One of the key challenges for molecular modelers is understanding the motion of specific components, such as ligands, residues, or domains, within a complex system. These motions can reveal critical insights into mechanisms like ligand unbinding, protein conformational changes, or diffusion, but visualizing them has historically been a cumbersome task.
Here’s how the Pathlines feature in SAMSON makes this process much more intuitive, allowing you to track the center of mass (COM) motion of selected atoms along specified paths. This guide will walk you through the streamlined capabilities offered by the Pathlines extension, designed to address this modeling challenge.
What Are Pathlines?
Pathlines are visual models that represent the trajectory of the center of mass (COM) of a group of atoms as they move along a path. This feature is particularly useful for:
- Tracking the motion of ligands, molecular residues, or atomic groups over time.
- Visualizing collective domain motions in protein assemblies.
- Evaluating diffusion, unbinding, or domain shift mechanisms in complex systems.
Quick Start: Create Pathlines for COM Motion
If you’re new to Pathlines, creating one is surprisingly intuitive. Here’s a step-by-step guide:
Step 1: Load the Sample System
Begin by loading the sample system provided with the Pathlines tutorial. This example includes Lactose Permease (PDB ID: 1PV7) and its ligand Thiodigalactoside (TDG):
- Navigate to Home > Download in SAMSON.
- Paste the document link provided: https://www.samson-connect.net/documents/046f1acd-c799-40f6-8185-cb4847eff795.
- Click Download.

This step will load a document with a protein-ligand system and pre-calculated unbinding paths, making it easy to begin exploring Pathlines immediately.
Step 2: Select Atoms and Paths
To create a meaningful Pathline, you’ll need to select the atoms and paths you wish to analyze:
- In the Document view, select a group of atoms, such as a ligand, for which you want to visualize their center of mass motion. You can hold Ctrl (or Cmd on Mac) to select multiple elements.
- Choose one or more paths from the options in the document. If no paths are selected, all paths will be used by default.

Step 3: Generate the Pathline Visual Model
Create the Pathline to visualize the motion:
- Go to Visualization > Visual model > More… or use the shortcut (Ctrl/Cmd + Shift + V).
- Choose Pathline of the center of mass from the dialog and click OK.
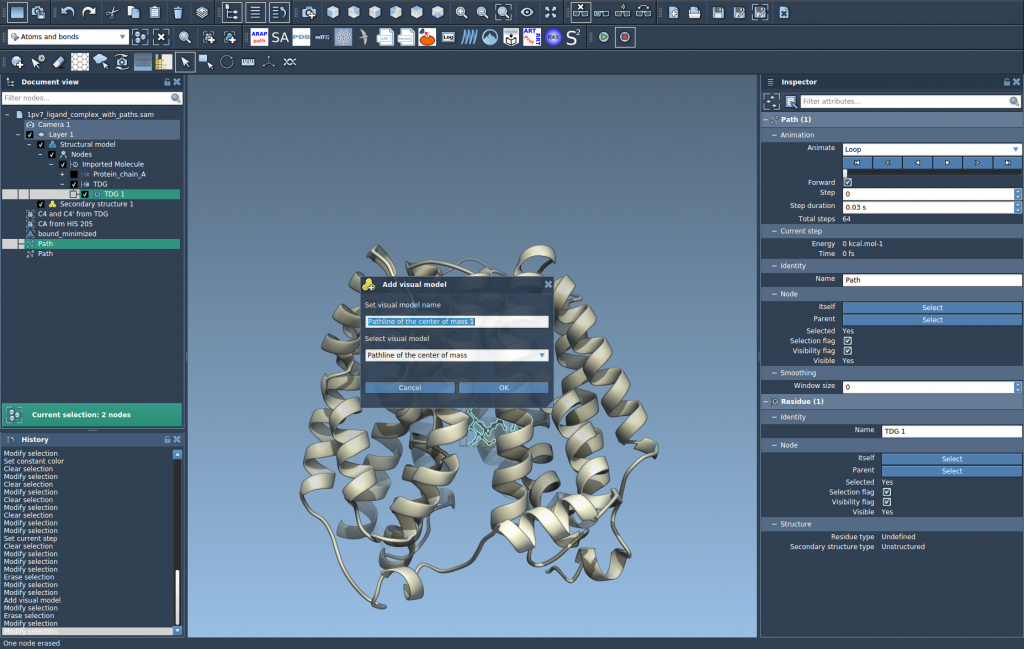
The visual Pathline will now display the center-of-mass motion of the selected atomic group along the defined paths. This enables detailed analysis of molecular mechanics.
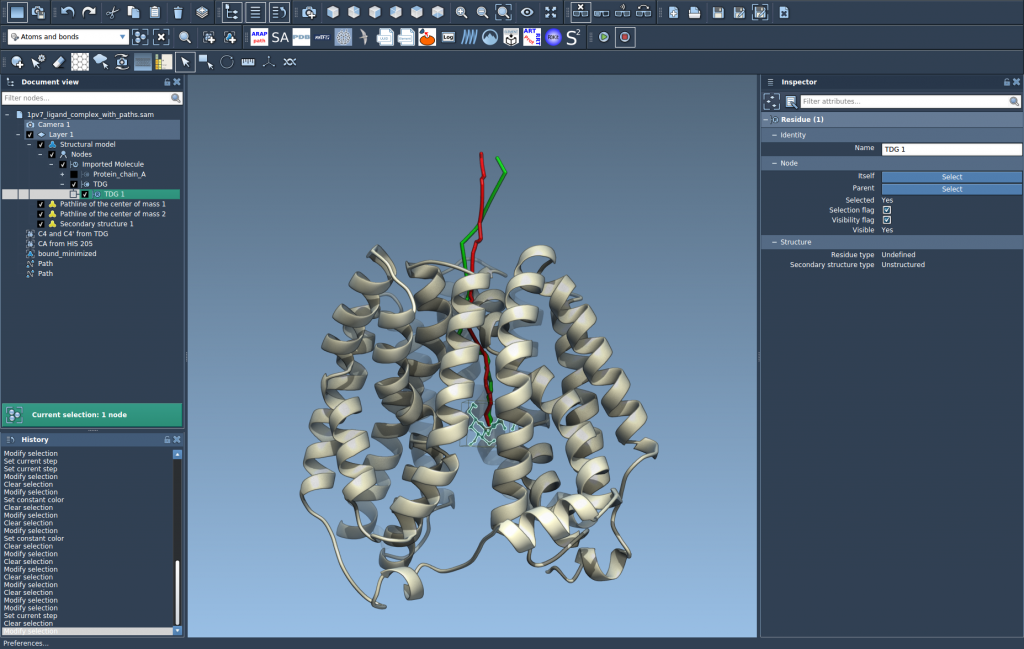
Step 4: Customize and Explore
You can now further refine and interact with your Pathline:
- Double-click on a path in the Document view to start or stop it.
- Access additional options by right-clicking on the path.
- Inspect and adjust properties like color and thickness through the Inspector (Ctrl/Cmd + 2).
Why Choose Pathlines?
By enabling a clear visualization of center-of-mass movement, Pathlines simplify complex analyses. Be it ligand unbinding routes or protein domain adjustments, the tool adapts seamlessly to your use case. For those looking to delve deeper, you can complement this with other SAMSON features like the Parallel Nudged Elastic Band (P-NEB) method to optimize paths further.
Ready to streamline your molecular motion analysis? Visit the full documentation page at Pathlines tutorial to learn more.
Note: SAMSON and all SAMSON Extensions are free for non-commercial use. You can get SAMSON at https://www.samson-connect.net.





