Conformational changes in proteins are critical to their functions, yet studying these transitions can be quite a challenge. Many molecular modelers face the pain of creating realistic transition paths between protein states for simulations, analysis, or structural studies. This is where the As-Rigid-As-Possible (ARAP) interpolation in SAMSON can significantly aid your workflow.
Simplifying Protein Transition Path Creation
ARAP interpolation provides a user-friendly way to compute smooth and realistic transition paths between two protein conformations. With this, molecular modelers can:
- Compute continuous structural transitions in just seconds.
- Visualize and export intermediate structures for further analysis.
- Integrate biologically meaningful transition pathways into free energy simulations.
This tool is particularly valuable in applications such as conformational analysis, transition state modeling, or setting up simulations like umbrella sampling. One remarkable example of its application is its use in studying the opening motion of the SARS-CoV-2 spike protein, showcasing its potential in understanding critical biological processes.
Steps to Generate a Transition Path
Here is a concise overview of how to use the ARAP interpolator in SAMSON:
1. Prepare Your Protein Structures
Ensure your starting and goal protein structures are properly cleaned:
- Fetch the required Protein Data Bank (PDB) files.
For instance, to study conformational changes in the diphtheria toxin: - Enter “
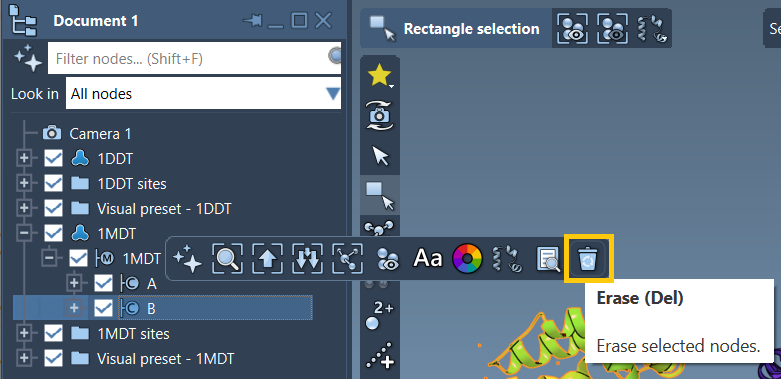
1DDT 1MDT” in PDB or PDB (mmCIF). - Load and delete non-essential chains, e.g., chain B from
1MDT. - Use the Home > Prepare tool to clean structures by removing water, ions, and ligands.
2. Create Conformations
Designate the starting (e.g., 1DDT A) and goal (e.g., 1MDT A) conformations for interpolation:
- Select and name one structure at a time using Edit > Conformation.
3. Run ARAP Interpolation
Launch the ARAP interpolation app in SAMSON under Home > Apps > Biology > ARAP Path Interpolation and adjust the required settings:
- Choose starting and goal conformations.
- Set parameters to match relevant atoms (e.g., “All except hydrogens”).
- Define how connectivity should be built, such as using α-carbon connections for missing residues.
- Enable alignment before interpolation and specify the desired number of intermediate conformations.
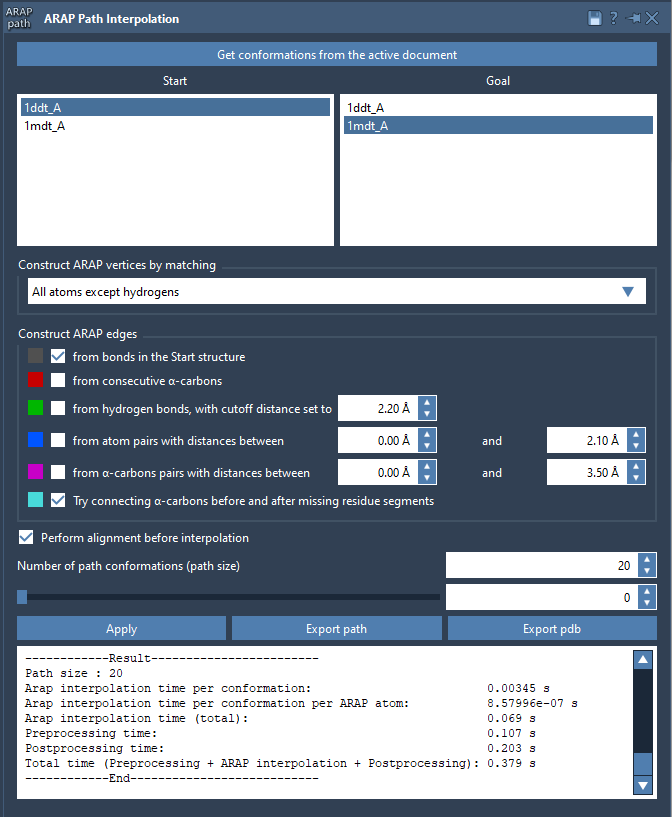
Hit Run, and the system will calculate a precise and biologically meaningful transition path in seconds.
