Molecular modelers often face the challenge of exporting atomic trajectories to analyze reaction coordinates, evaluate free energy changes, or visualize molecular pathways. SAMSON’s Export Along Paths extension simplifies this process, enabling users to export atomic coordinates along defined paths. This guide highlights how you can efficiently export atom trajectory data using two methods: exporting all atoms or a subset.
Why export atomic trajectories?
Atomic trajectories are essential for numerous molecular modeling applications. For instance, extracting ligand movement data can help analyze its exit/entry pathways from proteins. Similarly, exporting atomic coordinates along reaction paths enables molecular simulations like free energy calculations. Whether you’re studying ligand binding/unbinding or tracking reaction coordinates, precise trajectory export is vital for meaningful insights.
Start here: Basic setup
To begin with the Export Along Paths extension:
- Log into SAMSON Connect and add the extension from its extension page.
- Restart SAMSON to activate the extension.
- Select one or more paths in the Document view.
- Choose an export mode:
- All frames in a single PDB file.
- Each frame as a separate PDB file.
- Click Export atoms along paths to PDB files.
- Expand the Advanced panel within the app.
- Select your desired group of atoms in the Document view (e.g., ligand
TDG). - Click Add to define this selection as a model for export.
- Rename models by double-clicking their name.
- Add multiple models if you want to export various subsets of atoms.
- Use the context menu or buttons like Select and Reset to refine your selection.
- Generating reaction coordinate files for free energy studies.
- Exporting ligand entry/exit trajectories for enhanced sampling studies.
- Creating intermediate states for visualization and simulations.
- Tracking specific atom subsets (e.g., ligands, binding sites, or backbones).
Exporting trajectories: Two key methods
Option 1: Export all atoms
You’ll be prompted to select a destination folder and file prefix. For added precision, you can expand the Advanced section to configure frame export intervals.
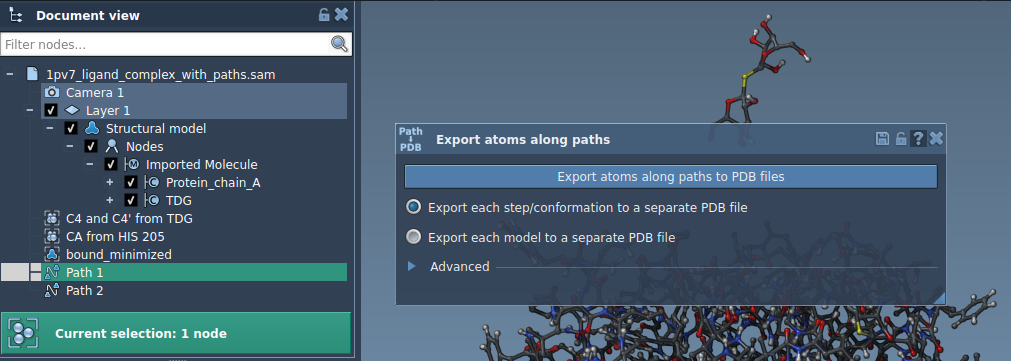
Option 2: Export specific atoms
In some cases, you may want to focus on specific parts of a structure—like a ligand or binding site. Here’s how to export a subset of atoms:
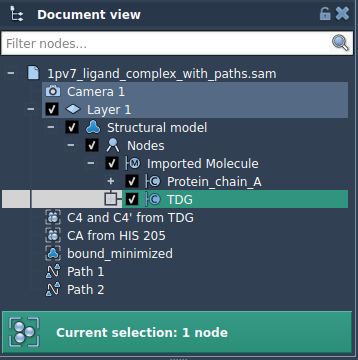
Once defined, the selected atoms will appear as a named model:
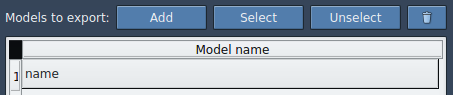
Finally, select an export format (single or multiple PDB files), choose paths, and click Export atoms along paths to PDB files. Choose a destination folder and file prefix when prompted.
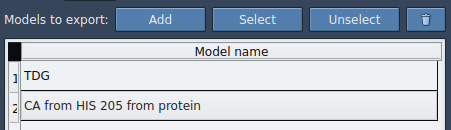
Applications in molecular modeling
Here are some practical uses for exporting atomic trajectories:
For a detailed, step-by-step walkthrough, visit the full documentation on exporting atom trajectories.
Note: SAMSON and all SAMSON Extensions are free for non-commercial use. Download SAMSON today at https://www.samson-connect.net.





