Docking libraries of ligands is a cornerstone task for many molecular modelers, especially those aiming to identify promising drug candidates. However, setting up such studies with traditional methods can be time-consuming and prone to errors. The AutoDock Vina Extended SAMSON Extension offers a streamlined solution to simplify this procedure while maintaining flexibility and precision. Here’s how it can make your workflow smoother and more efficient.
Why Docking Ligand Libraries Matters
Docking ligand libraries against a receptor allows researchers to analyze and predict multiple binding modes and affinities efficiently. Screening entire libraries can help uncover potential leads for further experimental validation, accelerating the drug discovery process. However, preparing the system and handling flexible ligands can be complicated.
The AutoDock Vina Extended SAMSON Extension integrates the popular AutoDock Vina program with enhanced functionalities, such as handling flexible ligands, setting rotatable bonds, and managing ligand libraries. Here’s a focused guide on effectively using it for a ligand library, along with actionable tips.
Step 1: Setting Up Your Ligand Library
Start by preparing the ligands you want to dock in a directory. The library can contain files of various formats, such as MOL2 or SDF, and may include multiple ligands per file. To load the library, check the Ligand library option and click on the browse button to select your directory:
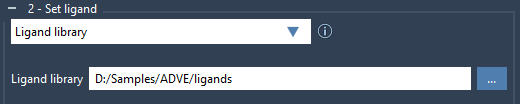
By default, all the ligands will be considered fully flexible. However, specific bond types can be locked to avoid unnecessary rotations. To do this, access the Locked bonds settings and specify which bonds to lock (e.g., C=N bonds).
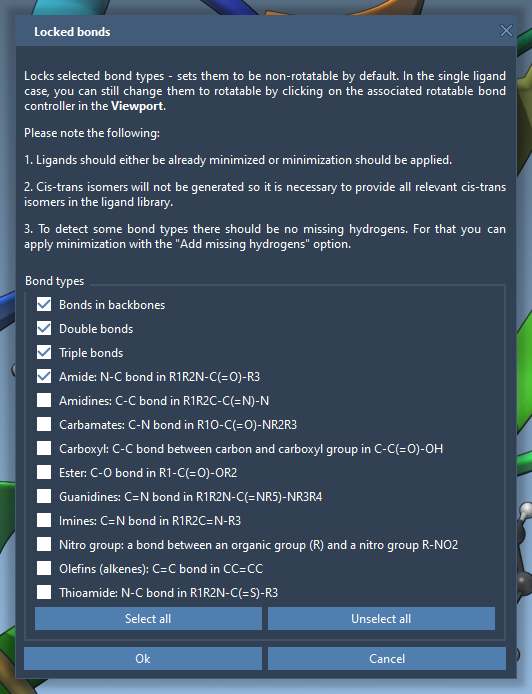
Once configured, select Lock specific ligand bonds to apply these settings to the entire library:
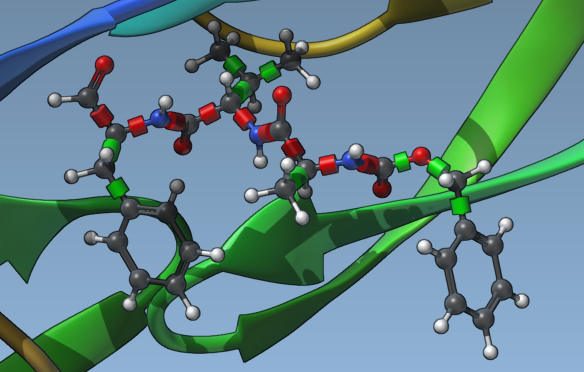
Step 2: Optional Ligand Minimization
If the ligands have not been pre-optimized, you can leverage the built-in minimization feature. Check the Minimize option and select a preset that determines the number of minimization steps and stopping criteria. This step can refine the ligands’ geometry before docking and add missing hydrogens if needed.

The minimization stops when either the energy difference between steps falls below a threshold for consecutive steps or a maximum number of steps is reached. Choose the option that suits your system’s needs for efficient preparation.
Step 3: Performing the Docking
Once your ligand library is set up, initiate the docking process. Under the Dock settings, select Dock to start the computations. You can also specify advanced parameters like exhaustiveness and the maximum number of binding modes to generate for a robust exploration of conformational space. To access these options, click the Options button.
