As a molecular modeler, navigating massive molecular structures can be daunting. Whether you’re analyzing a complex protein, editing residues, or searching for specific atomic arrangements, precision and efficiency are crucial. That’s where the Node Specification Language (NSL) and SAMSON’s powerful Find command make a significant difference.
Bringing Clarity to Molecular Models
Node Specification Language, or NSL, allows targeted selection of molecular nodes (atoms, residues, bonds, etc.) based on their properties and relationships. But did you know you can combine NSL’s power with SAMSON’s interactive tools to rapidly zero in on specific nodes? Let’s explore how to leverage this intuitive selection process in SAMSON.
Interactive Selection 101: The Find Command
The Find command in SAMSON takes full advantage of NSL, making node selection fluid and highly specific. You can access it directly in the search box and enter an NSL string to select nodes from the active molecular document.
For example, if you need to locate alanine atoms interacting with a receptor, you can just start typing a query like "ALA (with the opening quote and without the closing one). Then, press Tab, and SAMSON will suggest context-completion based on all entities starting with “ALA.” Options might include:
"ALA 22 Backbone""ALA 22 Side chain""ALA 28 Side chain"
Click your desired selection and proceed with your analysis. This combination of auto-completion and NSL string parsing saves time while ensuring accuracy.
Efficiently Filtering in the Document View
Besides the Find command, the Document View in SAMSON lets you filter nodes interactively using an NSL query. For example, if you want to filter structural groups (such as backbone or sidechains), you can use:
n.t sg
This short query will match nodes of the type structuralGroup, making your navigation within the document seamless. To select filtered results, press the Enter key after applying the filter.
Here is an example image demonstrating this powerful selection process:
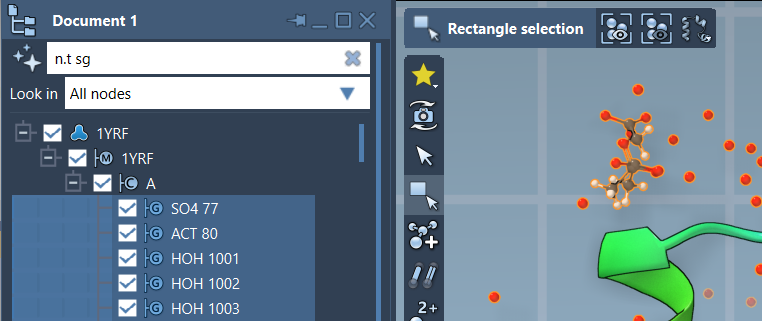
AI-Powered NSL Assistance
If you’re not sure how to craft the right NSL query, SAMSON’s AI Assistant can help! Simply click the ![]() button beside selection or filter fields. The AI Assistant understands your active molecular document’s hierarchy and can suggest NSL expressions tailored to your needs. It’s like having an expert assistant at your disposal.
button beside selection or filter fields. The AI Assistant understands your active molecular document’s hierarchy and can suggest NSL expressions tailored to your needs. It’s like having an expert assistant at your disposal.
Conclusion
SAMSON’s implementation of NSL empowers molecular modelers to navigate, select, and analyze molecular structures with ease and precision. From the intuitive Find command to robust Document View filtering and AI-guided queries, you have all the tools you need for streamlined workflows and informed decisions.
For more tips and advanced NSL examples, visit the official documentation page.
SAMSON and all SAMSON Extensions are free for non-commercial use. You can get SAMSON at samson-connect.net.





