Molecular modelers often face a challenging question when performing ligand docking: how to efficiently manage the flexibility of ligands? Excessive bond flexibility can significantly increase computational time, while insufficient flexibility might lead to unrealistic docking results. AutoDock Vina Extended, an advanced docking extension in SAMSON, offers tools to tackle this issue effectively by enabling users to refine ligand bond flexibility with precision.
Why Is Ligand Flexibility Important?
Ligand flexibility refers to the ability of certain bonds within the molecule to rotate, thus altering the molecule’s conformation. Allowing such rotations helps simulate realistic docking scenarios, as the ligand can adapt its shape to better fit the binding site. However, enabling all rotatable bonds indiscriminately can increase the docking computation time dramatically. Striking the right balance is key for both accuracy and speed.
Managing Ligand Rotatable Bonds in AutoDock Vina Extended
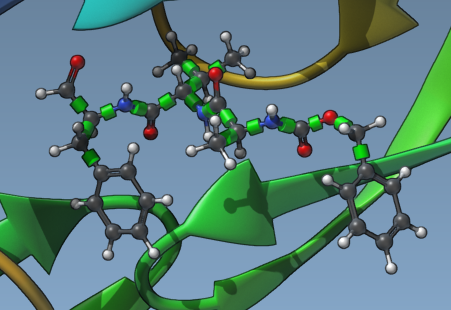
Setting up the flexibility of ligands is intuitive in AutoDock Vina Extended. After selecting your ligand in the Document view, you’ll notice green cylinders representing the rotatable bond controllers on superimposed bonds in the Viewport.
Here’s how to refine their flexibility:
- Visual Bond Control: By clicking on the green cylinders, you can toggle the bonds between rotatable (green) and non-rotatable (red).
- Locking Specific Bond Types: The extension allows you to automatically lock specific bond types (e.g., double bonds, aromatic bonds). Open the Locked bonds settings menu, select the types of bonds to lock, and click OK. Check the Lock specific ligand bonds option to apply these settings uniformly.
The ability to visual and customize bond flexibility gives modelers unprecedented control, ensuring that structural constraints of complex ligands are respected while maintaining computational efficiency.
Enhancing Ligand Preparation
If ligands in your library are not well-prepared or are in 2D, consider using the minimization features in AutoDock Vina Extended. By checking the Minimize option during setup, you can ensure that ligands are fully optimized for docking. Additionally, enabling the Add missing hydrogens option ensures proper hydrogen bonding potential, crucial for accurate docking simulations.
Tips for Efficient Ligand Library Docking
- When working with a large ligand library, apply the Lock specific ligand bonds option to default some bond types as non-rotatable. This dramatically reduces computation time.
- Ensure each ligand is minimized and properly protonated. Missing hydrogens or improperly assigned bond types could degrade docking results.
- Preview bond flexibility visually in the Viewport to confirm that all critical aspects of ligand flexibility are appropriately set before proceeding to docking.
Conclusion
By leveraging the customizable ligand flexibility tools in AutoDock Vina Extended, molecular modelers can achieve a more efficient and accurate docking workflow. Learning to refine these parameters will enhance your ability to achieve realistic simulations without unnecessary computational overhead.
For detailed instructions and to explore further capabilities, visit the full documentation page.
SAMSON and all SAMSON Extensions are free for non-commercial use. Get SAMSON at https://www.samson-connect.net.





