Understanding the spatial arrangements of proteins is critical for molecular modeling, design workflows, and simulations. One common challenge molecular modelers face is reconstructing biological assemblies and crystal structures from available data. If you’ve ever needed to visualize how proteins interact within their natural environments, or if you’re curious about how to quickly develop symmetric assemblies, the Symmetry Mate Editor in the SAMSON platform offers a robust, user-friendly solution.
The Symmetry Mate Editor allows molecular modelers to recreate symmetric replicas of proteins using transformation data embedded in PDB files. These replicas are particularly useful for analyzing biological assemblies and crystal packings stored in these files. Below, we’ll guide you through exploring symmetry mates effectively and discuss how this feature can significantly ease your workflows.
Preview and Explore Symmetry Mates in Real-time
The Symmetry Mate Editor helps you visualize symmetry transformations interactively. Here’s how it works:
- Activate the editor: You can open the editor through the Find everything search bar by pressing Shift + E, or via the Editors menu (… > General > Symmetry Mate Editor).
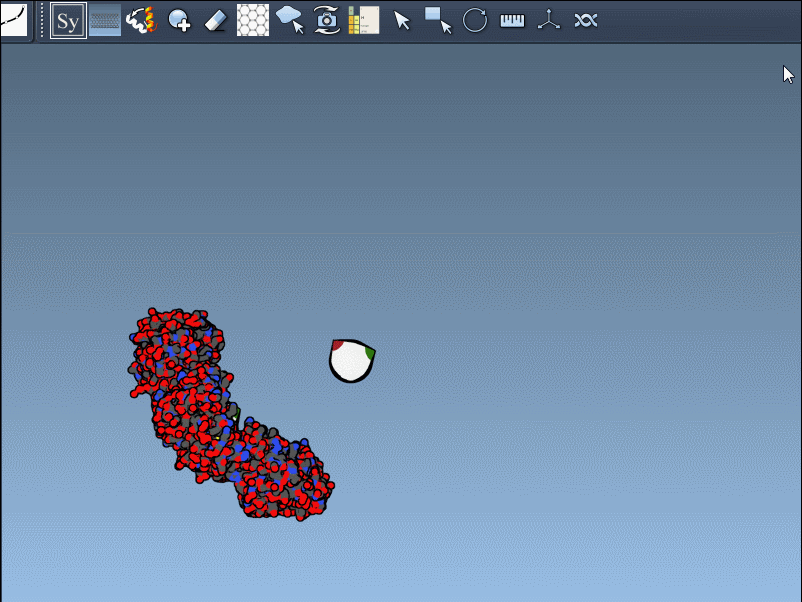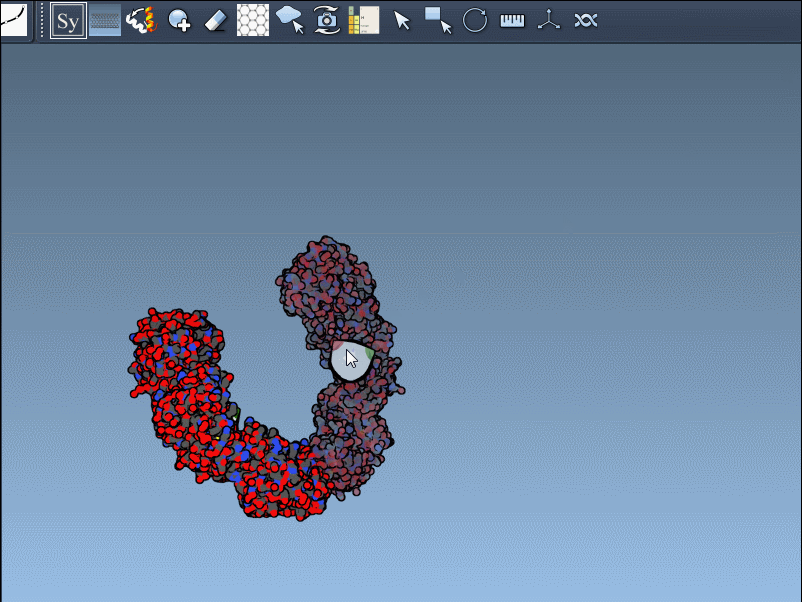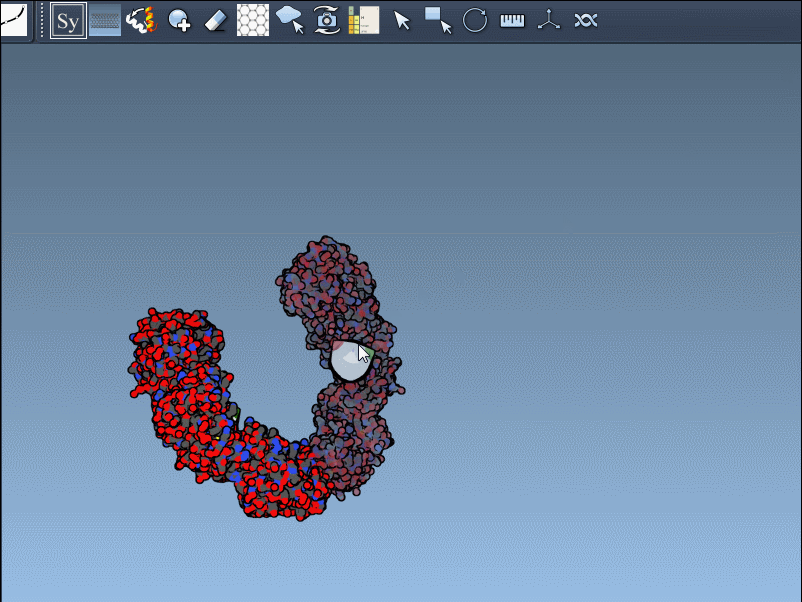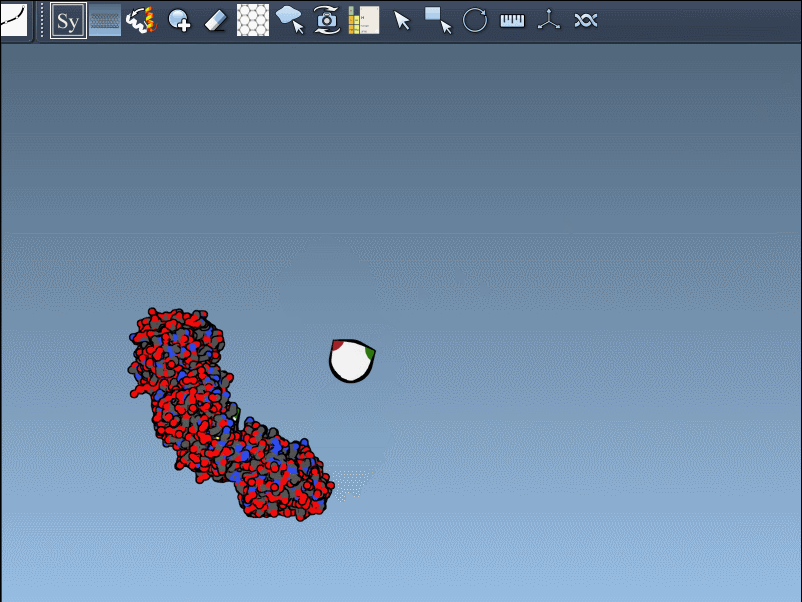
- Once activated, the viewport will display control nodes. Each node represents a transformation that can be applied to generate symmetry mates.
- Using Ctrl/Cmd + mouse wheel, you can scale the number of visible control nodes, making it easier to manage complex structures.
- Hover over a node to preview a symmetry mate interactively. These previews are real-time, giving you instant feedback on potential structural arrangements.
- Generate replicas: Left-click on a control node to permanently generate the replica in your model. This simple action allows you to build complete structures step by step.
