If you’ve ever faced challenges in building and editing molecular systems while preserving physical accuracy, the Interactive Modeling Universal Force Field (IM-UFF) could provide a helpful solution. IM-UFF is an extension of the Universal Force Field (UFF) that enables seamless topology modifications during molecular simulations. Here, we’ll explore how IM-UFF allows you to create, break, and modify bonds interactively, making molecular modeling more intuitive and dynamic.
Why Interactive Topology Handling Matters
Traditional molecular modeling tools often require static configurations, limiting your ability to experiment with structural changes mid-simulation. This becomes a bottleneck when working on systems that need iterative adjustments or involve topological changes, such as bond creation or breaking, changes in bond orders, or atomic retypizations.
IM-UFF addresses this pain point by introducing a dynamic, continuously adjustable interaction model. With this extension, you can simulate molecular systems while interactively altering their topology. It ensures a physically accurate representation of the system’s energy and forces throughout the process, allowing for smoother transitions between different configurations.
How It Works
To get started with IM-UFF, load a molecular system into SAMSON and add the simulator via Edit > Simulate > Add simulator. Select Interactive Modeling Universal Force Field from the interaction models list to activate this feature. Once set up, the parameter window gives you control over the model’s interactive options:
- Static topology (UFF only): This toggles between standard UFF behavior and the dynamic IM-UFF approach. In IM-UFF mode, bond energies shift so that non-interacting atoms are used as the zero energy reference, ensuring consistent comparisons with the original UFF implementation.
