Simplifying Center-of-Mass Pulling with GROMACS Wizard
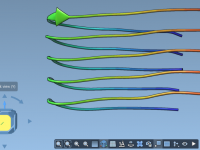
Center-of-mass (COM) pulling simulations can provide valuable insights into the molecular interactions and mechanical properties of biological systems. However, setting up such simulations can be tedious, especially when considering factors like system size, boundary conditions, and proper restraints. If you’ve…