For molecular modelers, exploring the chemical space of potential drug candidates or functional materials efficiently is a common challenge. Positional Analogue Scanning (PAS), a method for generating families of molecular analogues by modifying specific parts of a structure, can be a highly useful strategy for such tasks. Within SAMSON, an integrative molecular design platform, you can perform PAS seamlessly using the SMILES Manager extension. Here’s a step-by-step guide on how to accomplish this to explore molecular substitutions and prepare for docking or interaction analysis.
Why Positional Analogue Scanning?
In molecular design, small changes in a molecule’s structure can dramatically alter its properties and interactions. With PAS, you can replace specific groups or atoms in a molecule systematically, enabling you to generate a series of analogues quickly. This process aids in comparing analogue properties, refining docking studies, and identifying how substitutions might affect functionality or binding efficiency. Let’s dive into how this works in SAMSON.
Defining Your Starting Molecule
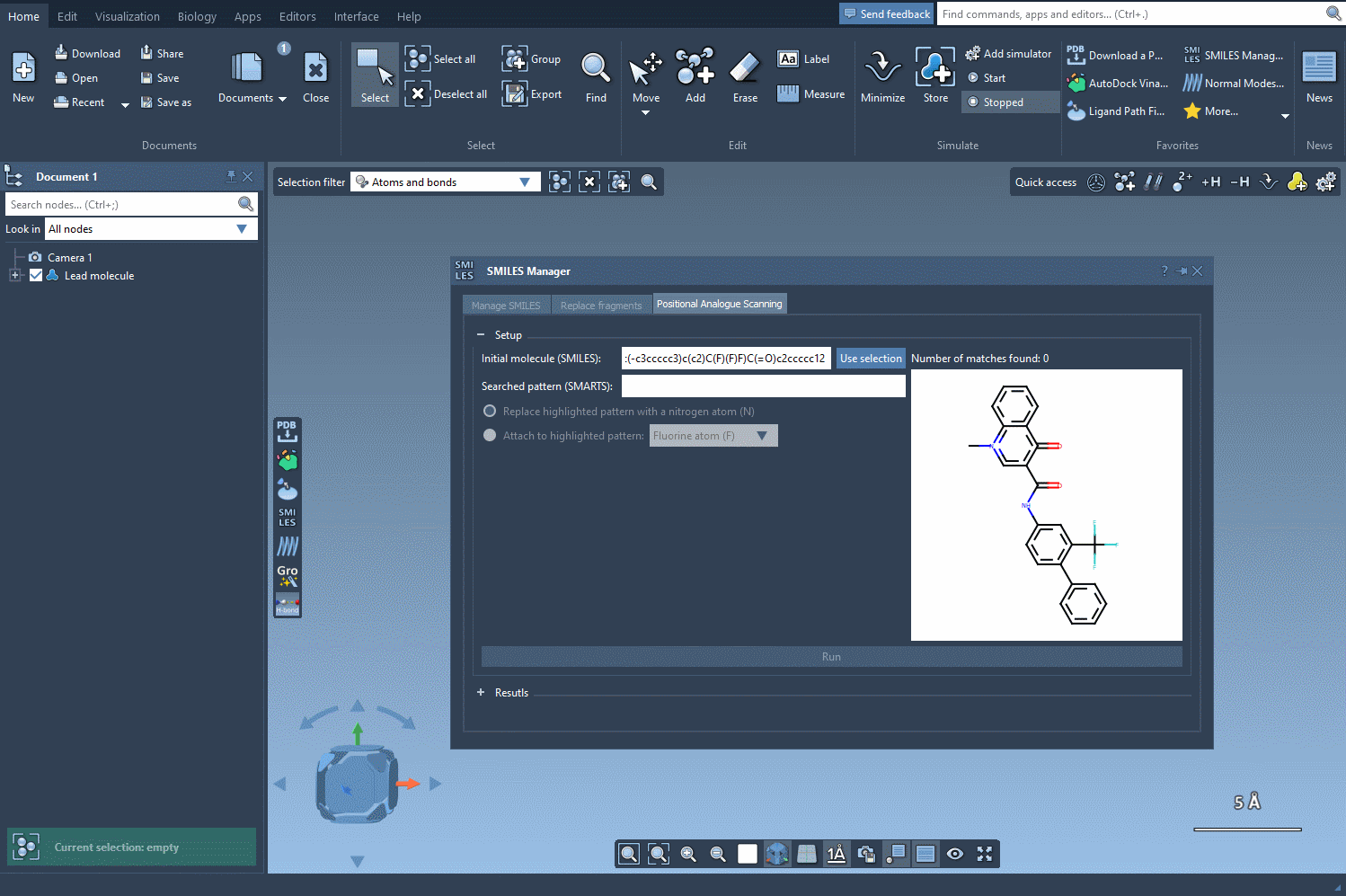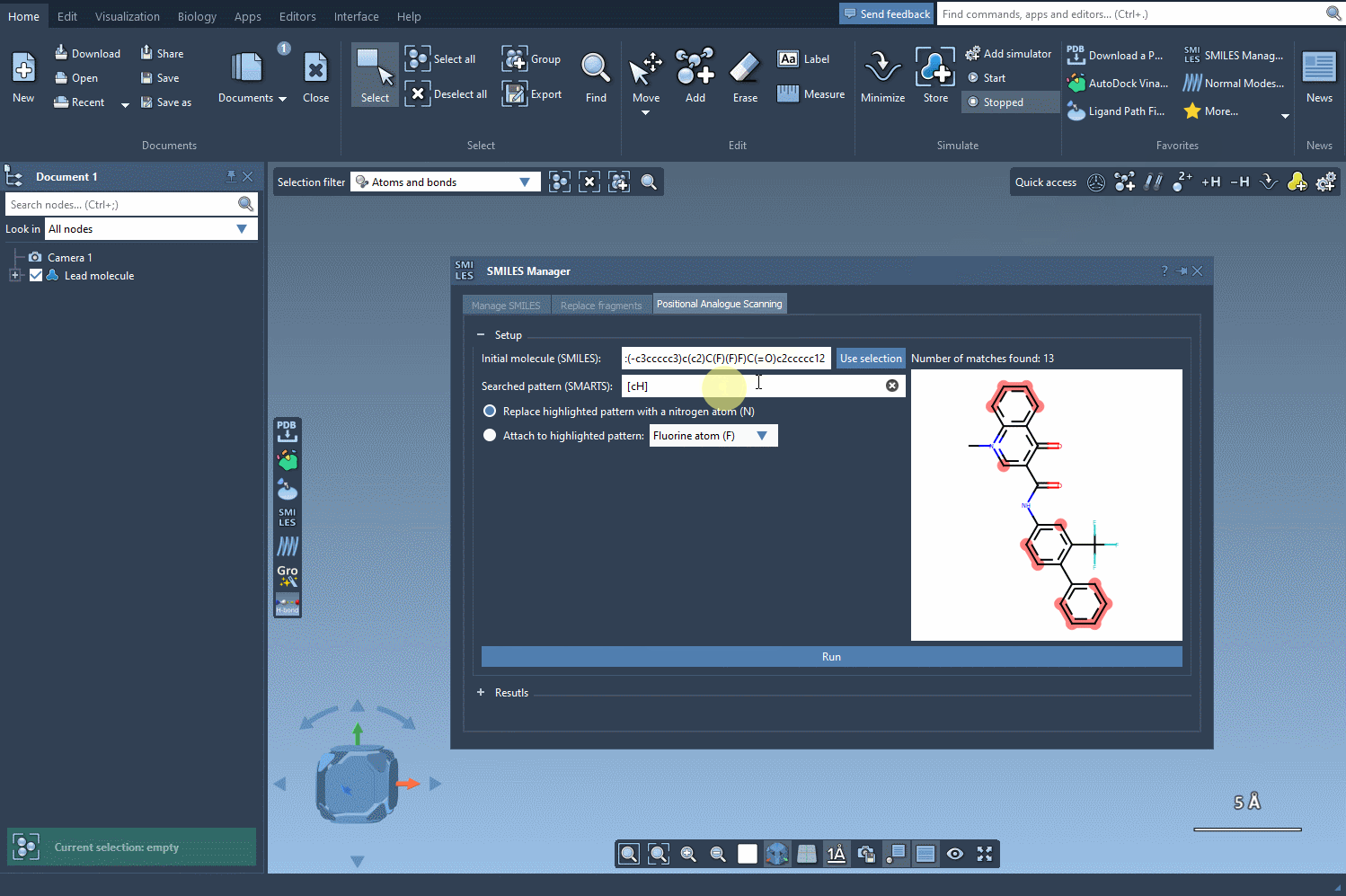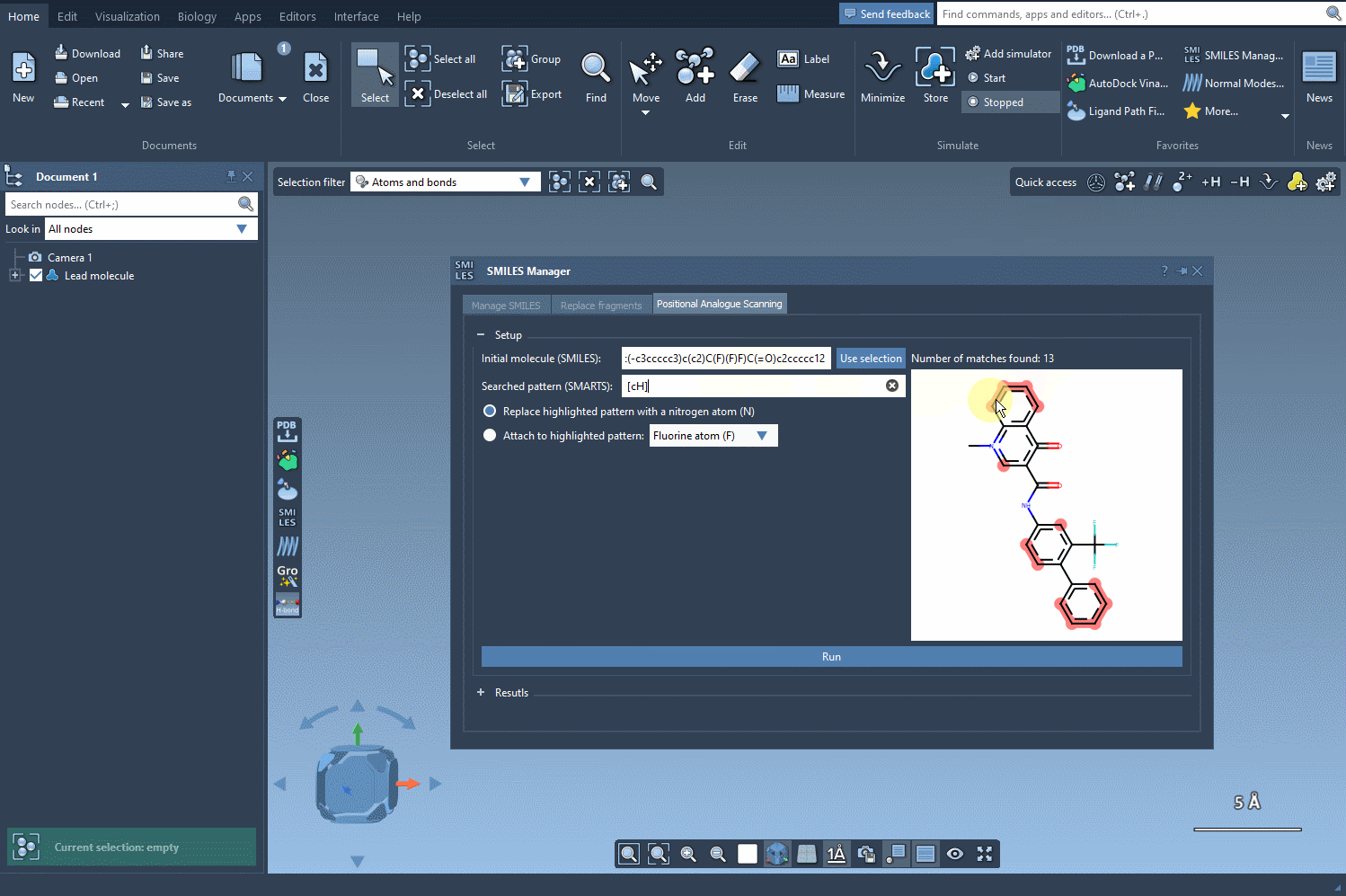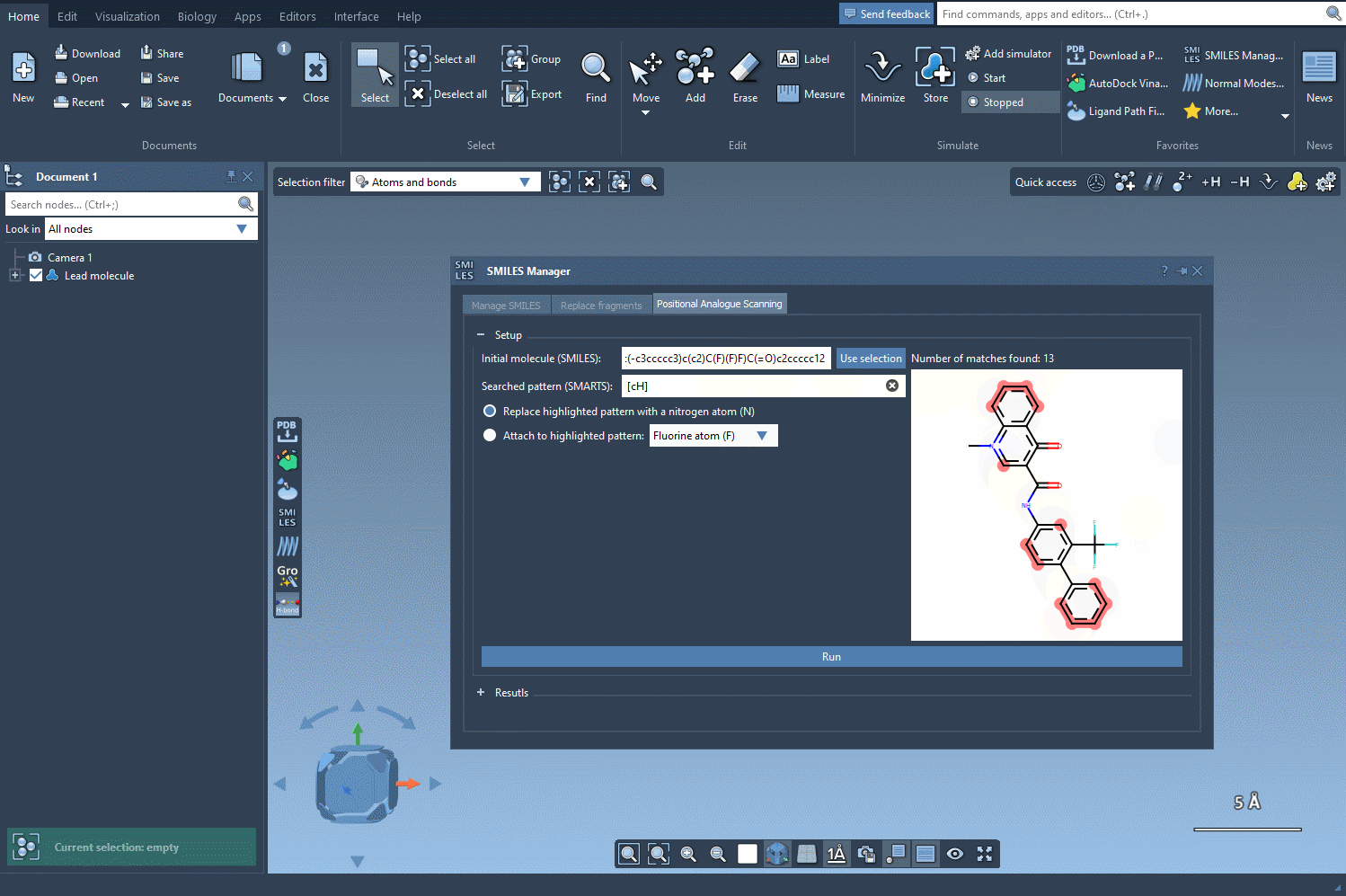
The first step in PAS is choosing a starting molecule. In the SMILES Manager interface, you can either enter the molecule’s SMILES code directly or select it from your SAMSON workspace. Once selected, click the Use selection button in the interface. With just a few clicks, you’re ready to move to the next step.