- Keep vdW for manipulated: When enabled, van der Waals energy and forces for all atoms, including manipulated ones, are calculated. Disabling it helps simplify manual manipulations by ignoring vdW interactions for atoms moved with the mouse.
As you manipulate atom positions, you’ll notice that bonds adjust dynamically. Moving an atom closer to others can form new bonds, while large displacements can break bonds, producing accurate topology updates in real time. This makes IM-UFF particularly useful for iterative molecular design and prototyping, where structures are frequently adjusted.
Understanding the Energy Feedback
During a simulation, the parameter window provides detailed insights into the system’s energies, from van der Waals interactions to total system energy. This allows for real-time tracking of structural stability and energy optimization while making interactive changes. The visual feedback, combined with energy data, provides actionable insights for informed decision-making during modeling tasks.
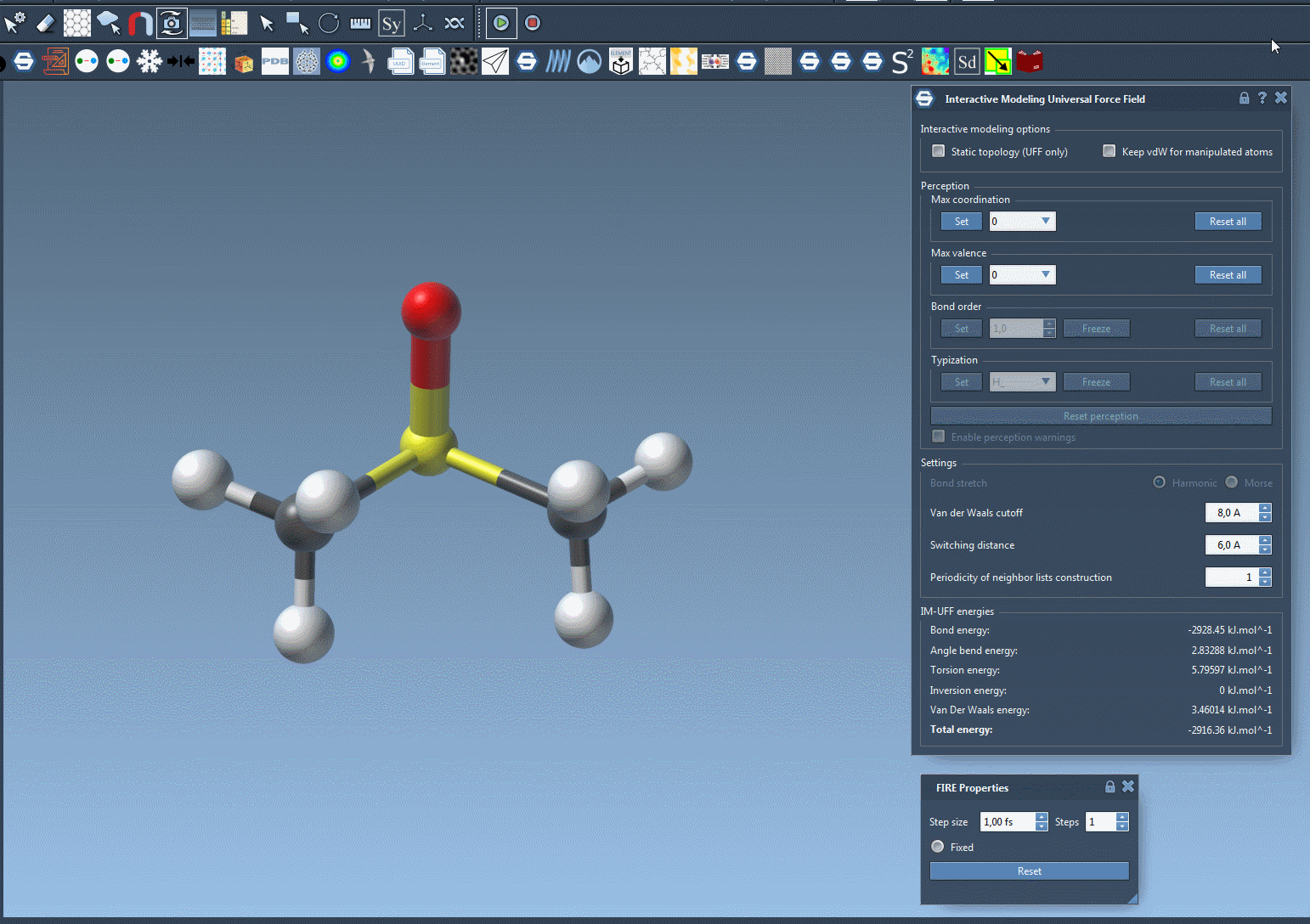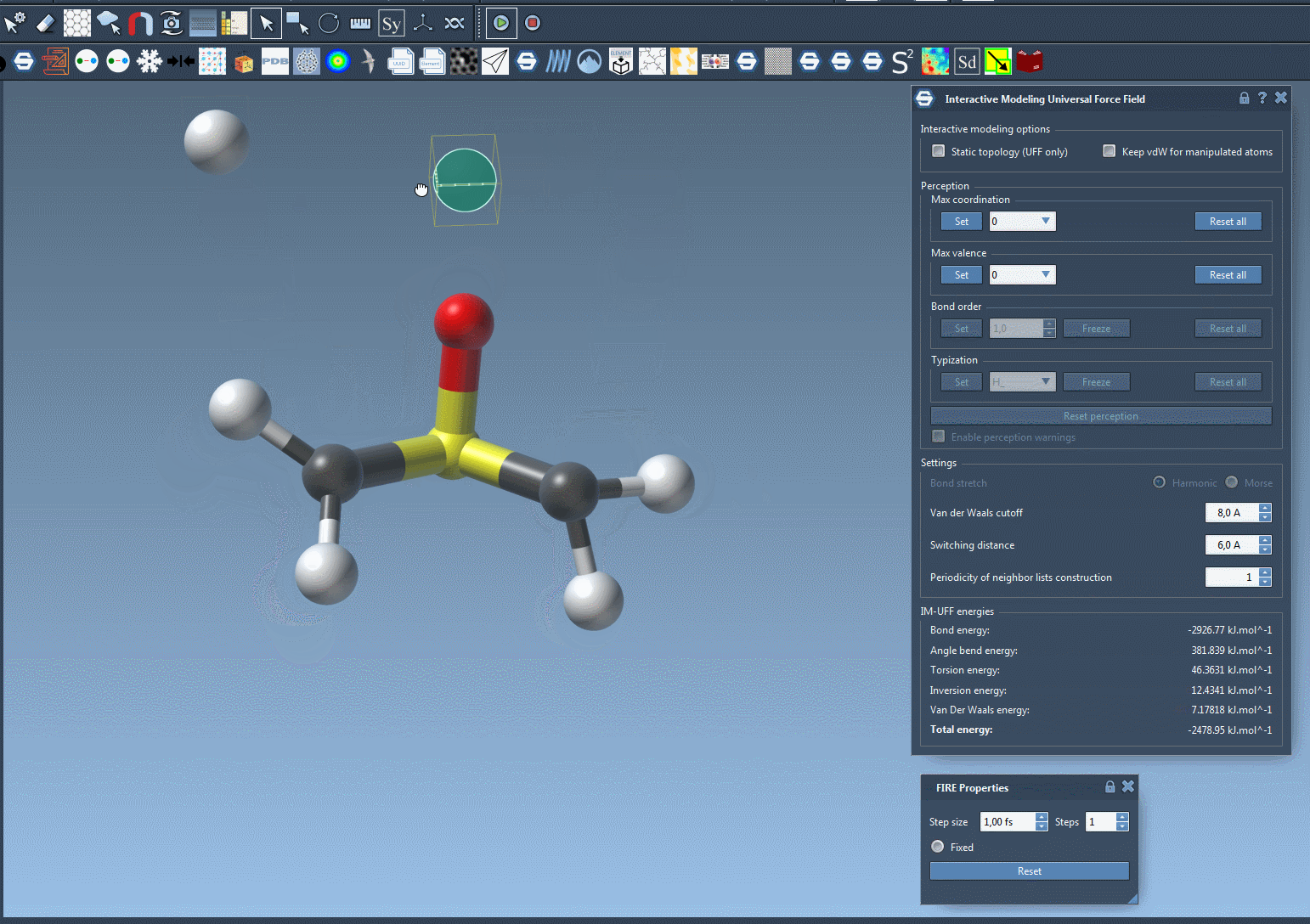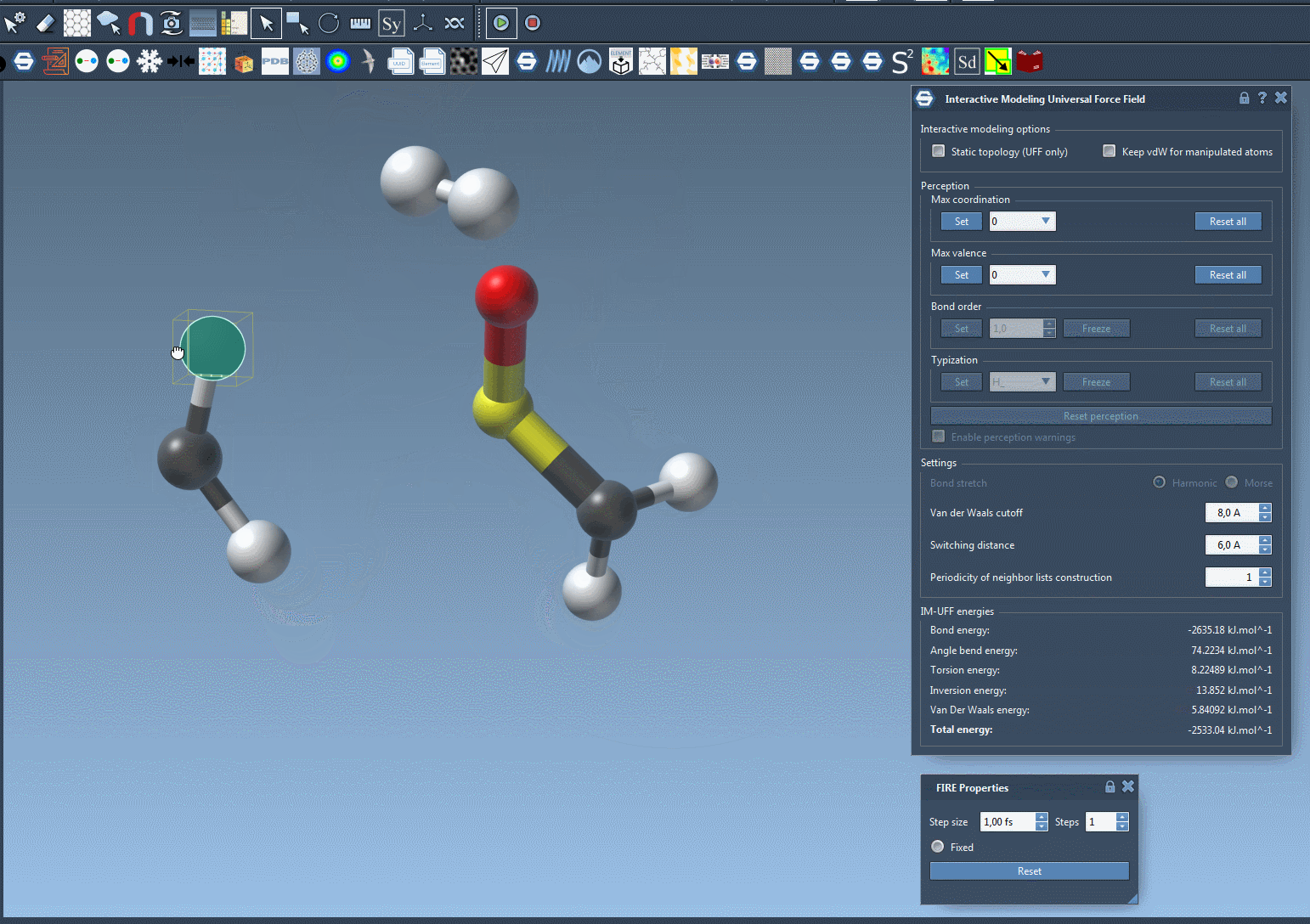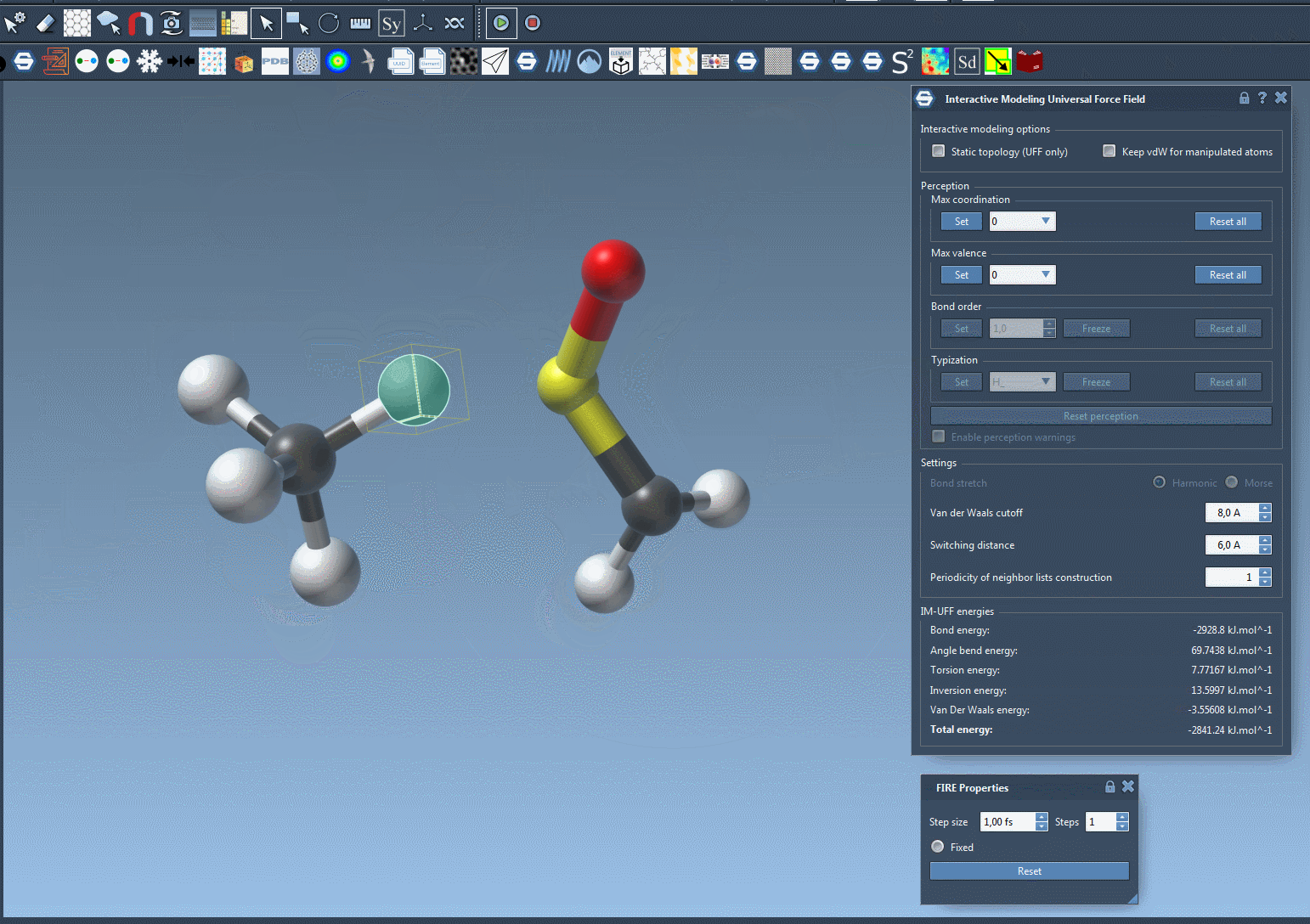
Customizing Parameters
IM-UFF also enables users to customize parameters such as van der Waals cutoff and switching distances, helpful for optimizing your simulations for specific systems. Additionally, the model handles smooth transitions in bond orders and atom typizations, allowing for continuous adjustments that reflect the physical environment of the atoms. These flexible settings reduce the complexity and provide precise control over the simulation’s behavior.
With features like these, IM-UFF is an essential tool for researchers and molecular designers aiming to refine complex structural models with interactive precision.
Want to Learn More?
For comprehensive details about IM-UFF and its configuration, head to the official documentation: Learn more about IM-UFF.
Note: SAMSON and all SAMSON Extensions are free for non-commercial use. To download SAMSON, visit SAMSON Connect.





