For molecular modelers, one common challenge is accurately analyzing ligand pathways or reaction coordinates while preparing data for applications like free energy calculations or visualization. The Export Along Paths extension in SAMSON offers a powerful solution: effortlessly exporting atomic trajectories along predefined paths.
Why Export Trajectories?
When studying ligand pathways, you often need the exact coordinates of specific atoms along a path for deeper analysis or computational tasks. For example, exporting a ligand’s step-wise exit trajectory may help you perform free energy calculations or visualize intermediate states. The capability to extract these data points efficiently can dramatically improve workflow efficiency and accuracy.
Step-by-Step Guide: Exporting Atom Trajectories
The process of exporting atomic trajectories is streamlined in SAMSON:
Step 1: Load a Sample System
Before utilizing the Export Along Paths feature, it helps to have a sample system to work with. Start by loading the provided Lactose permease (1PV7) model with its ligand, Thiodigalactosid (TDG), and unbinding paths generated through the Ligand Path Finder. To load the system:
- Visit Home > Download in SAMSON.
- Paste this link into the appropriate field and click Download.

This will import a structural model that includes the relevant protein, ligand, and unbinding paths.
Step 2: Open the Export Along Paths Extension
Access the Export Along Paths app via Home > Apps > All > Export Along Paths or by searching it in the Find everything… menu (shortcut: Shift + E).
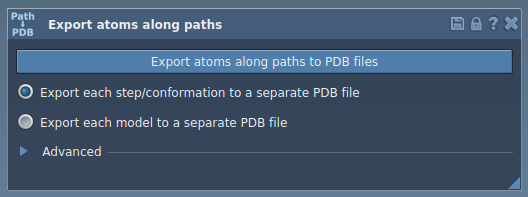
Step 3: Export Atom Trajectories
The app provides two main options for exporting trajectories:
Option 1: Export All Atoms
If you need trajectories for all atoms along a pathway:
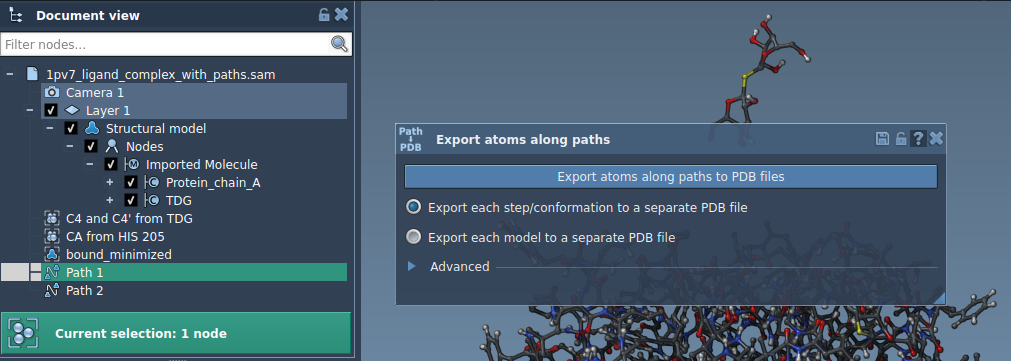
- In Document View, select the desired path(s).
- Choose how you’d like the data exported: All frames in a single PDB file or Each frame as a separate PDB file.
- Click Export atoms along paths to PDB files.
A file save dialog will appear where you can specify the destination folder and file naming convention.
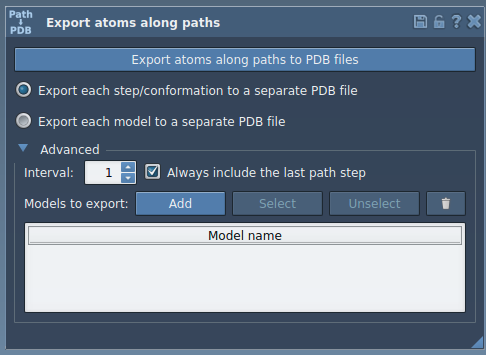
Optional: Use the Advanced panel to set parameters like the frame export interval.
Option 2: Export a Subset of Atoms
To focus on specific atoms, such as only the ligand:
- In Document View, select the atom(s) you wish to export (for instance,
TDG). - In the app, expand the Advanced section and add the selected atoms as a model.
- Customize the name or add multiple sets of atoms to define unique export models.
When ready, select the relevant path(s), export mode, and proceed as with the first option.
Applications in Molecular Modeling
Exported trajectories can aid in:
- Generating reaction coordinate files for free energy profiling.
- Visualizing intermediate states in ligand exit or entry processes.
- Tracking motion of specific areas (e.g., binding site or backbone).
These use cases demonstrate the flexibility and precision that trajectory exports can bring to your molecular design projects.
To learn more, visit the full documentation at this link.
Note: SAMSON and all SAMSON Extensions are free for non-commercial use. You can download SAMSON at www.samson-connect.net.





