Carbon nanotubes (CNTs) have taken the world of nanotechnology and materials science by storm. Their unique properties make them invaluable for designing nanoscale devices, studying molecular mechanisms, and much more. However, modeling them efficiently can be challenging, especially if you’re looking to create precise structures for simulations or designs. This is where SAMSON’s Nanotube Creator extension comes in. In this guide, we’ll walk you through how to model CNTs effectively within SAMSON, catering to molecular modelers’ need for streamlined, accurate design tools.
Why Modelers Need Efficient CNT Building Tools
When working with CNTs, molecular modelers often encounter pain points like:
- Difficulty setting precise parameters like dimensions and orientations.
- A lack of interactivity in many software tools for fine-tuning the design in real-time.
- Time-consuming processes for creating complex structures, such as multi-walled nanotubes.
The Nanotube Creator extension in SAMSON addresses these directly by offering both an interactive building mode and a graphical interface for precise control.
How to Build Carbon Nanotube Models in Two Methods
Method 1: Interactive Building in the Viewport
This method is for those who like a hands-on, real-time approach:
-
Set the nanotube axis and length by pressing and dragging the left mouse button in the viewport. As you do this, the SAMSON status bar provides live feedback on the tube’s orientation and length, helping you define the
nparameter precisely.
-
Set the nanotube radius (defining the
mparameter) by releasing the mouse button, adjusting the mouse position, and clicking again to confirm.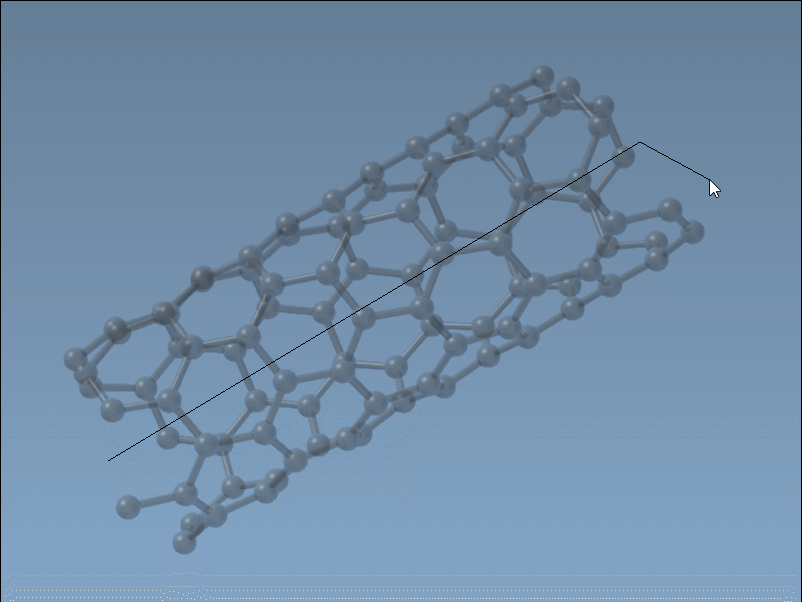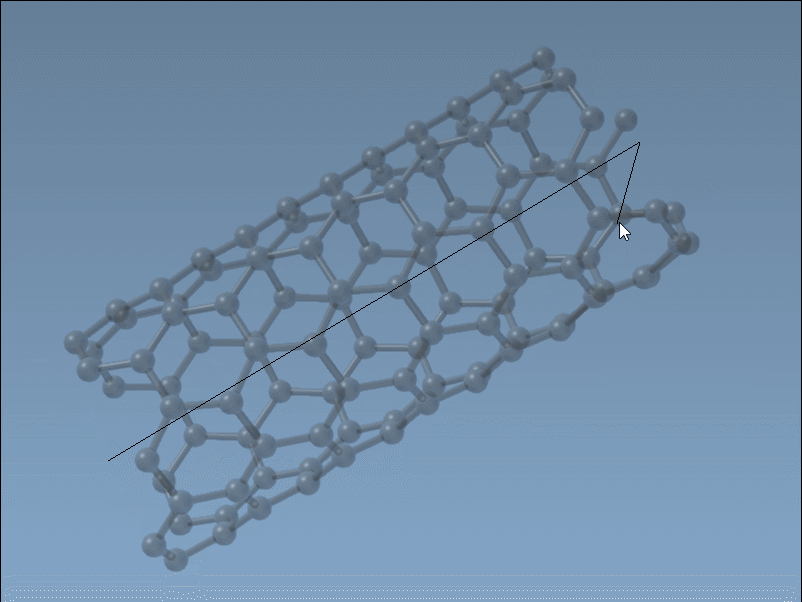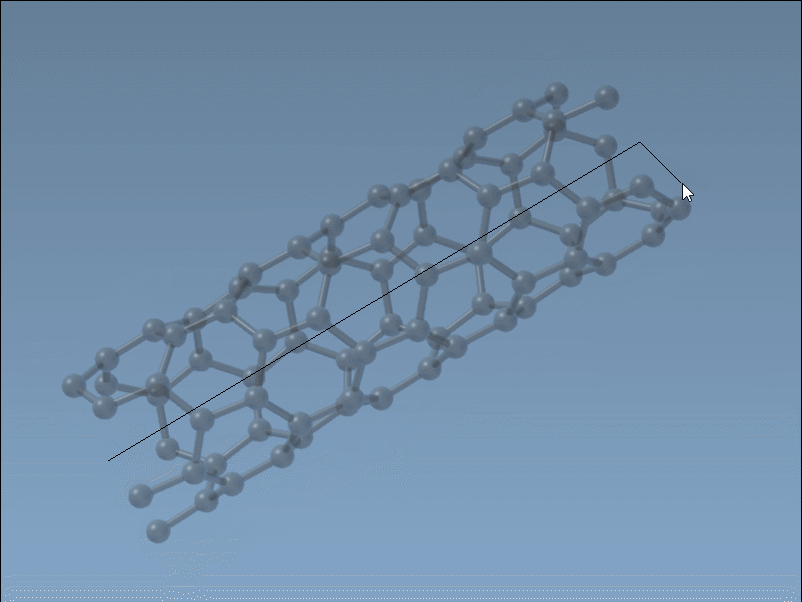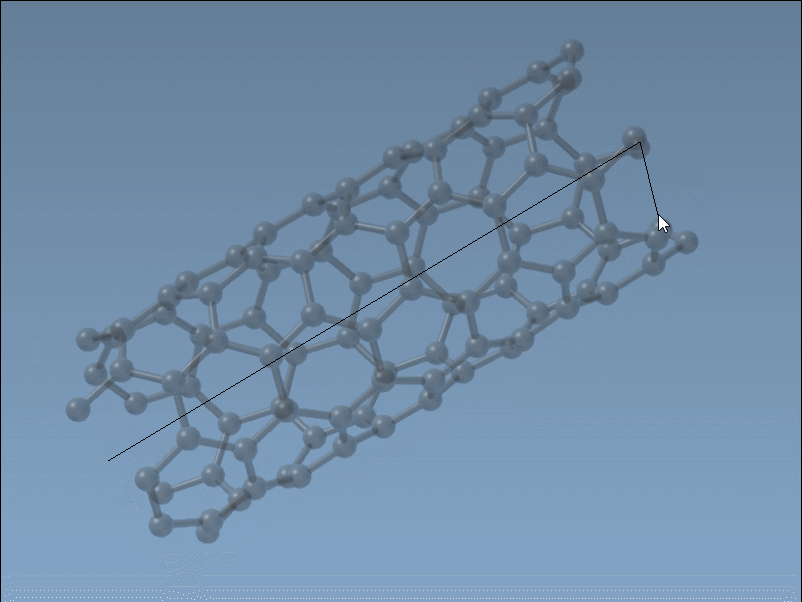
This intuitive and fast process is ideal for quick modeling tasks where visual feedback is key.
Method 2: Using the Graphical Interface
Need more control? Switch to the graphical user interface (GUI) of the Nanotube Creator editor:
- Activate the editor from the viewport menu (… > Materials > Nanotube Creator) or use the Shift + E shortcut and search for “Nanotube Creator.”
- If hidden, toggle the interface by reselecting the editor.
The GUI allows you to define exact parameters, such as start/end positions, n, and m values, and even enables the creation of complex multi-walled nanotubes. For instance, you can generate three concentric CNTs with controlled dimensions:
- Set start/end positions as
(0, 0, 0)and(40, 0, 0). - Create the first CNT with
n = 6andm = 6, then click Build. - Create the second CNT with
n = 10andm = 10, and the third one withn = 14,m = 14, clicking Build after each.
