For molecular modelers working with protein dynamics, understanding how a protein transitions between two conformations is often crucial. Yet, generating smooth and biologically meaningful transition paths between these states can be both time-consuming and complex. This is precisely where SAMSON’s ARAP Interpolator offers an elegant and efficient solution.
Why Focus on Transition Paths?
In protein modeling, transition paths are not merely visual aids; they are key for deeper insights. Whether you’re conducting conformational analyses, setting up umbrella sampling simulations, or studying transition states, having a continuous, realistic pathway enables better hypotheses and computational experiments.
SAMSON’s As-Rigid-As-Possible (ARAP) Interpolation tool is designed to streamline this process, generating smooth transition paths within seconds. Here’s how you can make the most of it!
Step-by-Step Guide to ARAP Protein Interpolation
Step 1: Prepare Your Protein Structures
To start, you’ll need two protein conformations, typically fetched from either experimental data or databases. For instance, let’s consider two conformations (1DDT and 1MDT) of the Diphtheria Toxin:
- Open the Home > Fetch menu in SAMSON.
- Input your structure IDs (e.g.,
1DDTand1MDT) and load them.
The fetched files might include non-structural elements such as water molecules or ions. These need to be cleaned to focus solely on the protein chain of interest:
- Select a specific chain, like chain
A, and remove unwanted chains (e.g., chainBin 1MDT). - Run Home > Prepare to clean up the structures.
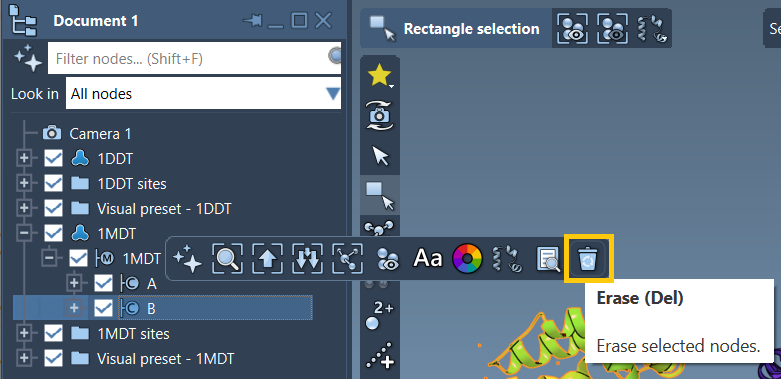
These actions ensure your two protein structures are prepped for interpolation.
Step 2: Define Protein Conformations
Once your structures are prepped, define them as conformations in SAMSON:
- Select a structure, say
1DDT, from the document view. - Create a conformation via Edit > Conformation and assign it a name like
1DDT A.
Repeat this for 1MDT, naming it appropriately. These conformations will act as your transition start and goal points.
Step 3: Run the ARAP Interpolation Tool
Access the tool via Home > Apps > Biology > ARAP Path Interpolation. Use the following key settings for a smooth path:
- Match atoms: Choose “All except hydrogens” for biologically meaningful alignment.
- Build edges: Check options like “from bonds in the Start structure” and “Try connecting α-carbons before and after missing residue segments”.
- Alignment: Enable “Perform alignment before interpolation” to ensure accurate overlays.
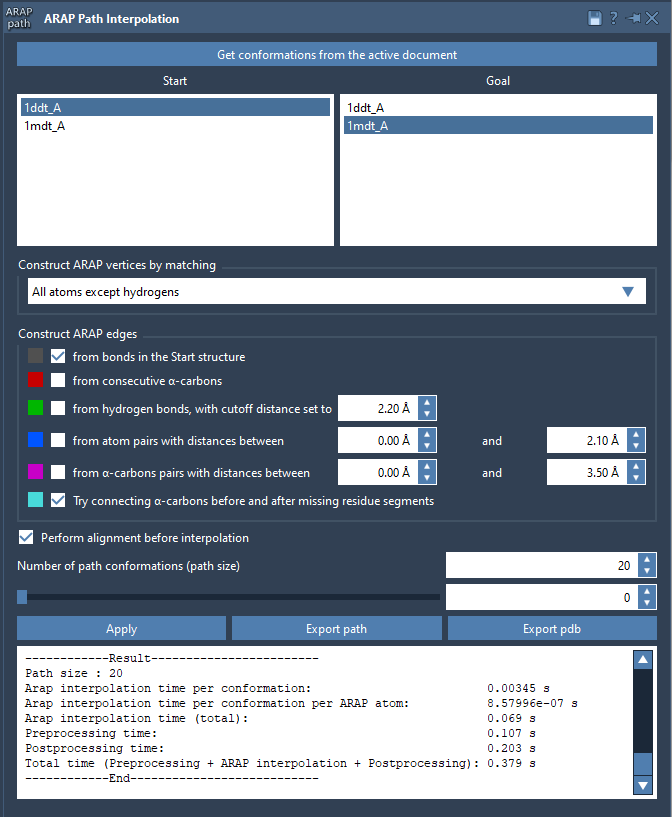
Set the “Number of path conformations” to define how detailed the transition pathway will be. For instance, selecting 20 generates 18 intermediate steps between the start and goal states, offering a detailed visual trajectory.
