Molecular modeling often involves analyzing complex motions within proteins, ligands, and other biomolecules. One significant challenge for molecular modelers is visualizing and understanding the collective motions, such as ligand unbinding or domain shifts. How do you elegantly capture and analyze these movements? This is where SAMSON’s Pathlines visual model steps in, offering an intuitive way to display the center-of-mass (COM) motion along selected paths.
In this post, let’s dive into how you can visualize the center-of-mass motion using Pathlines in SAMSON and make the most of this feature to solve real-world problems in molecular modeling.
What Problem Does Pathlines Solve?
Understanding the motion of ligands, residues, or entire protein domains is essential when studying biochemical behaviors like diffusion, unbinding, or structural transitions. Without a clear, dynamic visualization, it can be difficult to pinpoint exactly how these entities move or interact with their environments.
The Pathlines feature in SAMSON simplifies this process. It allows you to visually track the trajectory of selected atoms’ centers-of-mass. Whether you’re analyzing ligand unbinding pathways or collective domain motions, Pathlines provides an elegant overview of these motions, reducing the complexity of interpreting data from simulation results.
How to Start?
To use Pathlines effectively, begin by setting up your environment in SAMSON:
- Install Pathlines: If you haven’t already, log into SAMSON Connect, visit the Pathlines Extension page, and hit Add. Restart SAMSON to enable the feature.
- Load the sample system: For hands-on practice, load the sample system provided in the tutorial document. This contains a structural model of Lactose permease with its ligand and predefined unbinding paths.
Step 1: Select Atoms and Paths
To create Pathlines, start by selecting the group of atoms for which you want to visualize the center-of-mass motion. You can do this in the Document view within SAMSON:
- If you want to focus on a specific subset of the system, select those atoms and the corresponding paths. You can hold Ctrl / Cmd to select multiple nodes.
- Alternatively, if no specific atoms or paths are selected, SAMSON will use the entire molecular system and all paths in the loaded document.

Step 2: Create a Visual Model
Once selections are done, head to Visualization > Visual model > More… or use the shortcut Ctrl/Cmd + Shift + V. In the dialog that appears:
- Choose Pathline of the center of mass.
- Click OK to generate the visual representation.
This creates a pathline showing the COM motion along the selected pathways, offering a clear visualization of the molecular dynamics in action.
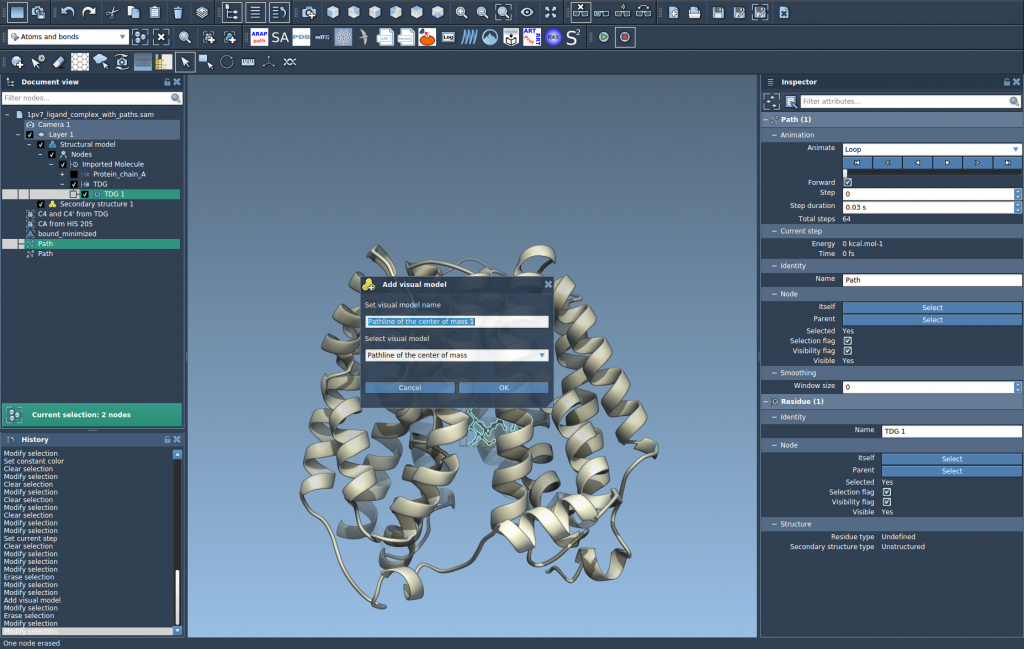
Step 3: Explore and Customize
After generating Pathlines, explore and adjust the visualization further:
- Double-click a path in the Document view to start or stop it.
- Access additional options by right-clicking a path and navigating through the Path > … menu.
If you want to tweak the aesthetics, select the Pathline in the Document view and open the Inspector using Ctrl/Cmd + 2. Here, you can adjust properties like color and thickness to tailor the visual model to your needs.
Use Cases
Pathlines is particularly useful for:
- Visualizing ligand unbinding and rebinding routes.
- Analyzing collective domain motions in protein complexes.
- Exploring center-of-mass movements in reaction coordinate workflows.
For a more detailed guide, explore the official documentation at this link.
Note: SAMSON and all SAMSON Extensions are free for non-commercial use. Experience the power of SAMSON today by downloading it from SAMSON Connect.