Here’s an example in action:
Customize Your View
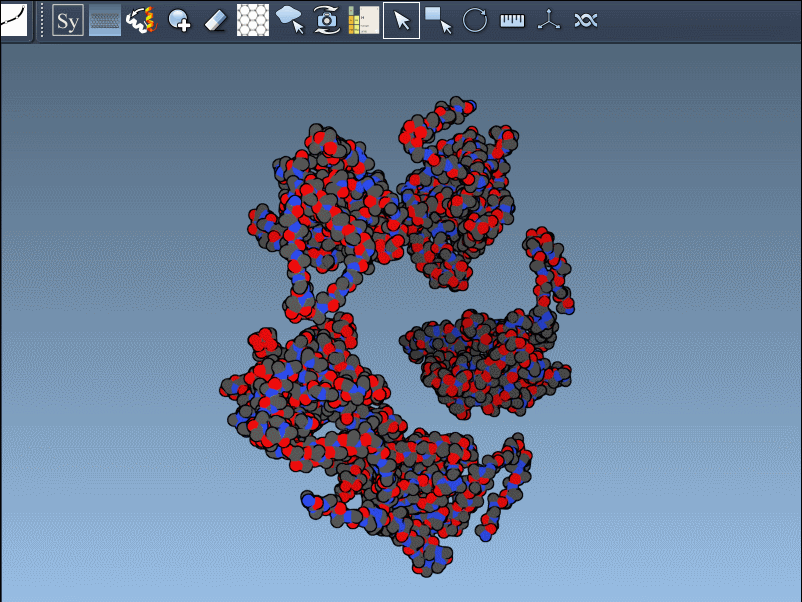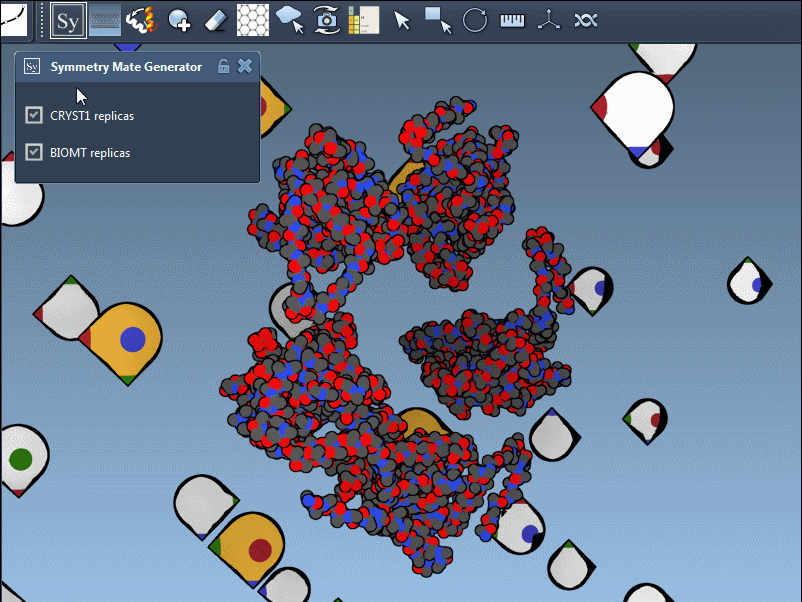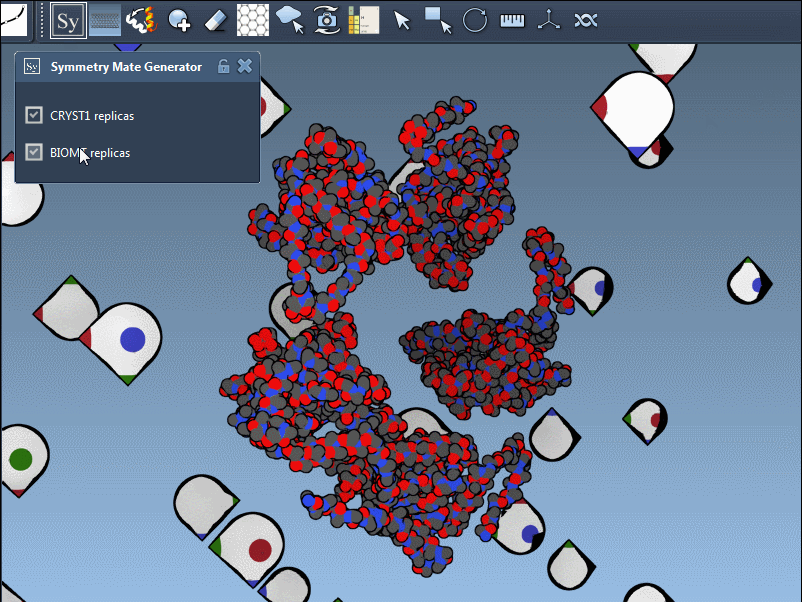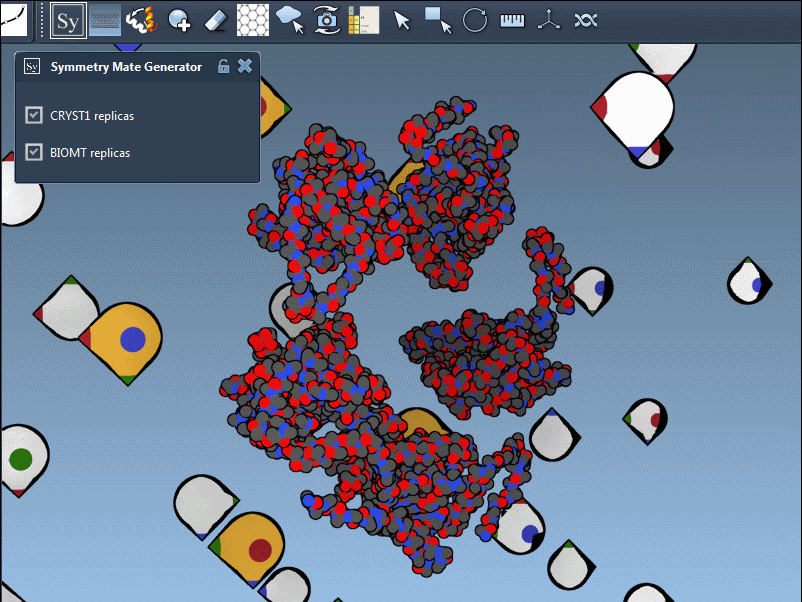
To fine-tune your exploration, you can toggle between symmetry types using the editor interface. The Symmetry Mate Editor supports two types of symmetry records found in PDB files:
- CRYST1: Represents symmetry from the crystal lattice.
- BIOMT: Represents symmetry from biological assembly annotations.
Each record type has a distinct widget color (CRYST1 in white, BIOMT in yellow) for ease of identification. Here’s an example of switching between the CRYST1 and BIOMT symmetries:
Applications for Molecular Modelers
The implications of using the Symmetry Mate Editor go beyond simple visualizations:
- Model oligomeric protein assemblies from asymmetric units.
- Design scaffolds or protein nanostructures with inherent symmetry.
- Analyze quaternary structures for molecular dynamics simulations.
- Map and test binding site repetition across different symmetric units.
These features provide critical insights into protein behavior and interactions within a molecular design environment, saving hours of manual reconstruction.
Conclusion
The Symmetry Mate Editor is a comprehensive tool that simplifies one of the most tedious aspects of molecular modeling. Its interactivity and ease of use make exploring symmetry mates not only productive but also intuitive.
To learn more about generating symmetry mates, visit the official documentation page. If you haven’t tried SAMSON yet, it’s worth noting that SAMSON and all SAMSON Extensions are free for non-commercial use. Get started today by downloading it from SAMSON’s website.
*Note: SAMSON and all SAMSON Extensions are free for non-commercial use. You can get SAMSON at samson-connect.net.