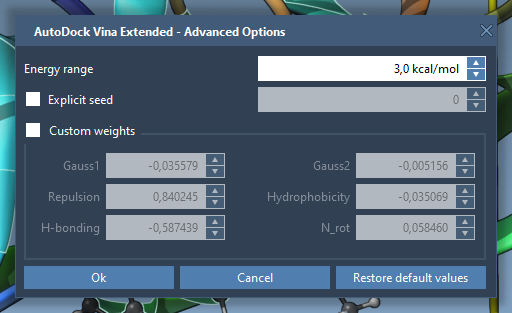
If needed, enable the Save results feature to ensure all docking outcomes are stored in a well-organized directory for subsequent analysis. Once the ligand library is processed, the extension will generate a results folder containing configurations, top binding ligands, and conformations.
Step 4: Analyzing Your Results
Post-docking, AutoDock Vina Extended provides tools to analyze and visualize docking results. The Results tab allows sorting ligands by affinity, filtering based on defined parameters, and generating plots, such as binding affinity vs. ligands. These data points are interactive, enabling users to restore conformations directly from the plot by double-clicking on them:
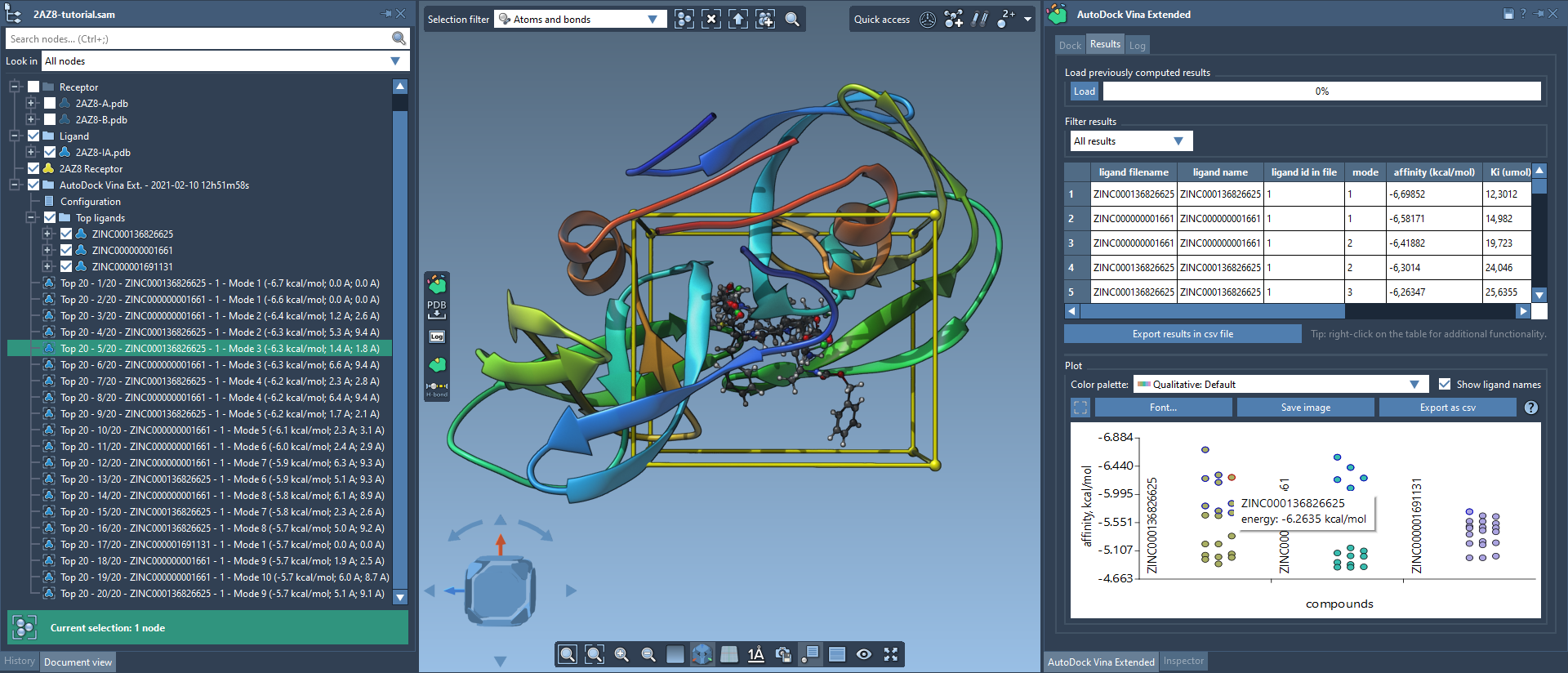
Moreover, export options allow researchers to save ligand conformations in multiple formats for further study. The extension’s intuitive interface ensures that insights are straightforward to extract.
To dive deeper and maximize the potential of your docking studies, make sure to consult SAMSON’s official documentation for the AutoDock Vina Extended extension.
Note: SAMSON and all SAMSON Extensions are free for non-commercial use. Download SAMSON at https://www.samson-connect.net.