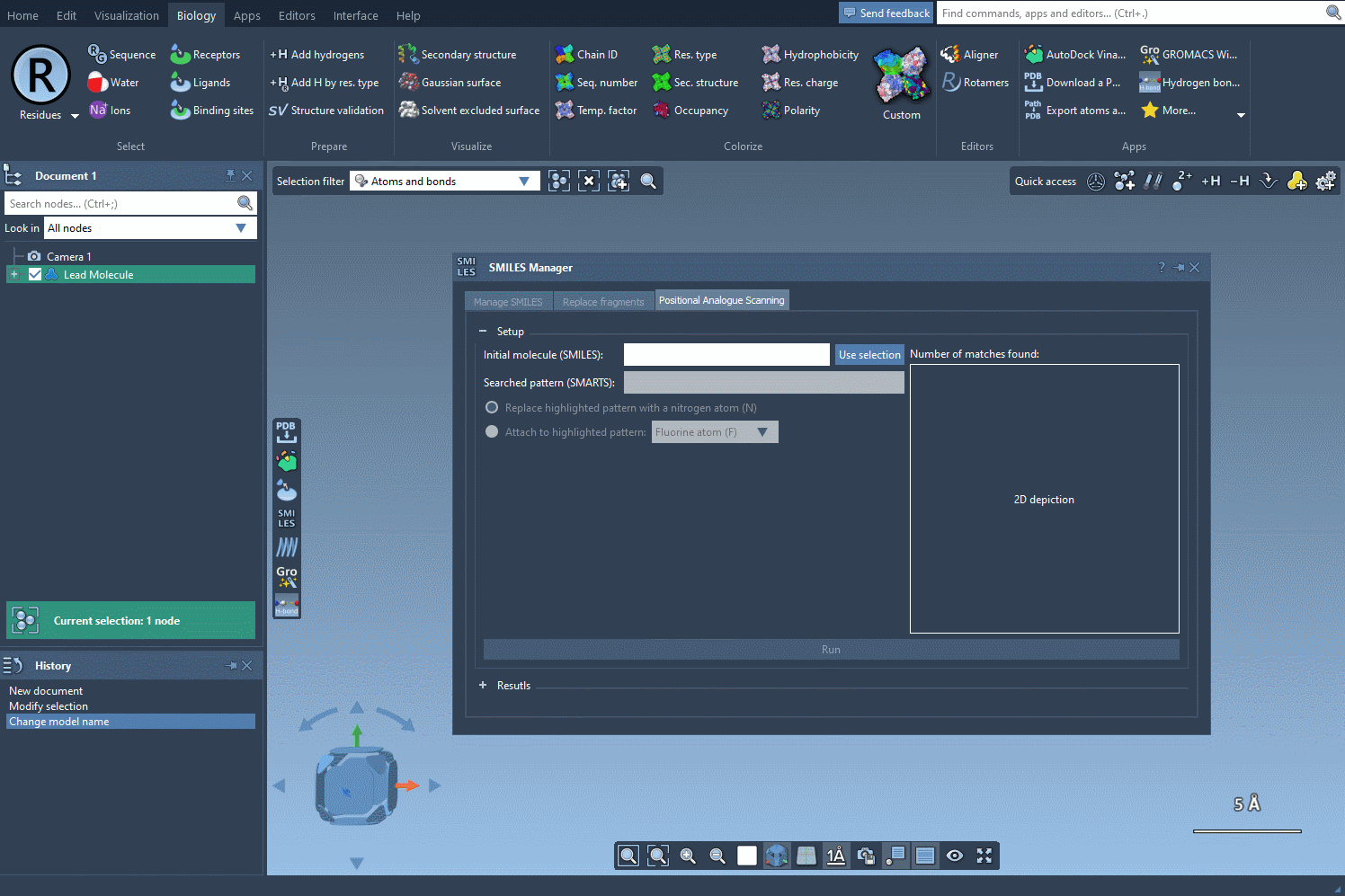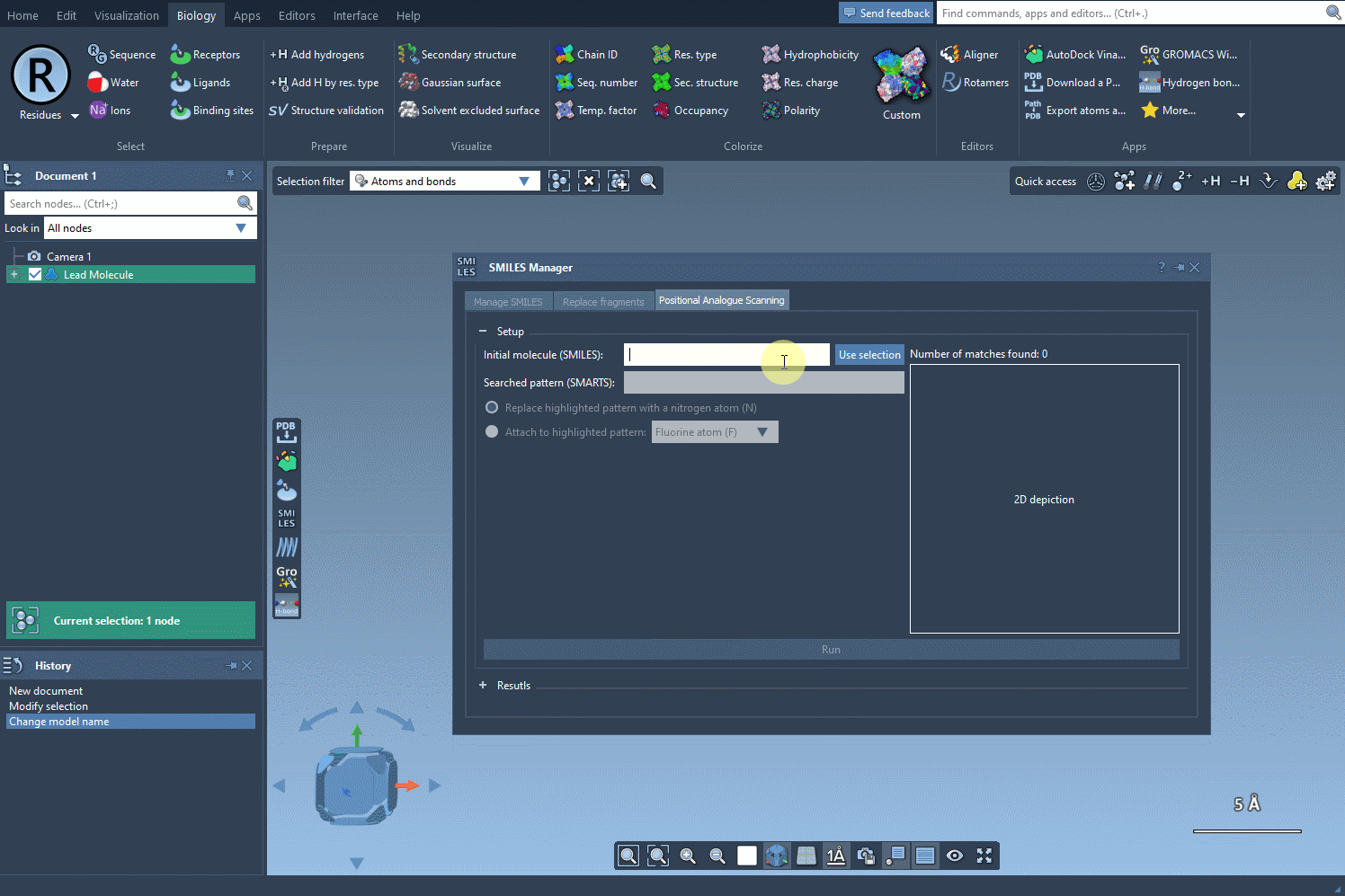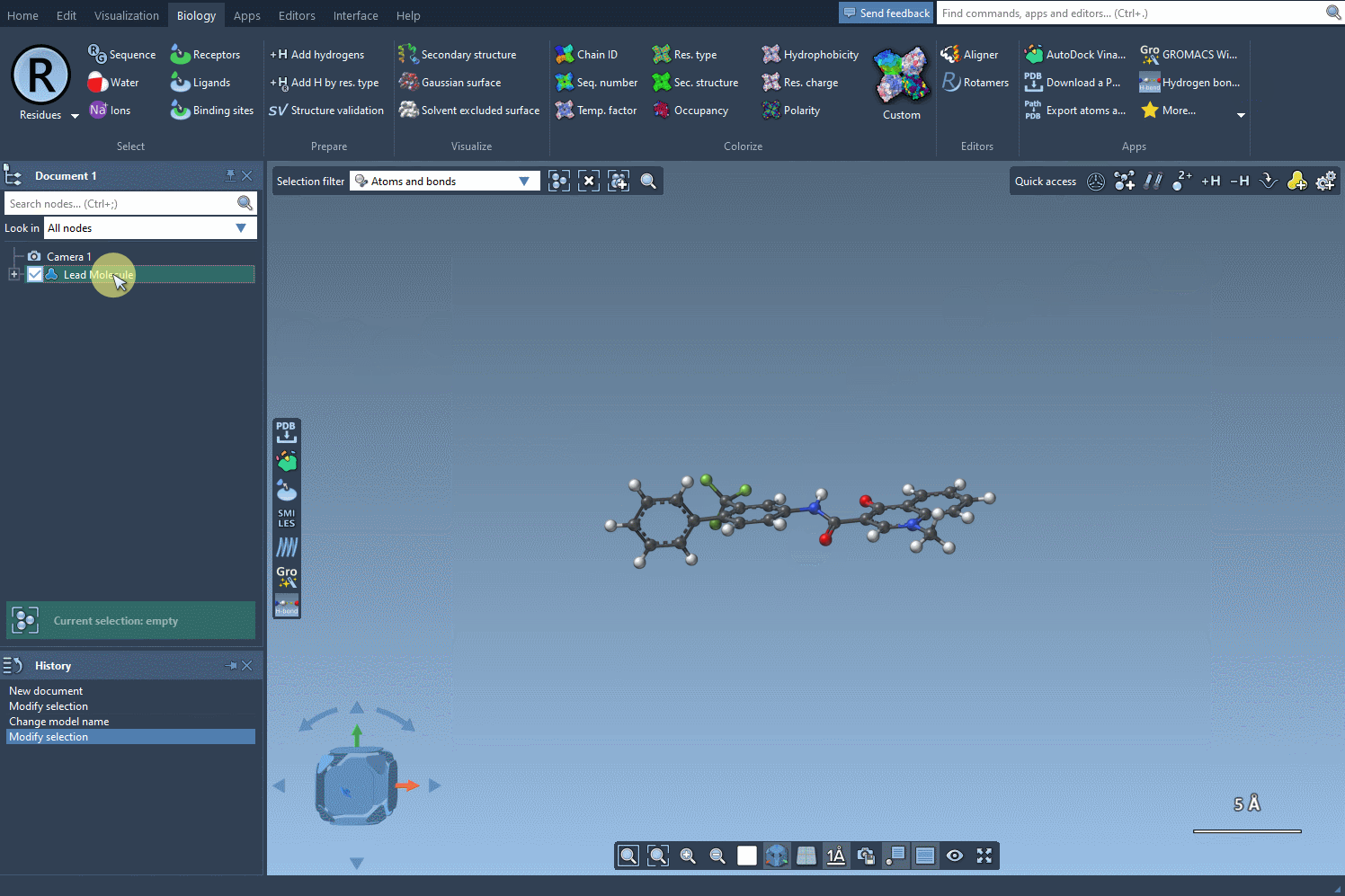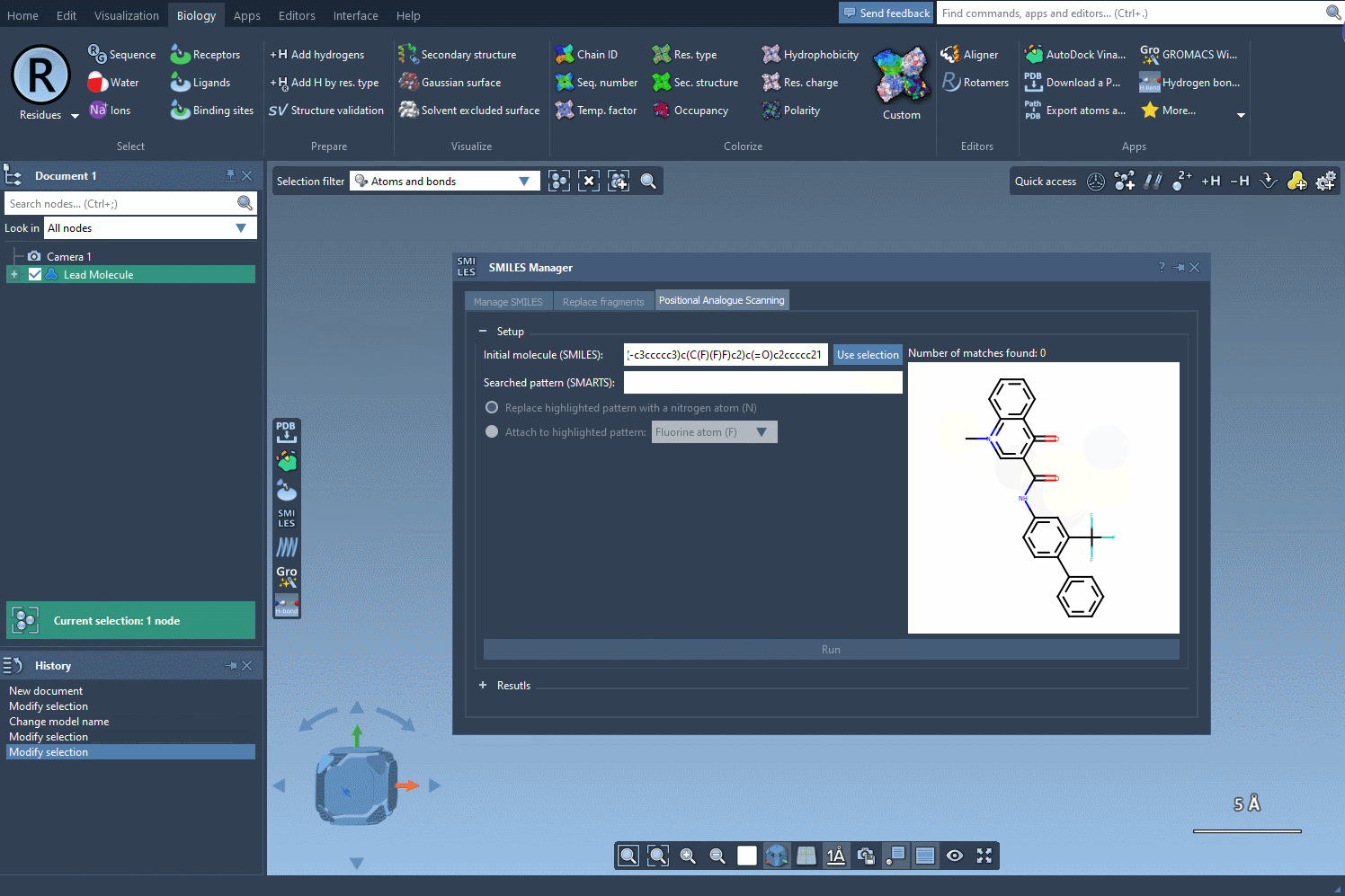
Finding and Defining the Pattern to Replace
After specifying your starting molecule, it’s time to identify which part of the molecule you want to alter. In PAS, this is often done using SMARTS, a syntax for identifying patterns in molecular structures. For example, you can search for all aromatic carbons in a molecule by entering the [cH] SMARTS pattern. The interface will highlight all occurrences of this pattern directly in your molecule.