Analyze and Export Your Results
Once the computation is complete, you’ll be able to:
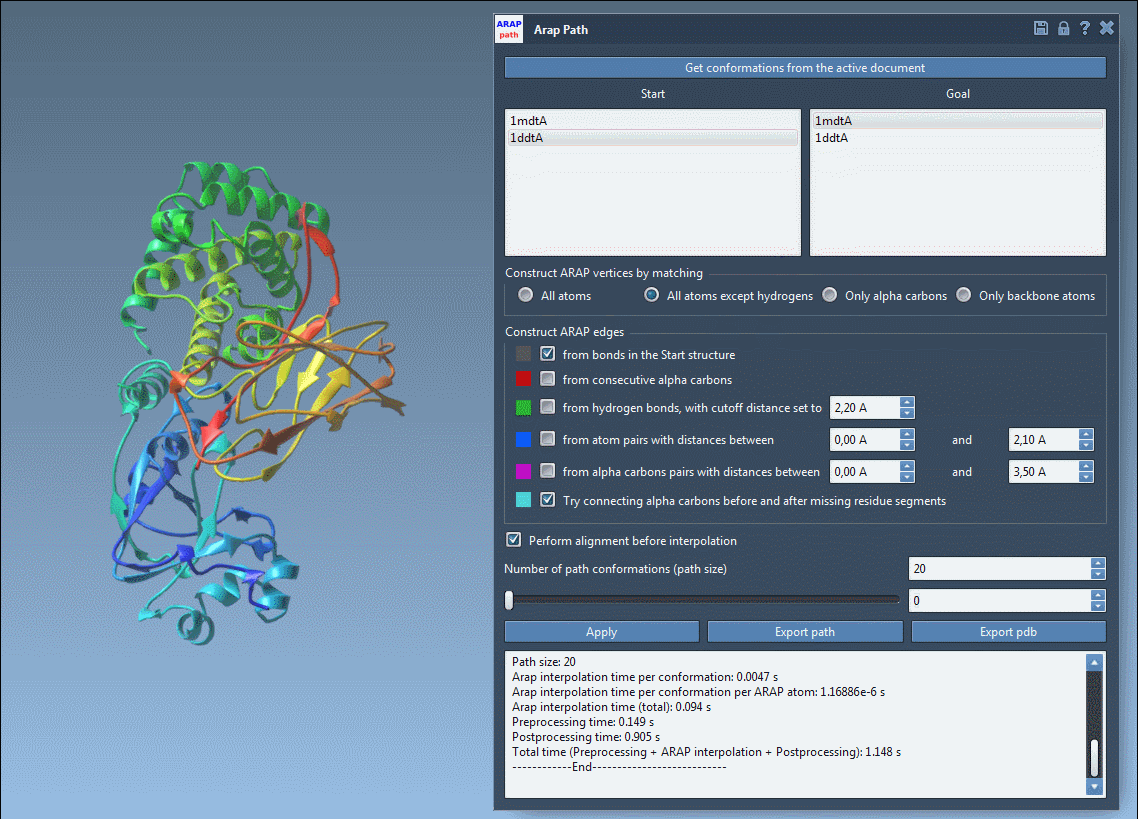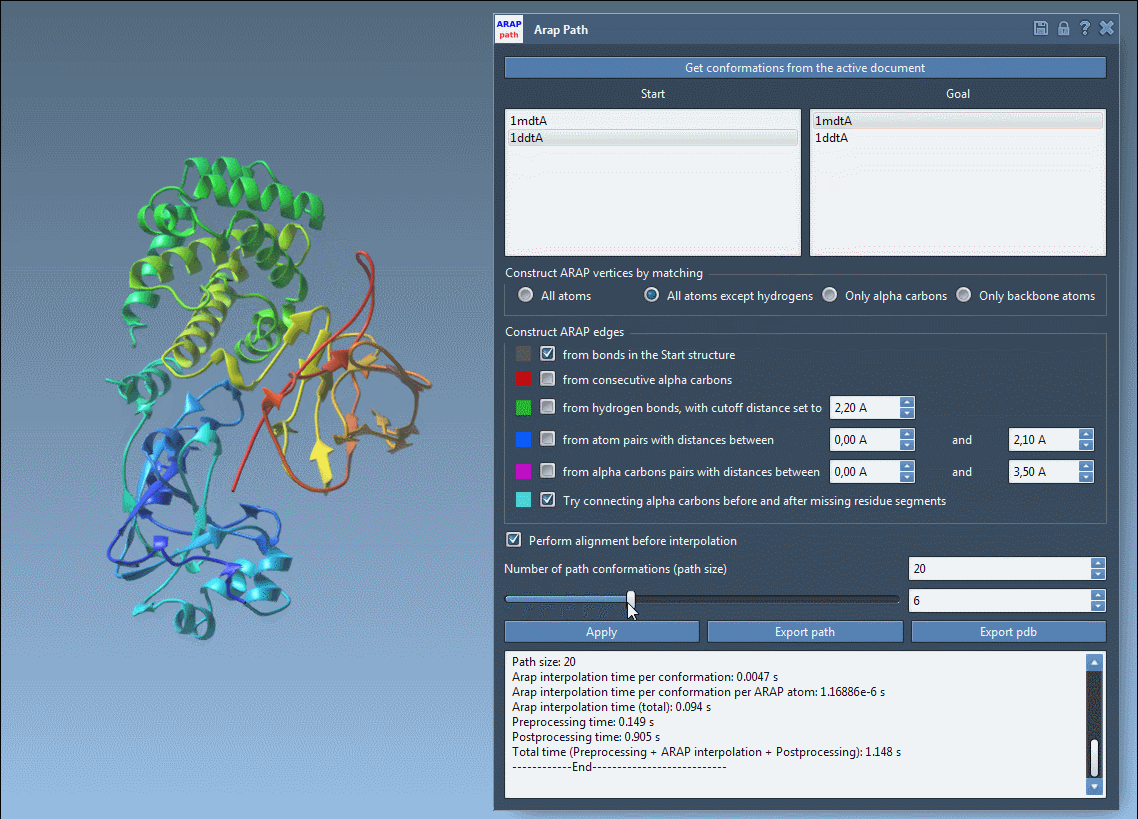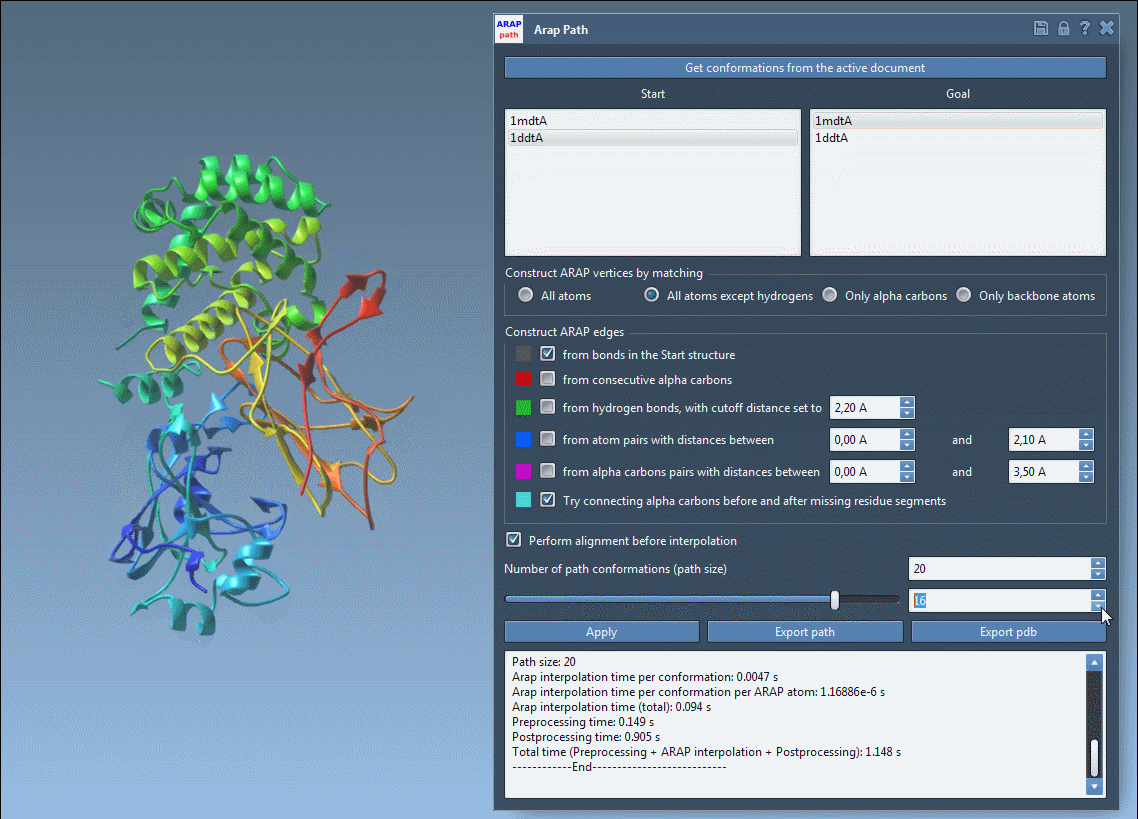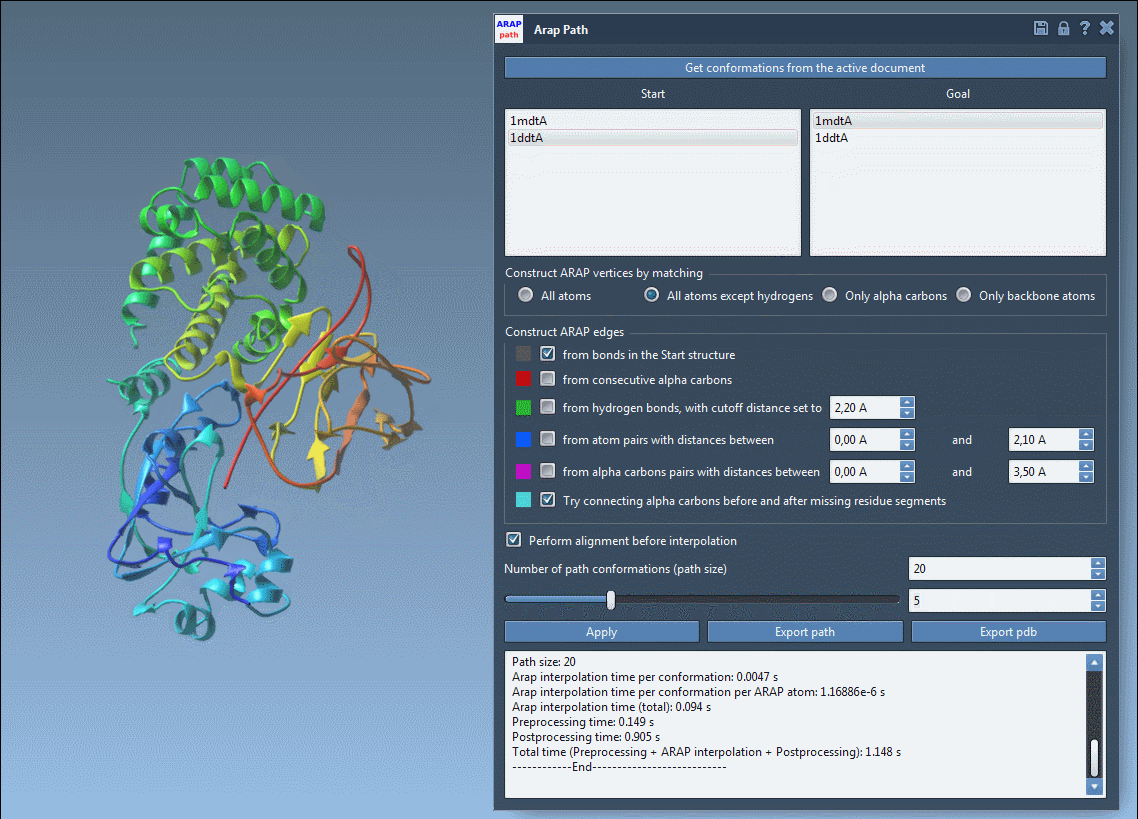
- Visualize the transition path using an interactive slider in the app.
- Analyze constructions like edge connections through intuitive visual models.
- Export the path as a trajectory object or as PDB files for each conformation.
Applications and Next Steps
Your ARAP-generated transition paths can be directly used as inputs for simulation techniques, including:
- Umbrella sampling and steered MD simulations.
- Refinement with parallel nudged elastic band (P-NEB).
- Visualization or dimensionality reduction of structural ensembles.
If you encounter any issues, remember to clean both structures beforehand using Home > Prepare and review the customizable options in the ARAP interpolation app settings.
For a more detailed tutorial, visit the official documentation page here.
Note: SAMSON and all SAMSON Extensions are free for non-commercial use. You can download SAMSON at SAMSON Connect.