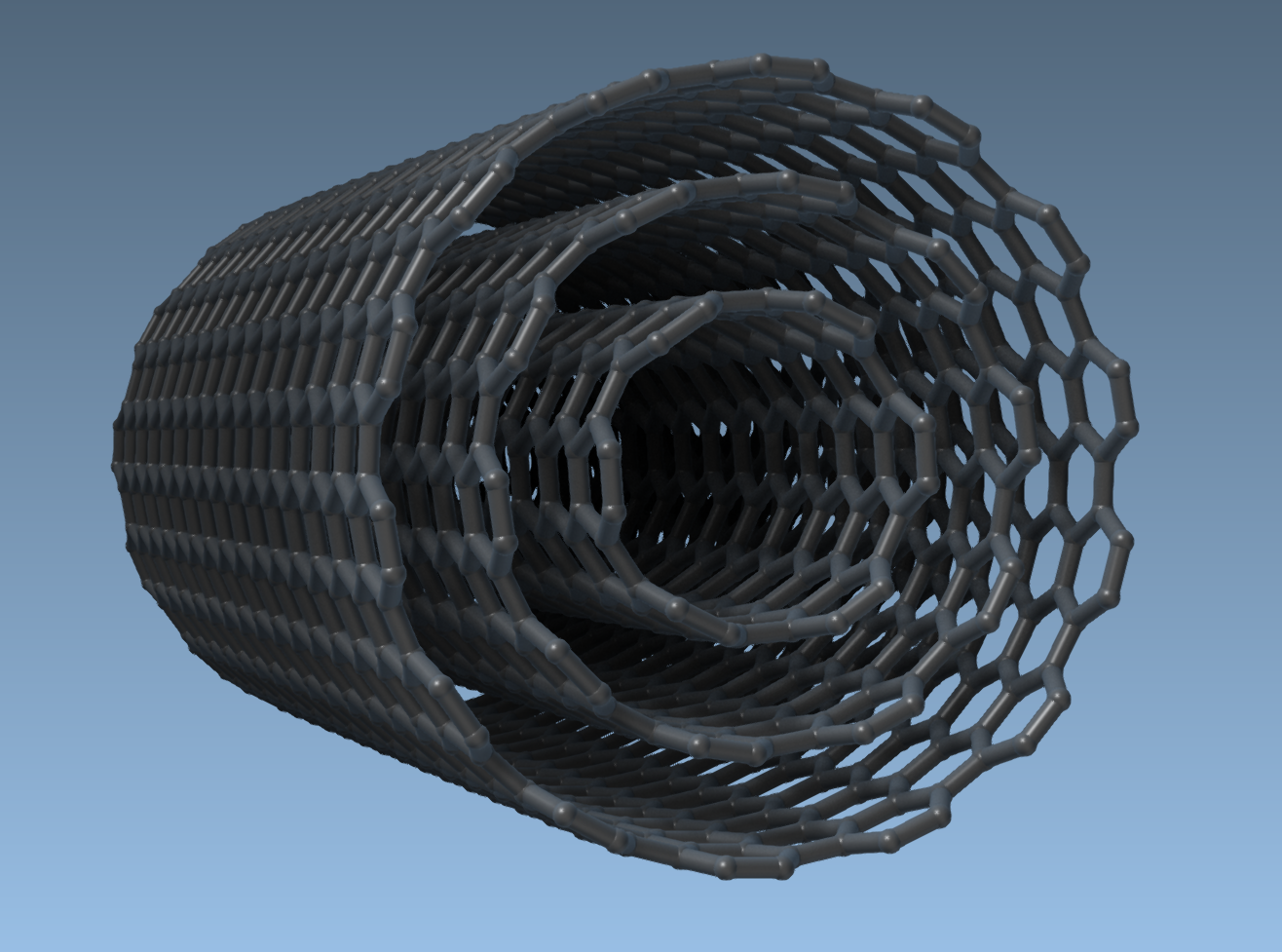

This will result in a multi-walled CNT model ready for applications in molecular simulations, materials design, and beyond.
Ready to Explore?
Once you’ve built your CNT models, SAMSON provides options to export visualizations, combine CNTs with other molecules to create hybrid structures, and even simulate their properties interactively. These features make it a one-stop solution for molecular and nanotechnology design.
For full details, reference the original documentation page at https://documentation.samson-connect.net/tutorials/nanotubes/building-nanotubes-models/.
Note: SAMSON and all SAMSON Extensions are free for non-commercial use. Get started today by downloading SAMSON from https://www.samson-connect.net.





