One of the recurring challenges for molecular modelers is efficiently navigating and refining their increasingly complex molecular datasets. Whether you’re working with thousands of atoms, residues, or structural groups, the ability to quickly filter and select relevant nodes based on specific criteria can save significant time and effort. SAMSON’s Node Specification Language (NSL) provides an elegant and powerful solution to this issue.
Why Filter Nodes?
Imagine identifying all atoms within 5 angstroms of a ligand in your model or isolating backbone nodes from a protein. These are just two examples of tasks that molecular modelers often face. Without a precise filtering mechanism, such tasks can be both tedious and error-prone. Fortunately, NSL allows you to accurately and efficiently filter nodes using expressions directly in SAMSON’s Document View.
How Does Node Filtering Work?
With NSL, you can specify a filtering expression to isolate nodes based on attributes such as type, proximity, or even relationships with other nodes. For instance, entering n.t sg (short for node.type structuralGroup) filters and displays only structural groups in your Document View. To apply your desired filter, you need to:
- Open the Document View in SAMSON.
- Enter your NSL string in the filtering panel.
- Press Enter to apply the filter and visualize only the relevant nodes.
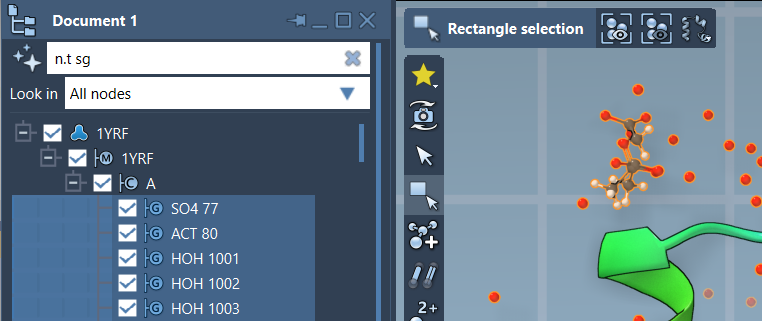
Below is an example of filtering structural groups using the NSL expression n.t sg:
Explore Context-Sensitive Assistance
Not sure which NSL expression you need? You can leverage SAMSON’s AI Assistant, available within the Document View. The Assistant is designed to understand the hierarchy of your active document and can generate specific NSL expressions tailored to your needs. Simply click on the Ask AI button alongside the filter nodes panel to get started.
Practical Examples for Everyday Use
Here are some scenarios where NSL filtering can simplify your workflow:
C within 5A of "Ligand"— Filters carbons located within 5 angstroms of any node named “Ligand” (useful for analyzing ligand-protein interactions).n.t r and not r.t ALA— Displays all non-alanine residues in your model.node.type residue linking r.t CYS— Identifies residues that have atoms bonded to cysteine residues.
For more complex applications, you can utilize NSL’s support for advanced topology and proximity operators. For example, the expression r.ss helix selects residues that are part of an alpha helix, which is particularly beneficial when studying secondary structures.
Conclusion
SAMSON’s NSL functionality ensures precision and efficiency when filtering nodes in large molecular datasets. This invaluable tool is not only intuitive but also customizable to your specific requirements. To learn more about NSL and its various capabilities, visit the official documentation at https://documentation.samson-connect.net/users/latest/nsl/.
Note: SAMSON and all SAMSON Extensions are free for non-commercial use. Click here to download SAMSON today.