Click Run, and within seconds, a smooth transition path is computed!
Step 4: Analyze and Export
Once the interpolation completes, the results are displayed interactively. Use the slider to review the intermediate steps and verify the biological plausibility of the transition.
- Export the path as a trajectory object or output it as sequential PDB files.
- Toggle the visualization of ARAP edges to explore the connectivity patterns.
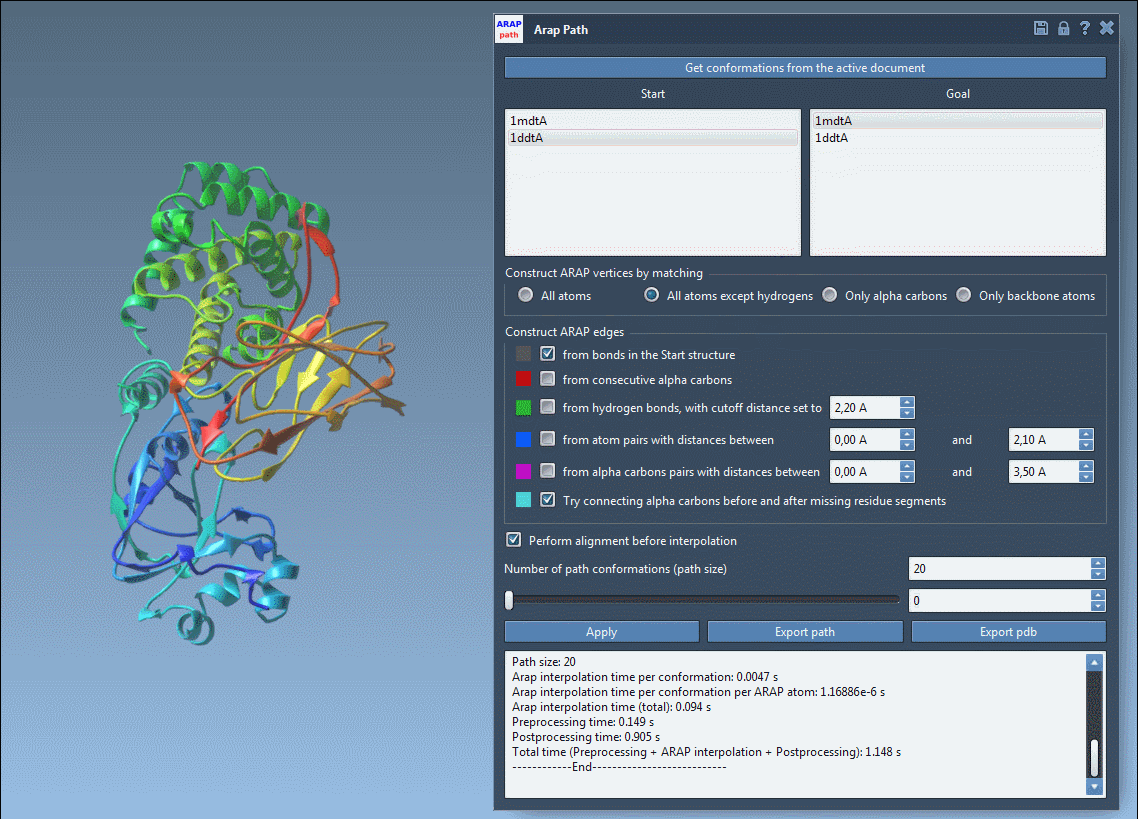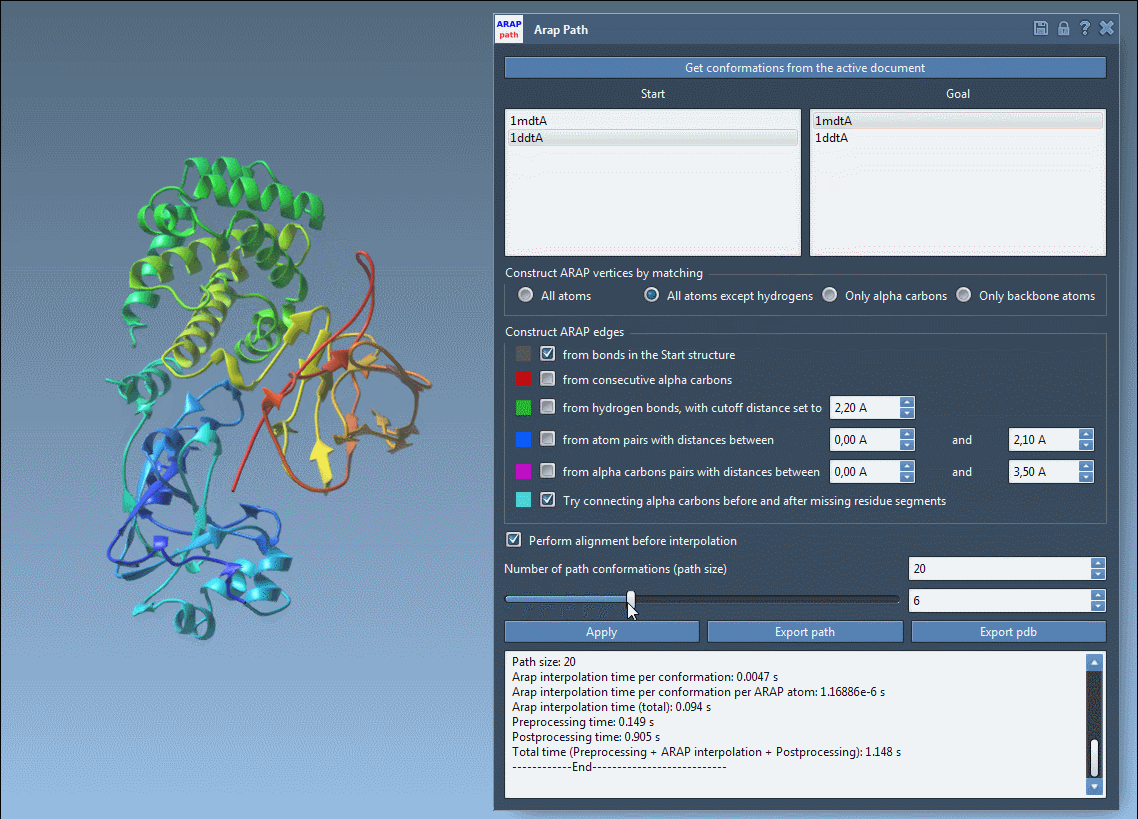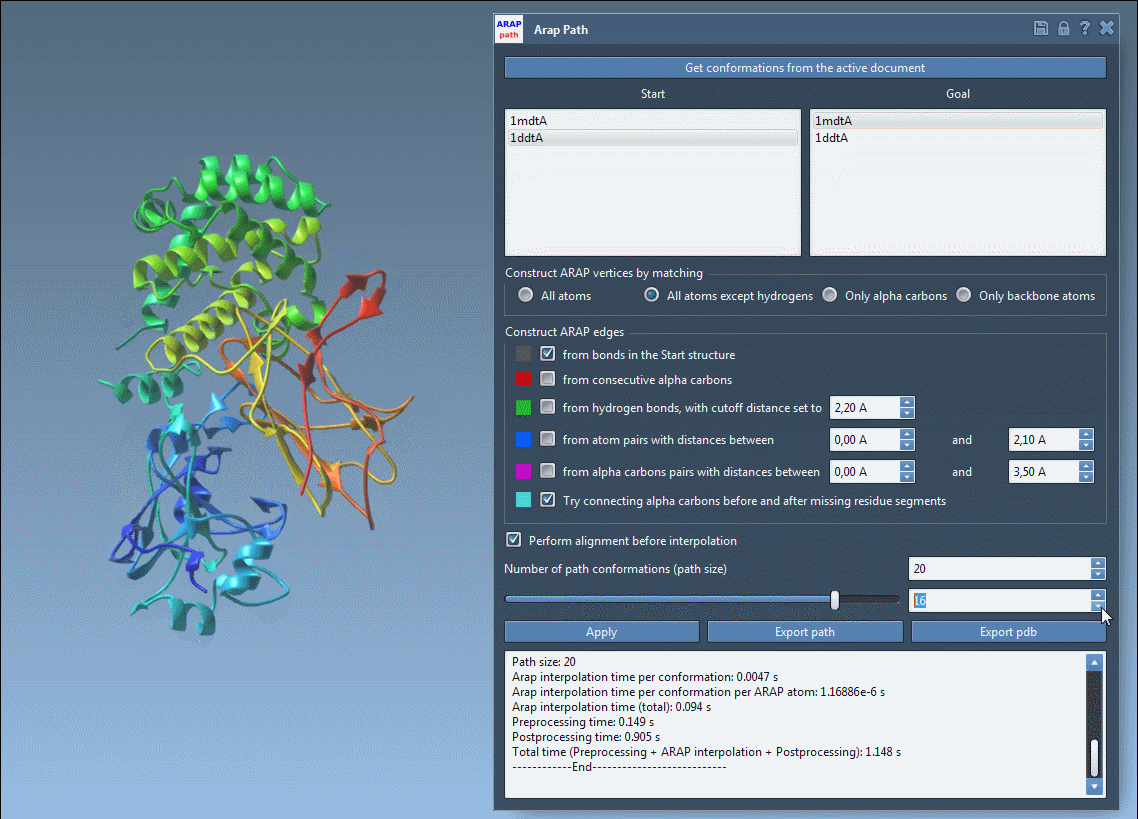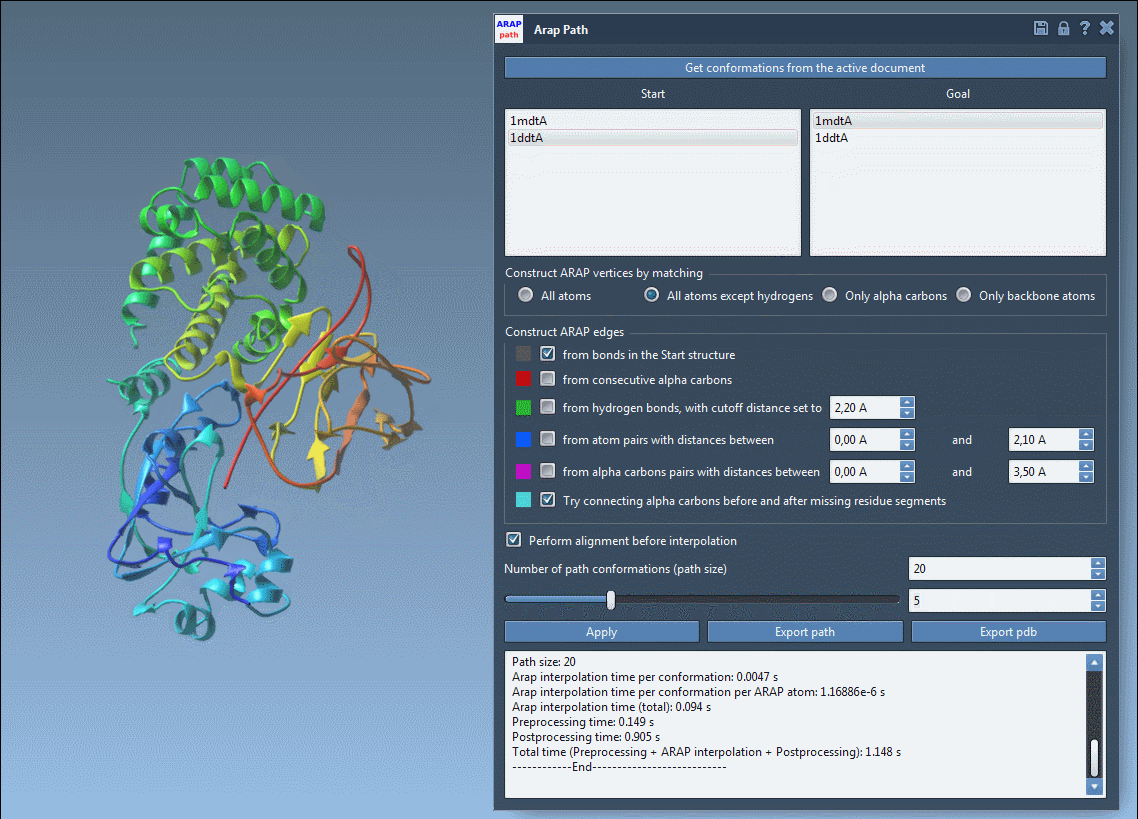
With SAMSON, these result files can directly inform further applications, such as umbrella sampling setup with the GROMACS Wizard, refinement of pathways using Parallel Nudged Elastic Band, or dimensionality reduction of the protein ensemble.
Conclusion
SAMSON’s ARAP Interpolator simplifies the often-complicated task of protein transition path modeling. By offering a fast, intuitive tool to construct biologically meaningful pathways, it empowers researchers to focus on analyzing dynamics rather than generating data.
To dive deeper, you can explore the comprehensive ARAP Interpolation documentation page.
Note: SAMSON and all SAMSON Extensions are free for non-commercial use. You can download it at SAMSON Connect.





