In molecular modeling, it’s not uncommon to end up with large amounts of trajectory data—often more than you need. Perhaps you’re studying just a ligand’s movement through a protein tunnel, or only the backbone motion during a conformational change. Exporting all atoms along a path doesn’t just create large datasets—it can make downstream analysis harder and slower.
If you’ve ever wanted to export trajectories for just a subset of atoms in SAMSON, there’s a simple tool that helps you do exactly that: the Export Along Paths extension.
Why Subset Export Matters
When preparing input files for free energy calculations, reaction coordinate constructions, or sampling intermediate states, simulation size matters. Exporting only what’s necessary—like just the ligand—reduces computation time, simplifies file parsing, and streamlines visualization in external software.
Export Along Paths – Subset mode makes this process intuitive and precise.
Step-by-Step: Exporting a Subset of Atoms
To use this feature, you’ll need the Export Along Paths extension installed in SAMSON. If you haven’t already, add it via Home > Apps > All then search for “Export Along Paths.”
1. Select Your Subset
In the Document view, locate and select a subset of atoms you’d like to export. For example, to follow a ligand like TDG, select it directly.
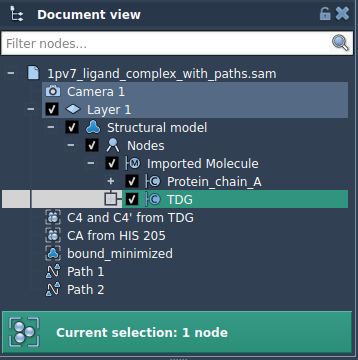
2. Define the Model to Export
Open the Export Along Paths app and expand the Advanced panel. Click Add to define the selected atoms as a model.
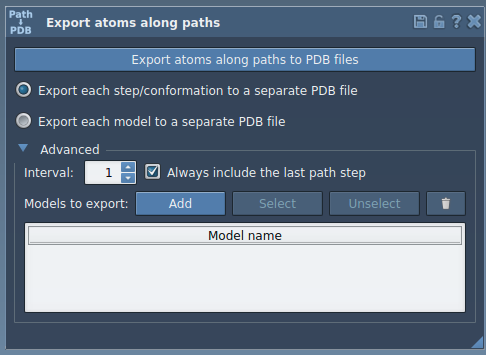
This creates a new model entry with a default name. You can rename it by double-clicking:
3. Customize and Export
Once defined, you can:
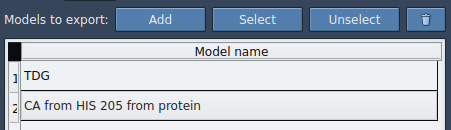
- Add additional subsets (e.g., binding site residues).
- Highlight or reset your selection anytime.
- Choose whether to export a single multi-frame PDB file or one file per frame.
When ready, select the path(s) in the Document view and click Export atoms along paths to PDB files. You’ll be prompted to choose the destination folder and a file prefix.
Useful Scenarios
- Sampling ligand unbinding pathways for enhanced sampling.
- Focusing on loop dynamics without exporting the full system.
- Reducing data size for iterative data analysis.
This small but powerful feature gives you more control over your exports, helping you work efficiently with only the data you actually need.
To learn more about using the Export Along Paths extension, visit the official documentation page.
SAMSON and all SAMSON Extensions are free for non-commercial use. You can get SAMSON at https://www.samson-connect.net.