Customizing Substitutions
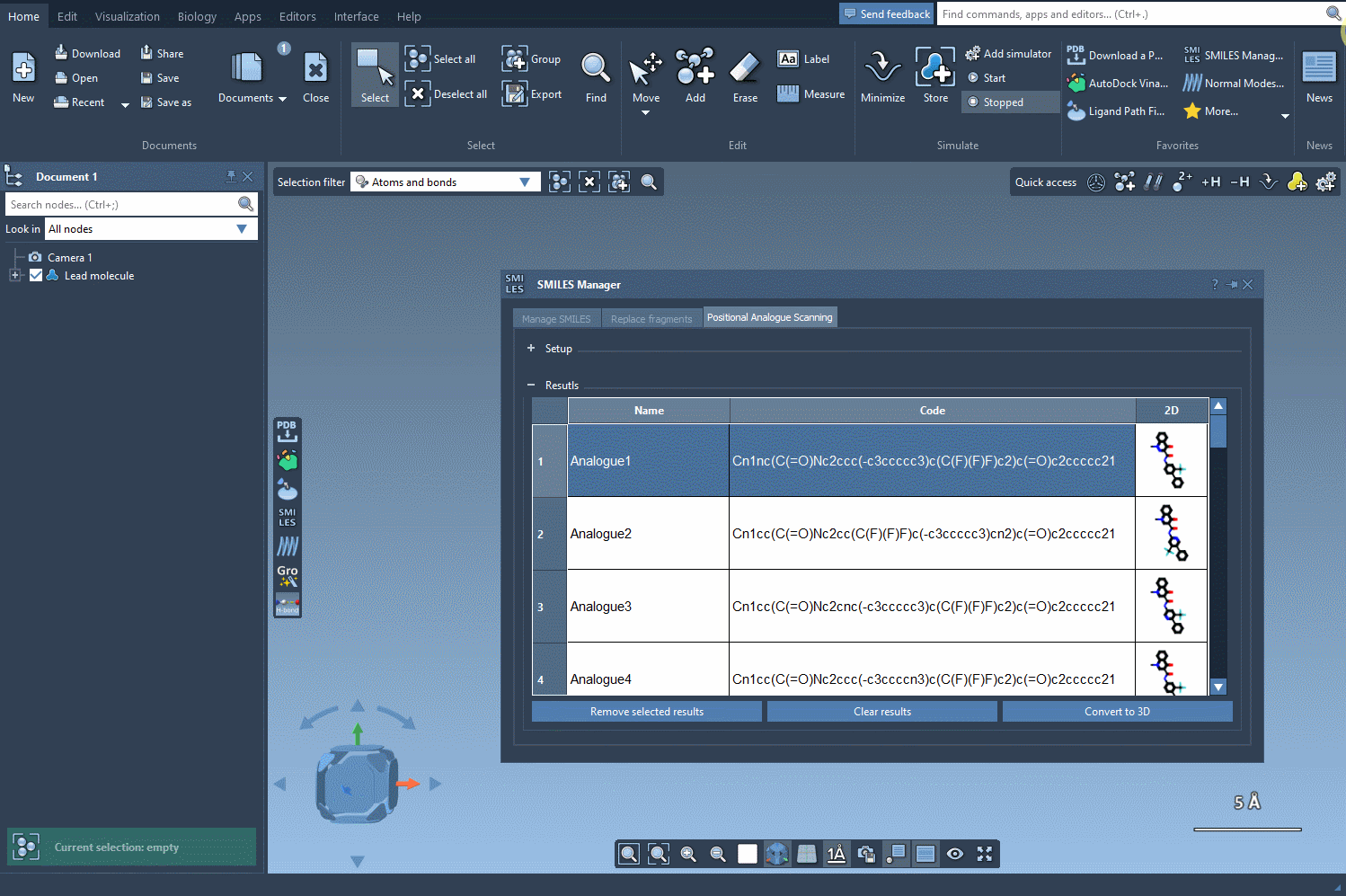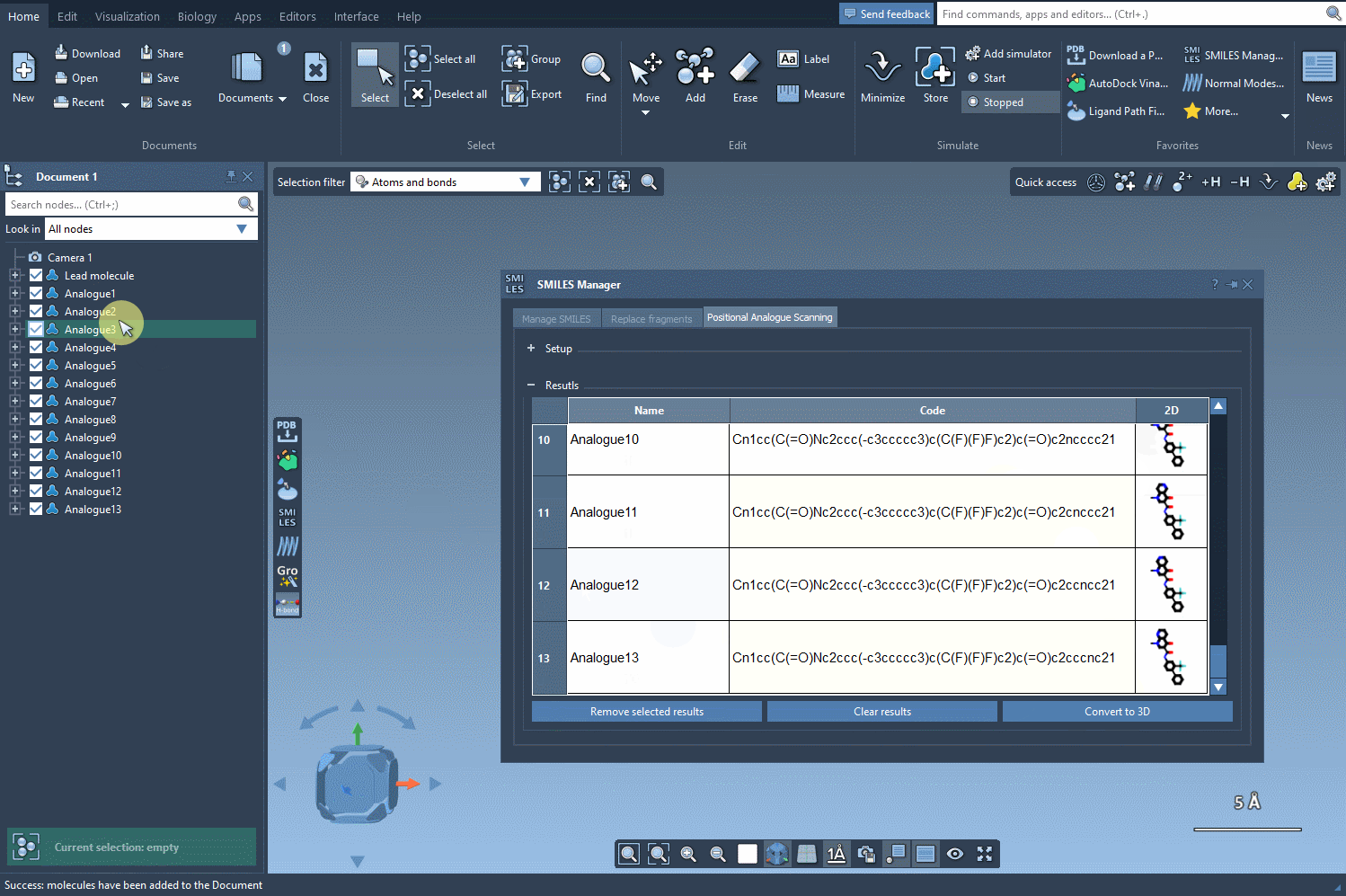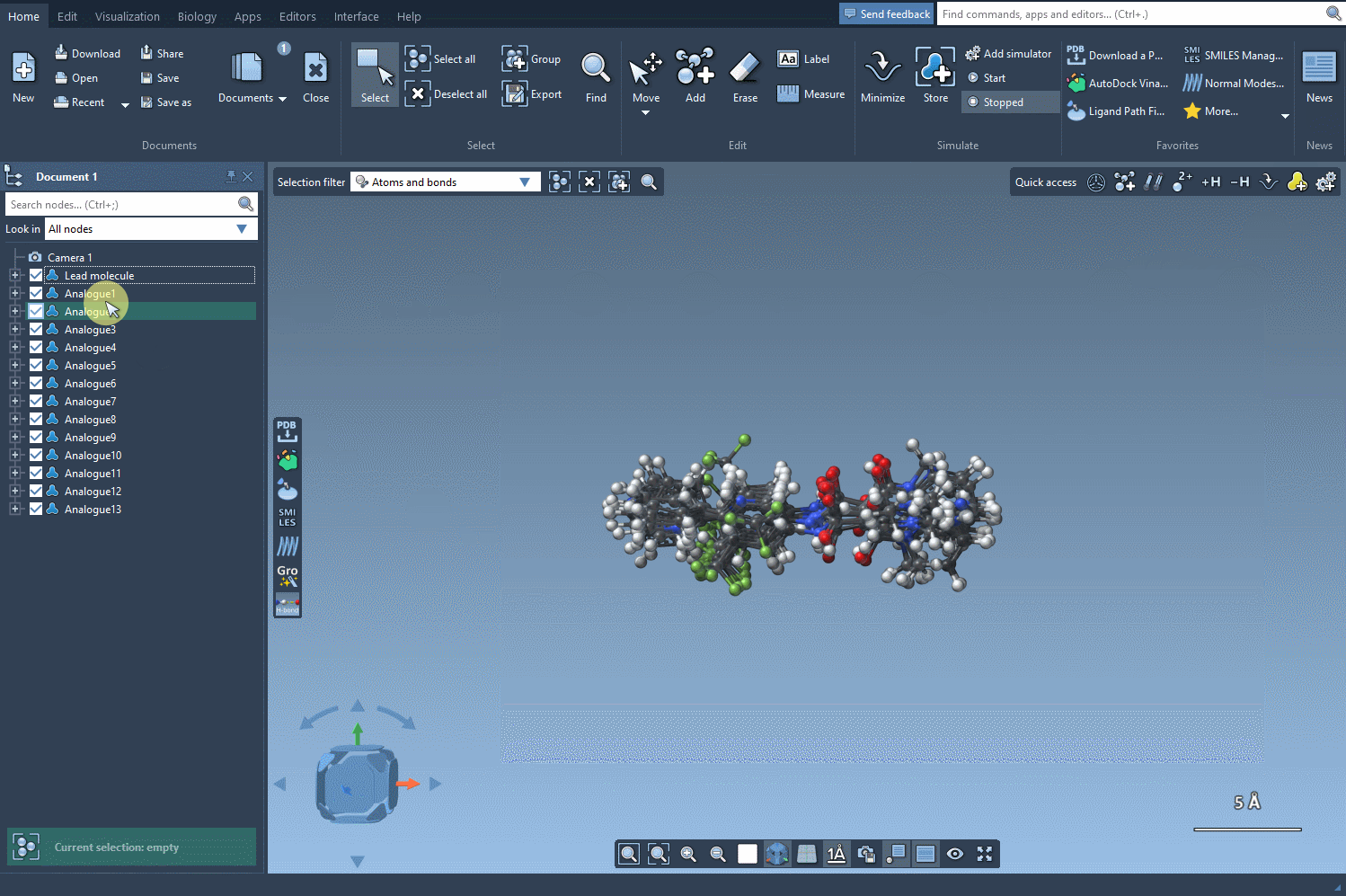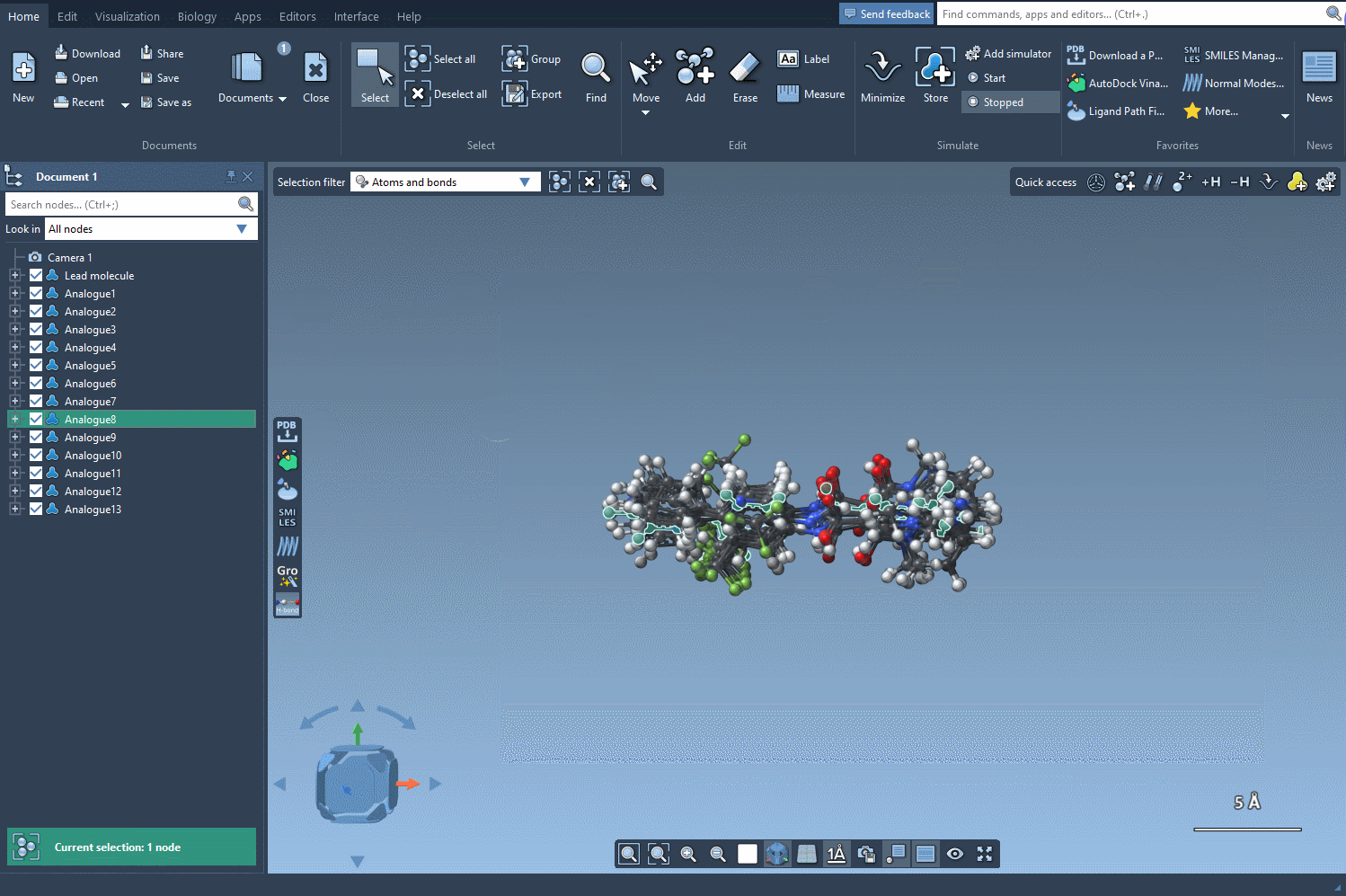
Once you’ve defined the pattern, you can specify what modifications to make. Replace a highlighted pattern with a nitrogen atom (N), attach a new group such as fluorine (F) or methyl (CH3), or test other substitutions. To execute the PAS process, simply click Run. SAMSON will then generate the analogues, displaying them as a list complete with their SMILES codes and 2D depictions.
