One of the challenges molecular modelers often face is identifying physically meaningful transition paths between molecular conformations. If you’ve ever struggled with refining a rough transition path, the Parallel Nudged Elastic Band (P-NEB) app in SAMSON offers an efficient and intuitive solution to improve your pathways.
Why Transition Path Optimization Matters
Transition paths help model essential molecular mechanisms like ligand binding/unbinding or protein conformational changes. However, rough paths, such as those generated from linear interpolation, often lack physical accuracy. This can result in poorly optimized pathways, making simulations or further analyses unreliable. With P-NEB, you can fine-tune these paths to ensure they represent the actual transition with minimized energy discrepancies along the trajectory.
What is the P-NEB Method?
The Nudged Elastic Band (NEB) method is a popular approach to optimization, allowing the refinement of intermediate states while preserving equal distribution along the path using spring forces. P-NEB improves upon traditional NEB by integrating parallel computing (processing one thread per conformation) to accelerate the optimization process.
With SAMSON’s P-NEB app, you can:
- Relax transition paths into more physically meaningful ones.
- Improve transition paths for specific studies, like pathways between local energy minima.
- Leverage features like the climbing image method for advanced saddle point detection.
Getting Started with P-NEB in SAMSON
Before diving into the P-NEB app, make sure you have:
- Added the P-NEB Extension and the FIRE state updater.
- A prepared rough path or set of conformations ready for refinement.
Access the P-NEB Extension via Home > Apps > All > P-NEB and configure the following settings:
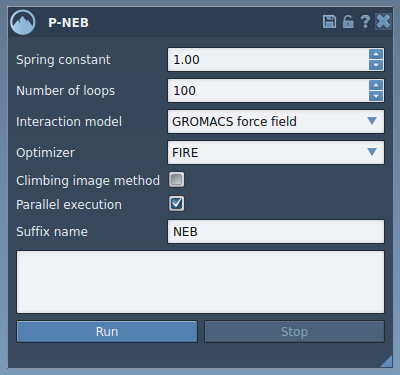
The app allows customization of parameters, including:
- Spring constant: Define the spring coefficient (1.00, for example).
- Number of loops: Specify the number of optimization cycles (e.g., 100).
- Interaction model: Choose a force field model, such as “Universal Force Field”.
- Optimizer: Select an optimization algorithm (FIRE is recommended).
- Parallel execution: Enable for better performance on multi-conformation paths.
- Climbing image method: Use this for saddle point detection if needed.
Applying P-NEB to a Path
In the Document view, select the path node you want to optimize and click the Run button within the P-NEB interface.
During the process, you’ll be prompted to configure the Universal Force Field settings. Simply confirm to use existing bonds, and your optimization will begin. The progress can be monitored in the app’s status bar.

After optimization, the refined path will appear in the Document view ready for further inspection, visualization, or uses in simulations.
Optimize Sets of Conformations
If you’re working with individual conformations rather than pre-generated paths, P-NEB can optimize these as well. Note that this approach may take longer since conformations are optimized separately. It’s often more efficient to combine conformations into a path before applying P-NEB. SAMSON provides a simple option to do this via Conformation > Create path from conformations.
Take Your Study Further
Once your path is optimized, you can inspect it using SAMSON’s Inspector, export trajectories, or continue your investigation with other apps. P-NEB’s versatility ensures compatibility across a wide range of studies, from enzyme mechanisms to drug design workflows.
To go deeper into the P-NEB app’s full capabilities and learn more details about the optimization process, visit the official documentation page at this link.
Note: SAMSON and all SAMSON Extensions are free for non-commercial use. You can download SAMSON at samson-connect.net.





