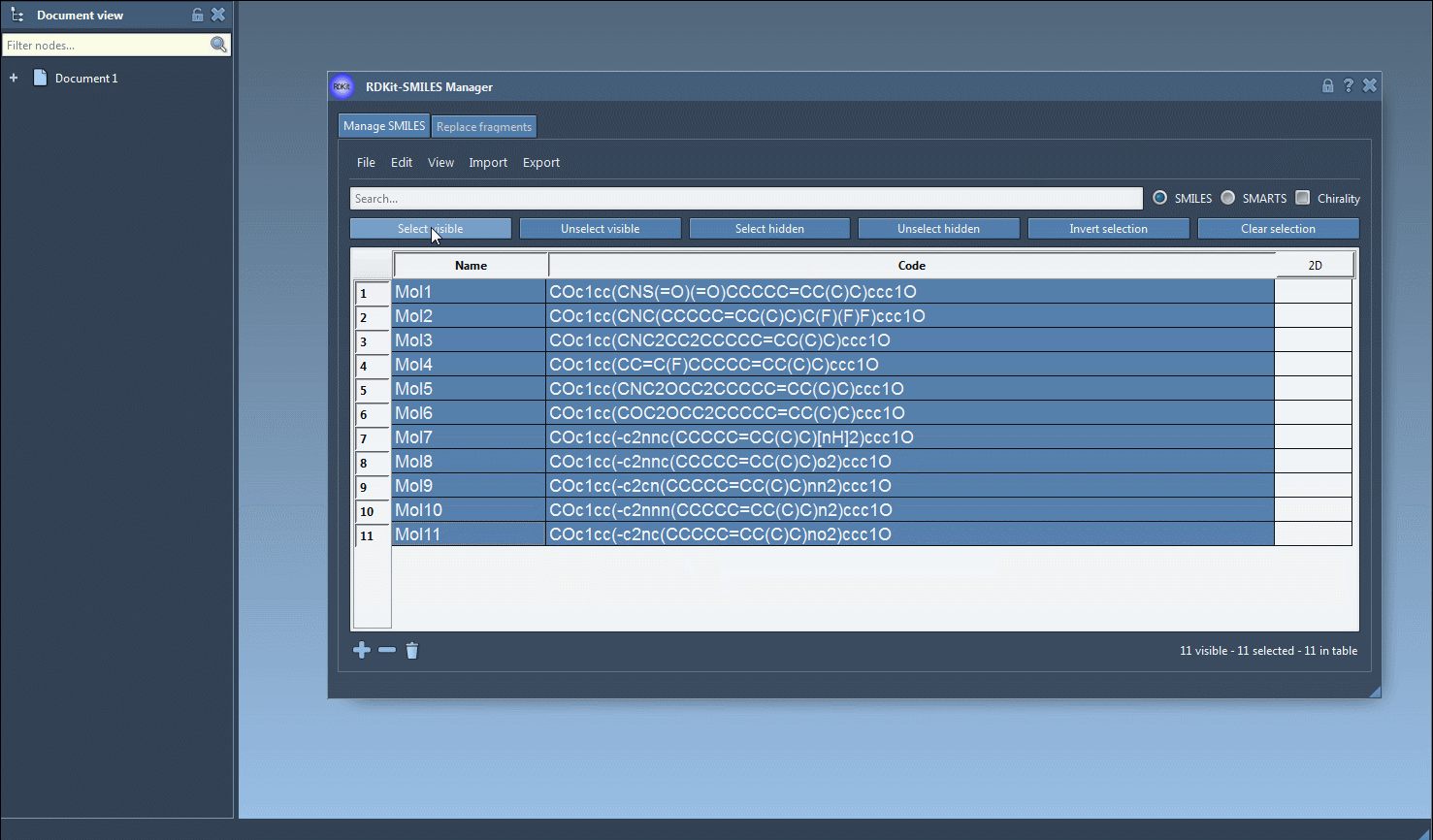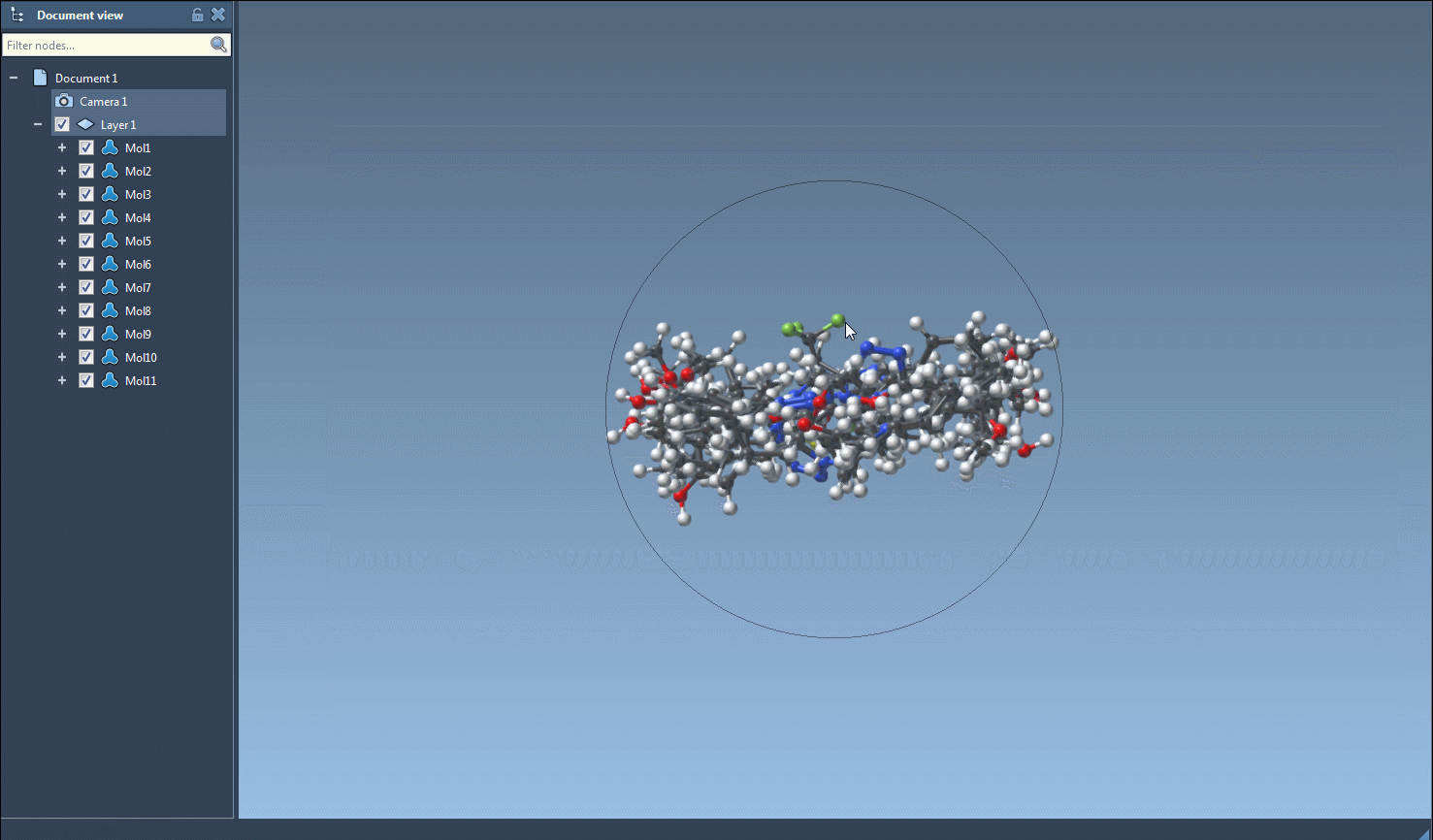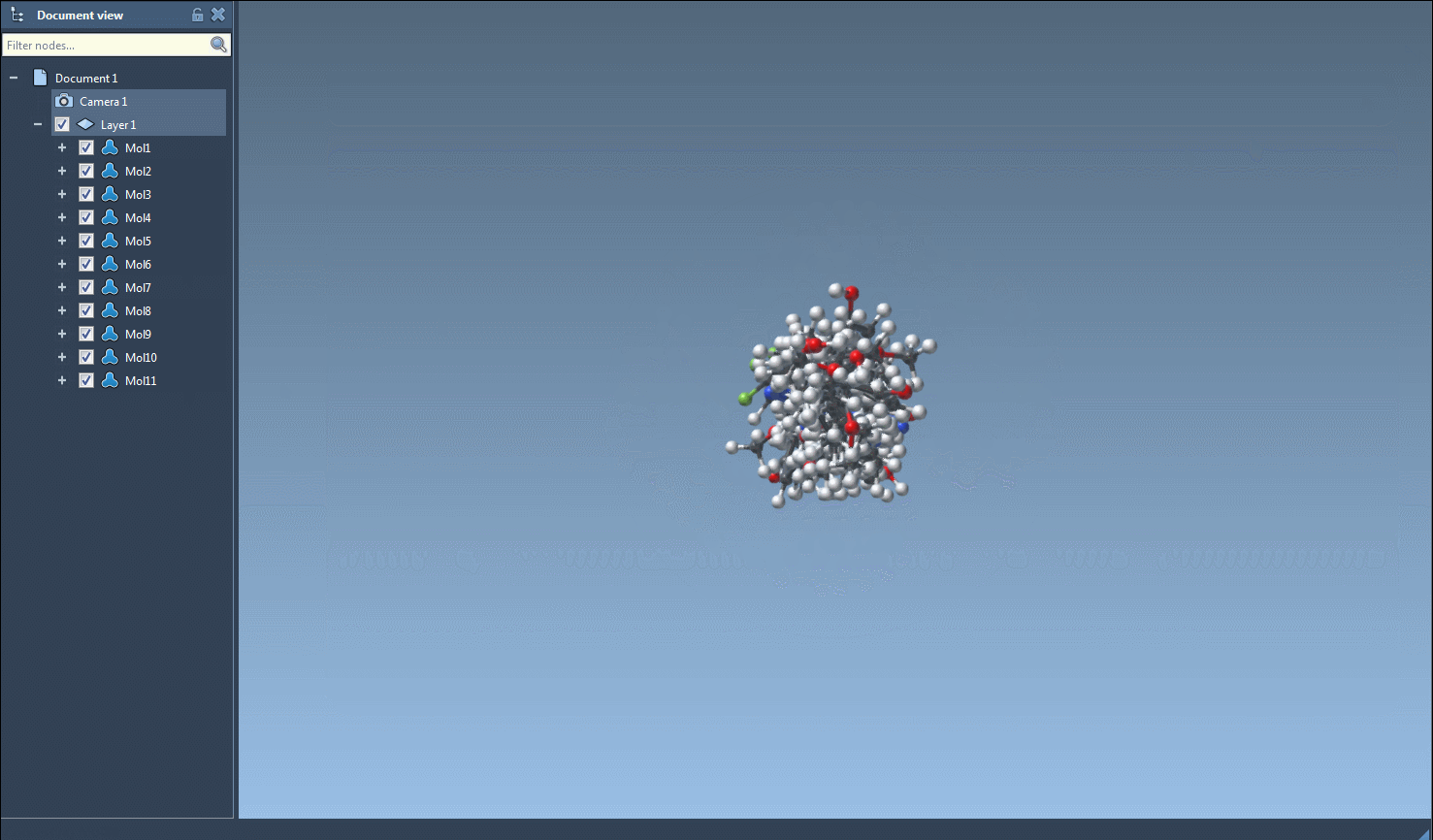
- Import Your SMILES Data: Import your SMILES strings into the SMILES Manager using .smi or .txt files. Just go to the File menu and click Open. The tool supports standard RDKit formats, making it compatible with widely-used datasets.
- Select Molecules in the Table: Once imported, your SMILES strings along with their 2D conformations will appear in a table. For 3D conversion, simply select the molecules you’re interested in.
- Initiate 3D Structure Generation: From the Export menu, select the Selected SMILES string to Document option. This triggers the conversion algorithm, generating 3D structures that are added to your active SAMSON document with corresponding molecule names.
- Single-Molecule Conversion: Prefer to focus on one molecule? You can right-click a SMILES entry in the table and use Generate 3D structure, or open its large view window and use the dedicated Generate 3D structure button.
Additionally, the SMILES Manager handles invalid SMILES strings gracefully, highlighting errors and providing feedback to maintain a smooth workflow.
Why This Matters
The 3D generation process is fully integrated into the SAMSON environment, minimizing the need for external tools and data shuffling. This makes it easier to batch-process large datasets or focus on specific molecules without workflow interruptions.
Moreover, SAMSON offers additional functionality, such as visualizing molecules, exporting 3D structures, and even enabling further structural manipulation. Combined with RDKit’s robust algorithms, this ensures accurate and efficient 3D conformer generation.
Next Steps
If you are a molecular modeler looking for an easy and reliable way to generate 3D structures from SMILES, the SMILES Manager Extension for SAMSON could significantly enhance your workflows. It’s a streamlined solution designed for professionals managing molecular data at scale.
Learn more about these features on the official documentation page at https://documentation.samson-connect.net/tutorials/smiles-manager/using-the-rdkit-smiles-manager/.
Note: SAMSON and all SAMSON Extensions are free for non-commercial use. You can download SAMSON at https://www.samson-connect.net.





