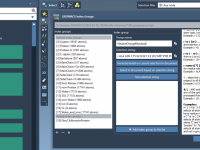
When working with large biomolecular systems using GROMACS, it’s often necessary to define custom index groups—for example, to specify pull groups for umbrella sampling, analyze specific subsets of atoms, or apply restraints. But creating these groups manually using gmx make_ndx…