As a molecular modeler, you might often face challenges while dynamically editing molecular systems. Standard molecular simulation tools aren’t always equipped to handle seamless topology changes like breaking or forming covalent bonds during interactive modeling. This is where the Interactive Modeling Universal Force Field (IM-UFF), an extension of the Universal Force Field (UFF), comes into play. In this post, we’ll explore how IM-UFF can empower you to model complex molecular structures with unprecedented flexibility.
The IM-UFF interaction model, introduced by Jaillet, Artemova, and Redon, extends UFF by providing interactive capabilities to handle topological changes smoothly. Whether you’re editing molecular structures or simulating complex transformations, IM-UFF allows significant modifications to molecular systems in real-time, guided by physically-based inter-atomic forces.
Why Use IM-UFF for Interactive Modeling?
The key strength of IM-UFF is its ability to adapt topology dynamically. Here’s what it brings to the table:
- Covalent Bond Manipulation: IM-UFF enables you to create and break covalent bonds or modify bond orders during modeling sessions.
- Dynamic Typization: The atom types adjust dynamically based on their positions, ensuring smooth topological transitions.
- Ease of System Modification: Combined with an intuitive user interface, IM-UFF provides the freedom to add or delete atoms on-the-fly while simulating, automatically updating the molecular structure.
Getting Started with IM-UFF
To simulate a molecular system with IM-UFF, follow these steps:
- Open a molecular system in SAMSON.
- Add the IM-UFF simulator by navigating to Edit > Simulate > Add simulator or using the shortcut
Ctrl + Shift + M / Cmd + Shift + M. Select Interactive Modeling Universal Force Field in the list of models. - Choose a state updater for your simulation, such as FIRE (Fast Inertial Relaxation Engine), and confirm your selection.
IM-UFF does not require a separate setup window, distinguishing it from the standard UFF. Instead, its parameter window offers additional Interactive modeling options, such as:
- Static topology (UFF only): Toggle between standard and interactive modes. Note that IM-UFF adjusts the zero-energy reference to align with non-interacting atom states. For detailed energy considerations, refer to the original reference.
- Keep vdW for manipulated: When unchecked, atoms manipulated with the mouse are excluded from van der Waals (vdW) calculations, making large structural modifications easier to orchestrate.
Once selected, you can initiate the simulation from Edit > Simulate > Start, and begin interacting with your molecular system. You’ll notice that small atomic displacements prompt local adjustments while maintaining the topology. Conversely, significant displacements either break bonds or form new ones, updating the topology dynamically.
Visualizing the Impact
The ability to seamlessly transform topologies can accelerate your workflow while eliminating common frustrations. For instance, IM-UFF simulations allow you to:
- Build complex molecular assemblies from simpler modules interactively.
- Easily simulate reactions involving bond formation and breaking.
- Experiment with manipulated atom positions without affecting vdW forces, in cases where connections may be otherwise hindered.
