Category: Uncategorized
Easily Visualize Reverse Molecular Trajectories in SAMSON
Turning Molecular Animations into Slides: Using the Stop Effect in SAMSON
When presenting molecular mechanisms and designs, clarity is everything. Whether you’re teaching, presenting your research, or building interactive lessons, you want your audience to follow key steps of molecular dynamics or conformational changes without distractions or information overload. This is…
One Click to Clarity: Simplify Molecular Visualization with SAMSON Visual Presets
When You Can’t See It: Managing Label Visibility in Molecular Models
Dragging Atoms to Simulate Molecular Behavior in Real Time
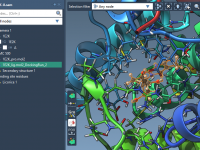
Customizing Protein-Ligand Structure Predictions with Boltz-2 in SAMSON
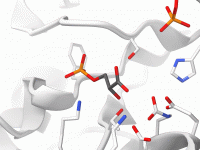
Speeding Up NMR Structure Prediction by Assigning Methyl Groups
For molecular modelers working with protein-ligand complexes, Nuclear Magnetic Resonance (NMR) remains a go-to experimental method when X-ray crystallography becomes impractical. However, interpreting NMR data in silico, especially ambiguous NOESY peaks involving unassigned methyls, can quickly turn computationally expensive. When…
Creating Multiple Protein Replicas in SAMSON without Errors
Working on molecular simulations often means scaling up. Whether you’re modeling protein aggregation, crowded cellular environments, or simply running multiple replicas for statistical accuracy, duplicating protein structures is essential. But, if you’ve ever tried this manually, you’ve likely encountered issues…
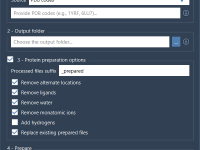
Cleaning Hundreds of Protein Structures Just Got Simpler
Easily Visualize Reverse Molecular Trajectories in SAMSON
Turning Molecular Animations into Slides: Using the Stop Effect in SAMSON
When presenting molecular mechanisms and designs, clarity is everything. Whether you’re teaching, presenting your research, or building interactive lessons, you want your audience to follow key steps of molecular dynamics or conformational changes without distractions or information overload. This is…
One Click to Clarity: Simplify Molecular Visualization with SAMSON Visual Presets
When You Can’t See It: Managing Label Visibility in Molecular Models
Dragging Atoms to Simulate Molecular Behavior in Real Time
Customizing Protein-Ligand Structure Predictions with Boltz-2 in SAMSON
Speeding Up NMR Structure Prediction by Assigning Methyl Groups

For molecular modelers working with protein-ligand complexes, Nuclear Magnetic Resonance (NMR) remains a go-to experimental method when X-ray crystallography becomes impractical. However, interpreting NMR data in silico, especially ambiguous NOESY peaks involving unassigned methyls, can quickly turn computationally expensive. When…
Creating Multiple Protein Replicas in SAMSON without Errors

Working on molecular simulations often means scaling up. Whether you’re modeling protein aggregation, crowded cellular environments, or simply running multiple replicas for statistical accuracy, duplicating protein structures is essential. But, if you’ve ever tried this manually, you’ve likely encountered issues…