For molecular modelers, a significant challenge lies in refining docking calculations to achieve accurate and efficient outcomes. One of the key parameters influencing this process is the “search domain”—the area within which the docking algorithm searches for potential binding modes. A well-defined search domain can not only save computational time but also pinpoint regions of interest within a receptor structure. In this blog post, we’ll walk you through how to adjust and optimize the search domain using the AutoDock Vina Extended SAMSON Extension.
Why Adjust the Search Domain?
By default, the search domain is set to encompass the entire receptor structure. While this is suitable for general docking, it may lead to unnecessary computation if the binding site is small and well-known. Reducing the domain to focus on a specific binding site benefits by:
- Reducing computational overhead.
- Focusing resources on the most relevant interaction regions.
- Enhancing the accuracy of predictions within a predefined target area.
Step-by-Step: Setting the Search Domain
Here’s how you can set the search domain for your docking project:
1. Basic Adjustments
The search domain’s center and size can be modified directly within the “AutoDock Vina Extended” interface. If you are new to docking or working with a large receptor, this is the simplest way to refine the search area.
2. Fine-Tuning in the Viewport
If you prefer hands-on adjustments, you can alter the search domain box directly in the Viewport. Grab the spherical controllers at the corners of the yellow search box and adjust them to fit the intended binding site. This approach allows you to visually align the domain with specific receptor regions.
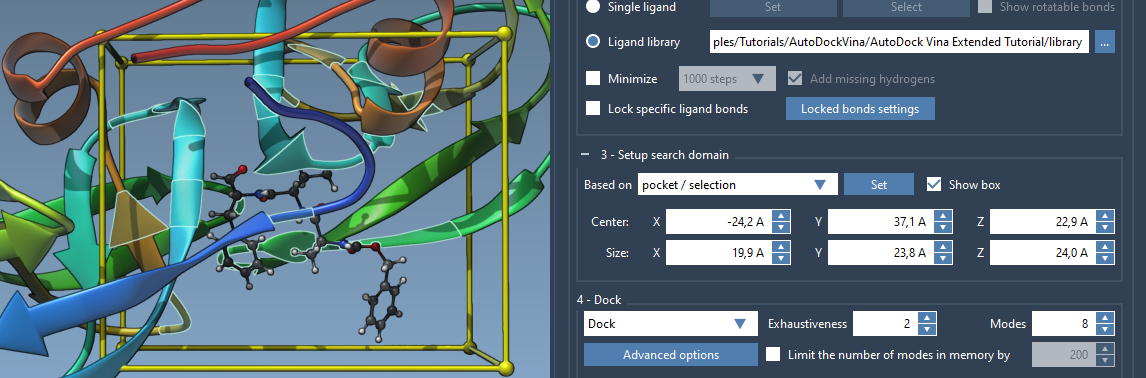
3. Selection-Based Domains
Another effective way to set the search domain is by leveraging selections. For instance, if you have prior knowledge of the binding site:
- Choose nodes or atoms surrounding the binding site.
- Select “Based on pocket/selection” from the interface.
- Click the Set button to adjust the search domain accordingly.
Alternatively, if your receptor includes a bound ligand, select it and use the Select > Biology > Binding sites option. This automatically determines the binding site and adjusts the search domain to match.
4. Verifying Your Changes
Once the search domain is set, verify its placement and size in the Viewport. Use the Show box toggle to hide or reveal the search box as needed. If necessary, adjust your receptor and ligand alignment through SAMSON’s Move editors.
Key Considerations
Keep in mind:
- The search domain is defined in global XYZ coordinates, so ensure your system is properly oriented.
- Over-restricting the search domain may exclude potential binding regions and impact docking results.
- You can always readjust the domain if the initial setup does not deliver optimal conformations.
Explore More
Accurately defining the search domain is a critical step to improving docking success. If you’re interested in learning more about docking ligands with AutoDock Vina Extended, visit the full documentation at this link.
SAMSON and all SAMSON Extensions are free for non-commercial use. You can get SAMSON at this link.