Going Further: Analyzing and Refining Results
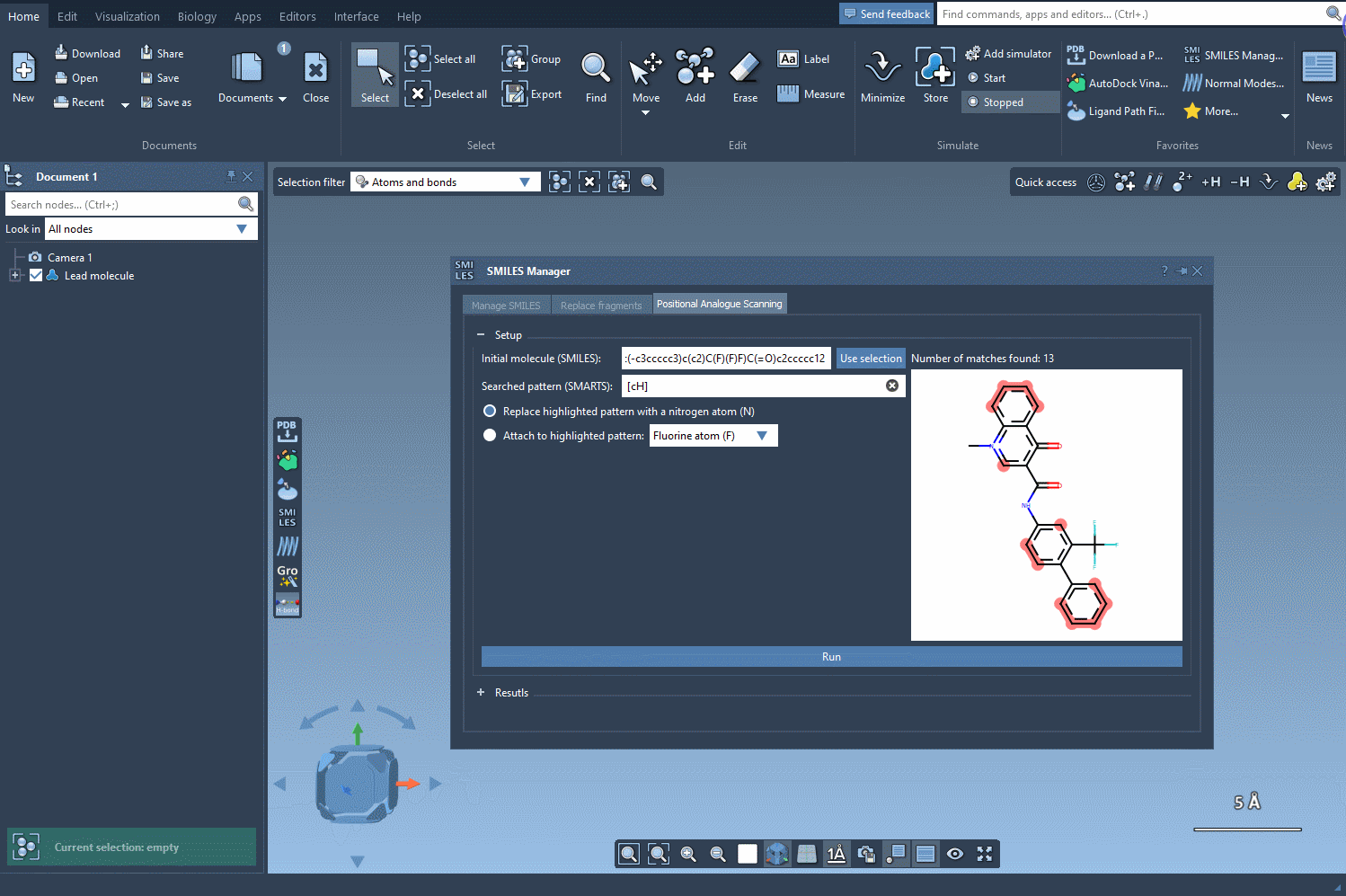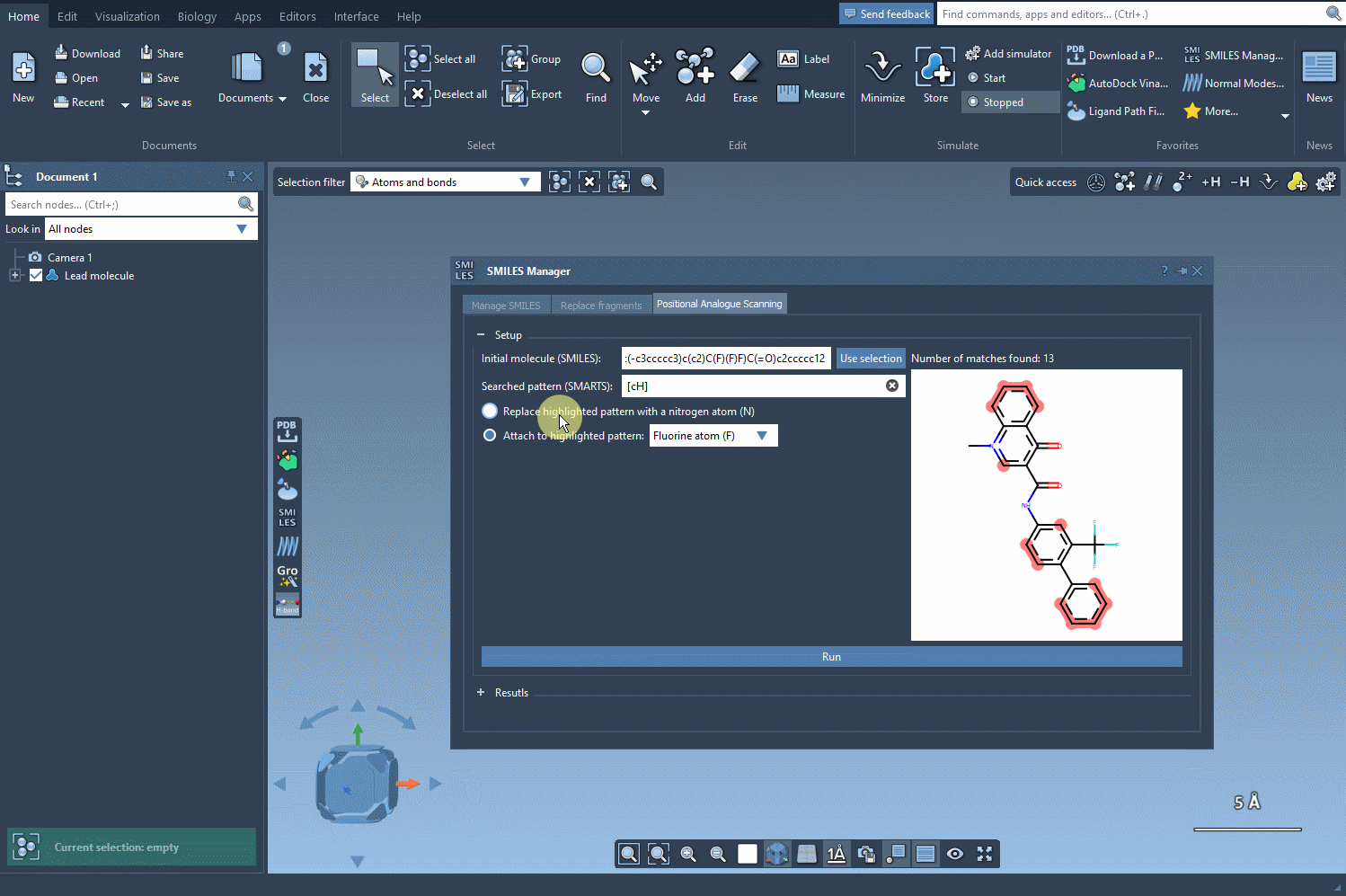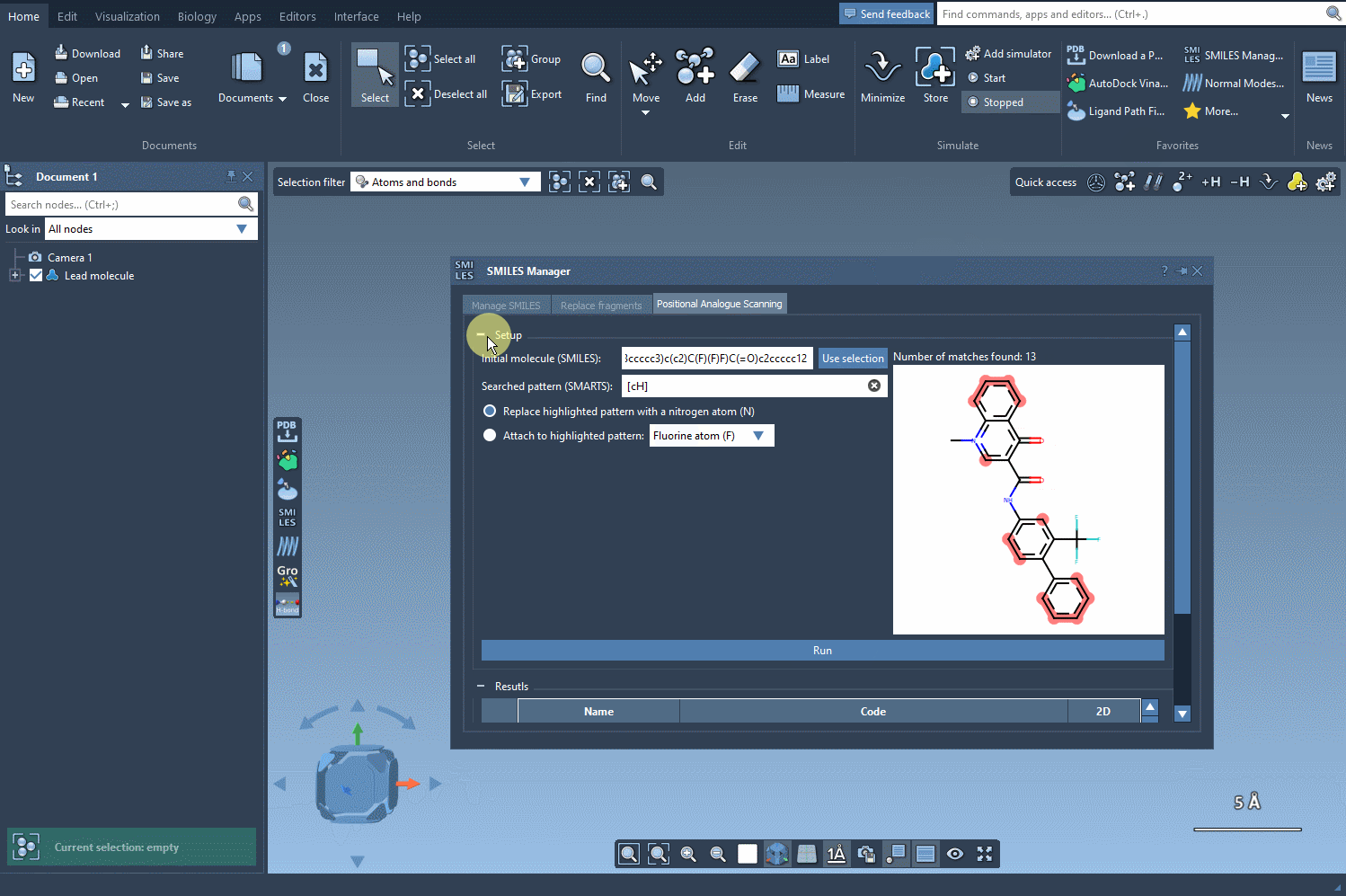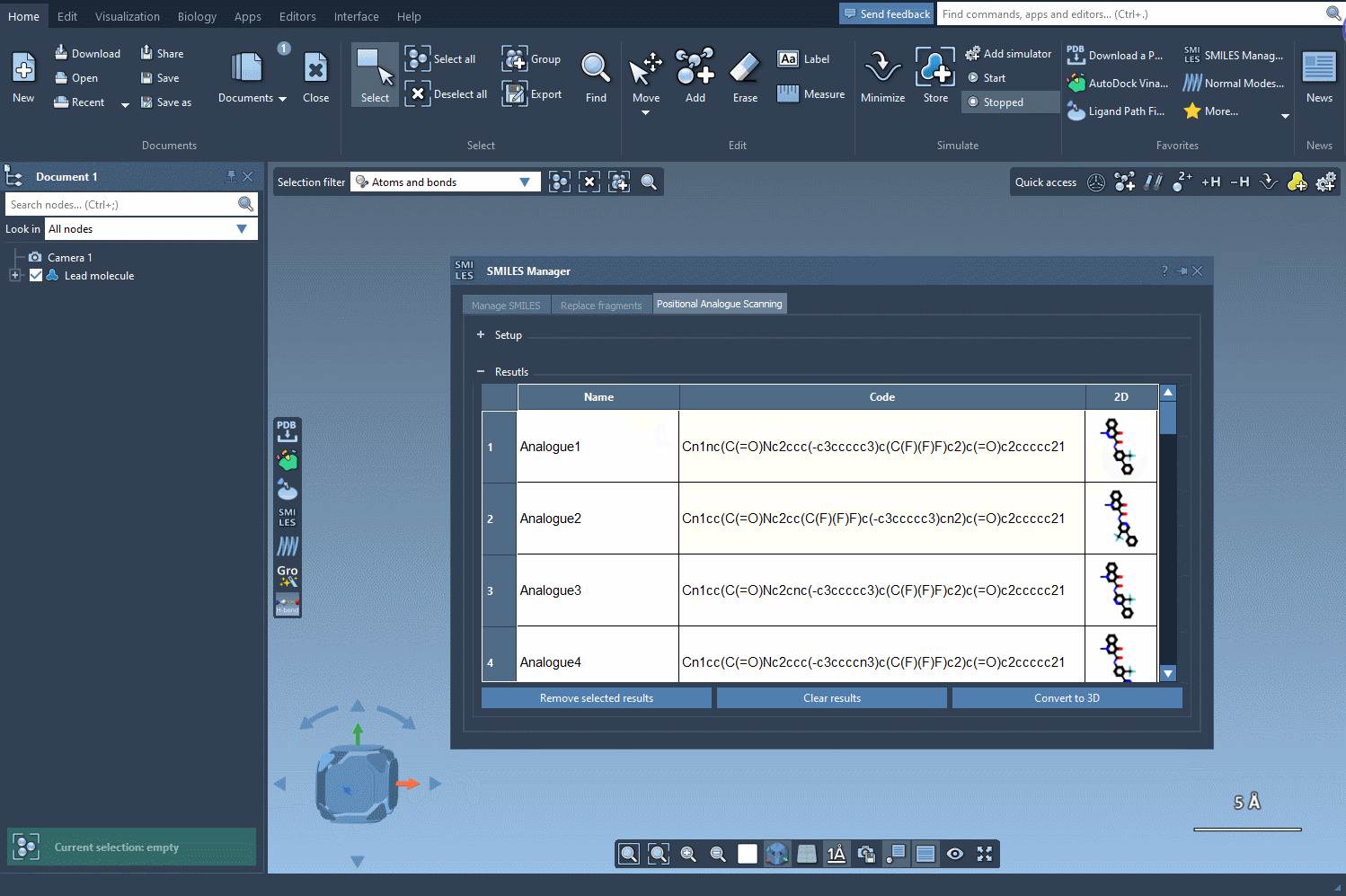
The results table in the SMILES Manager enables you to organize and refine your newly generated analogues efficiently:
- Modify analogue SMILES: Change SMILES codes or names directly by double-clicking on table cells.
- Open or hide 2D structures: View larger versions of 2D images or toggle their visibility.
- Generate 3D structures: Right-click a molecule’s image and select Convert to 3D for further studies.
Bringing 3D Into the Picture
After generating a series of promising analogues, your next step might involve converting them into 3D structures. Clicking on the Convert to 3D button prepares them for use in docking tools such as AutoDock Vina Extended, available as another SAMSON extension. This workflow allows you to evaluate which chemical modifications enhance interactions with target proteins and which are less optimal.
Conclusion
Positional Analogue Scanning in SAMSON’s SMILES Manager empowers researchers to efficiently generate and analyze analogue series. Whether you’re screening for a specific property, preparing molecules for docking, or exploring a target’s interaction space, this tool streamlines vital modeling workflows. To learn more, visit the full tutorial on the SAMSON documentation platform.
Note: SAMSON and all SAMSON Extensions are free for non-commercial use. Download SAMSON at https://www.samson-connect.net.





