Author: OneAngstrom
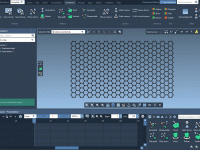
Reduce Eye Fatigue During Molecular Modeling with Dark Mode in SAMSON
Reading molecules like sentences: the Sequence View in SAMSON
Progressively Revealing Atoms: A Clearer Way to Build Molecular Animations
Understanding visibility settings in SAMSON’s Property Models
How to Efficiently Select Render Presets in SAMSON with NSL
Organizing Molecular Structures with Folders in SAMSON

As molecular modeling projects grow in complexity, so does the need to keep structures, simulations, scripts, and datasets well-organized. With multiple molecules, different simulation conditions, and embedded metadata, managing a project in a streamlined way isn’t just convenient—it’s essential. If…