To see the process in action, refer to the image below from the IM-UFF documentation:
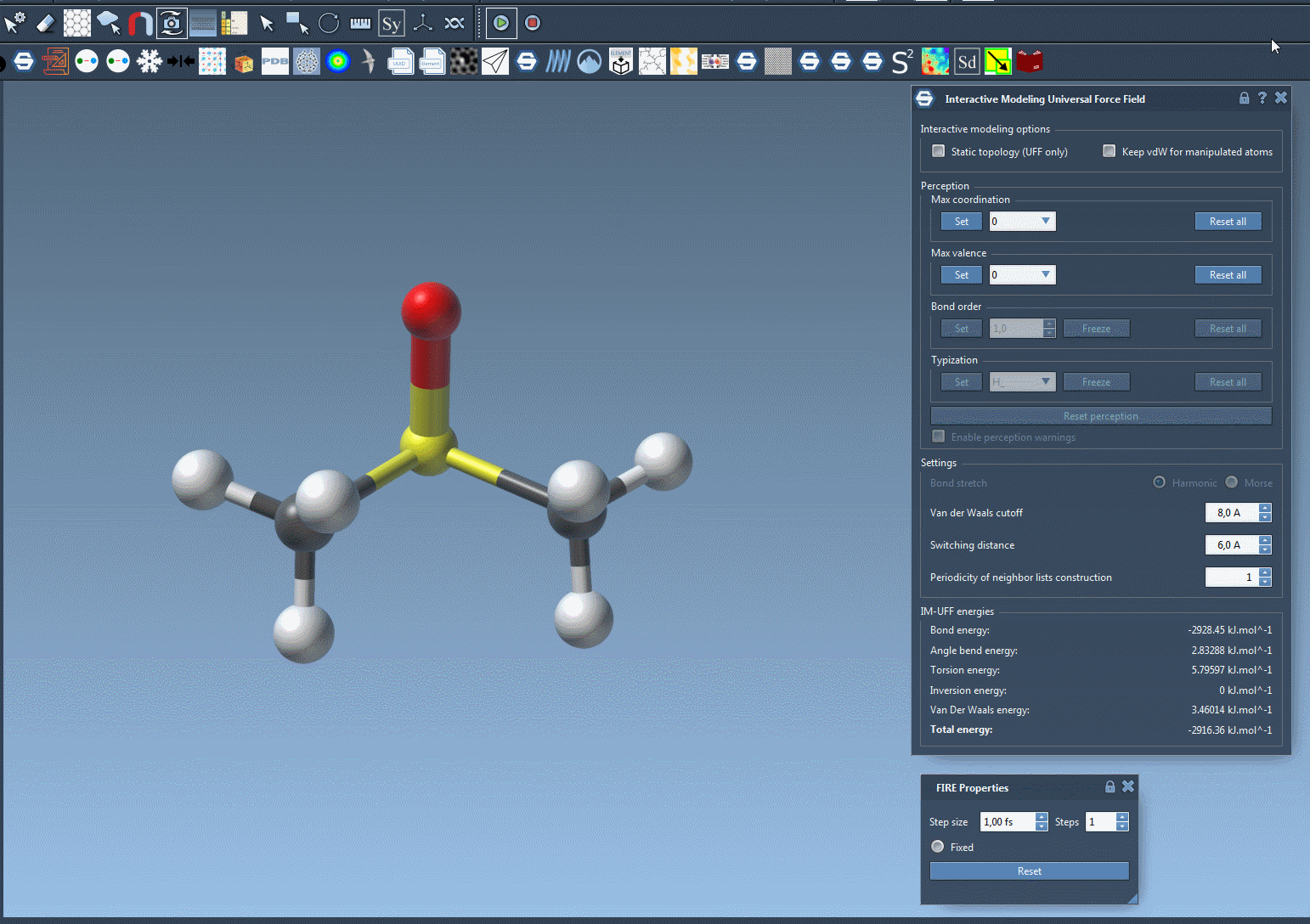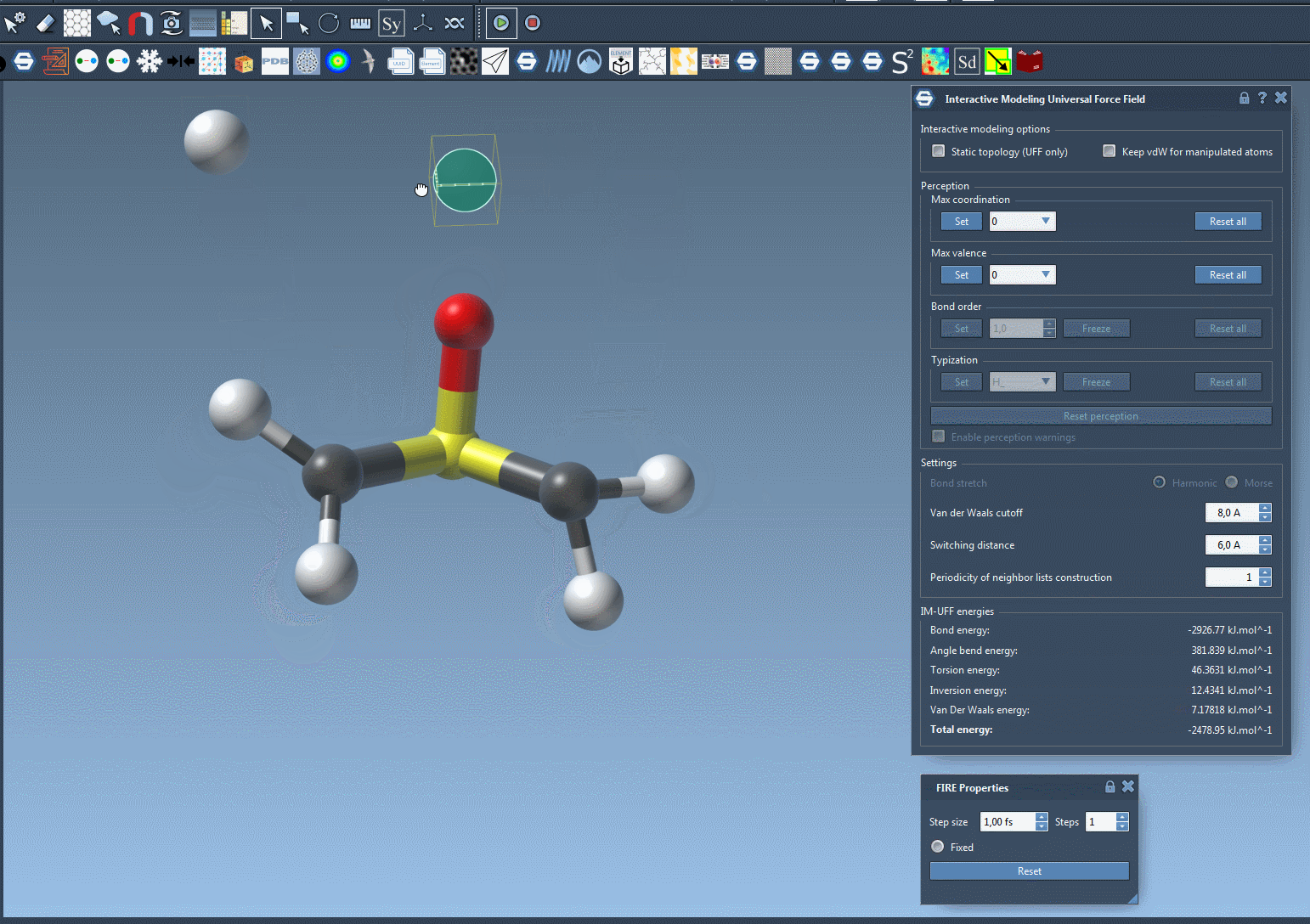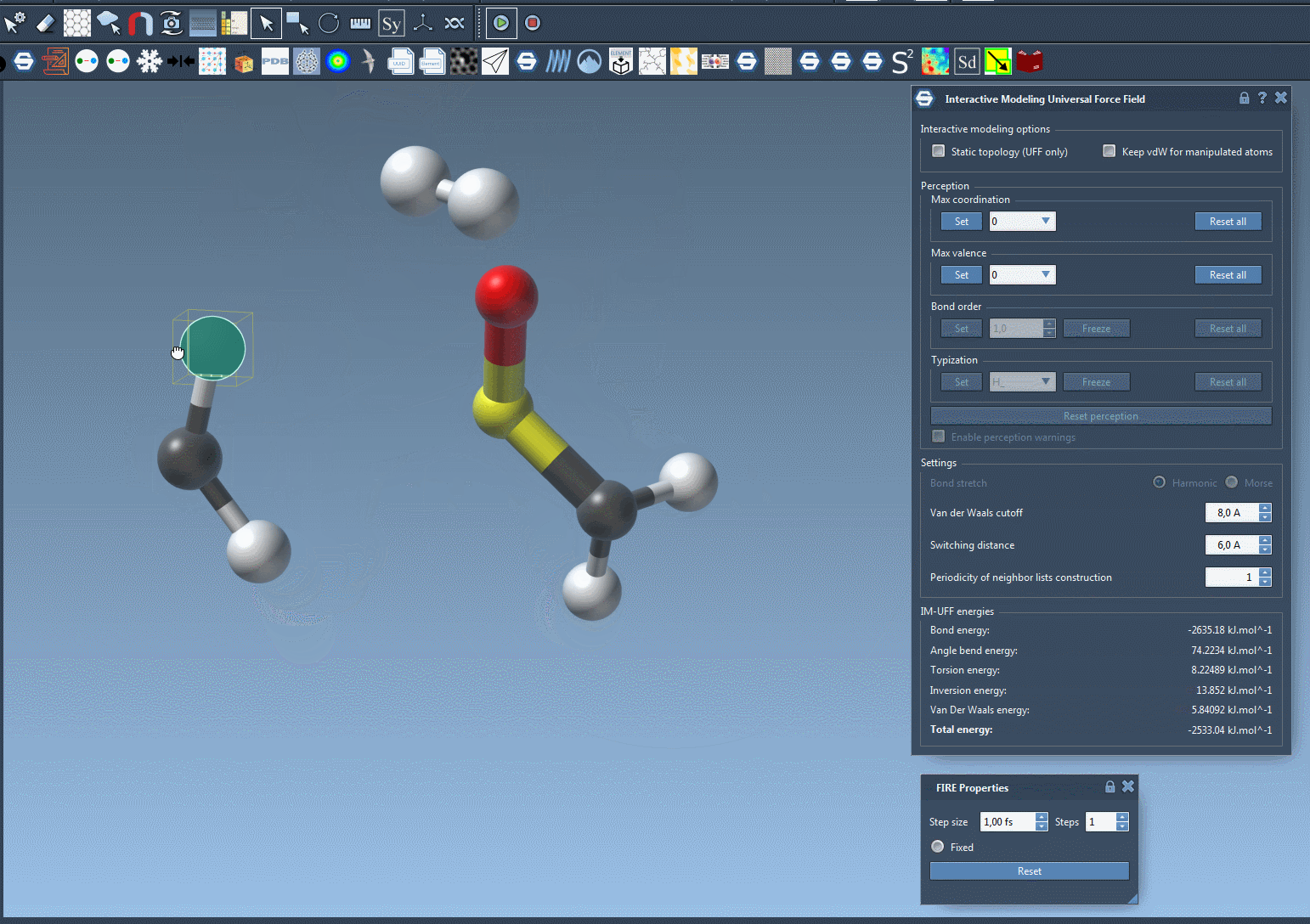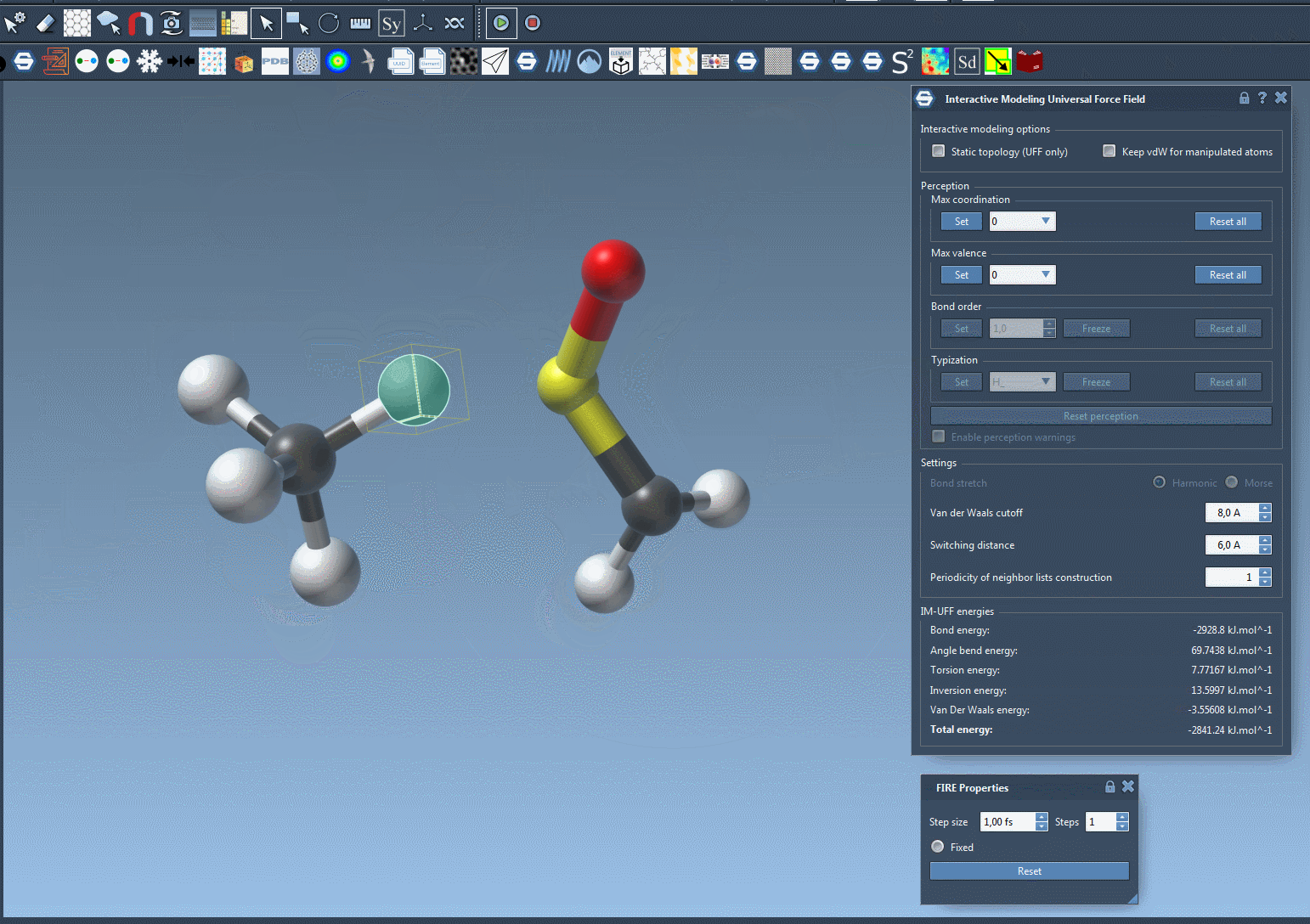
Customizing IM-UFF Simulations
IM-UFF also supports parameter customization depending on your needs. You can adjust:
- van der Waals cutoff and switching distances.
- Automatic typization parameters (e.g., maximum coordination or valence).
Keep in mind that with active IM-UFF simulations (static topology unchecked), bond order transitions become continuous functions of atomic positions for smooth topology updates. For an in-depth guide, explore the full documentation.
Interactive modeling with tools like IM-UFF unlocks extensive possibilities in molecular design, whether you’re conducting research on chemical reactions or refining biological systems. With its robust and user-friendly approach, IM-UFF proves to be an indispensable tool for the molecular modeling community.
Learn more about how to get started and access additional details in the full documentation.
Note: SAMSON and all SAMSON Extensions are free for non-commercial use. You can download SAMSON at https://www.samson-connect.net.





