For molecular modelers tackling complex protein dynamics, one common challenge is generating smooth and realistic transition paths between two protein conformations. Whether you’re performing conformational analyses, modeling transition states, or setting up umbrella sampling simulations, defining biologically meaningful pathways can be tricky and time-consuming. This is where SAMSON’s As-Rigid-As-Possible (ARAP) Interpolation Extension comes in.
The ARAP Interpolator enables you to compute a realistic conformational transition path between two protein structures in just a few seconds. This blog post will guide you through the workflow and highlight why ARAP can be a game-changer in your molecular modeling projects.
Why Use the ARAP Interpolator?
Here are some of the key benefits ARAP provides:
- It computes continuous structural transitions with a biologically meaningful geometric model, making it ideal for realistic simulations.
- You can visualize and export intermediate conformations for further analysis or sharing.
- It assists in building reaction coordinates that are critical for free energy simulations.
In essence, it’s fast, intuitive, and specifically designed to enhance molecular workflows.
How to Get Started
Getting started with the ARAP Interpolator is easy and follows a few simple steps:
- Install the Extension: Log into SAMSON Connect, visit the ARAP Interpolator extension page, and click Add. Restart SAMSON to enable the extension.
- Prepare Protein Structures: In SAMSON, use Home > Fetch to load the protein conformations (e.g.,
1DDTand1MDT). Remove any unnecessary chains, water, ligands, and ions using Home > Prepare. - Create Conformations: Label your starting and goal conformations using SAMSON’s Edit > Conformation tool. For instance, name the cleaned structures
1DDT Aand1MDT A.
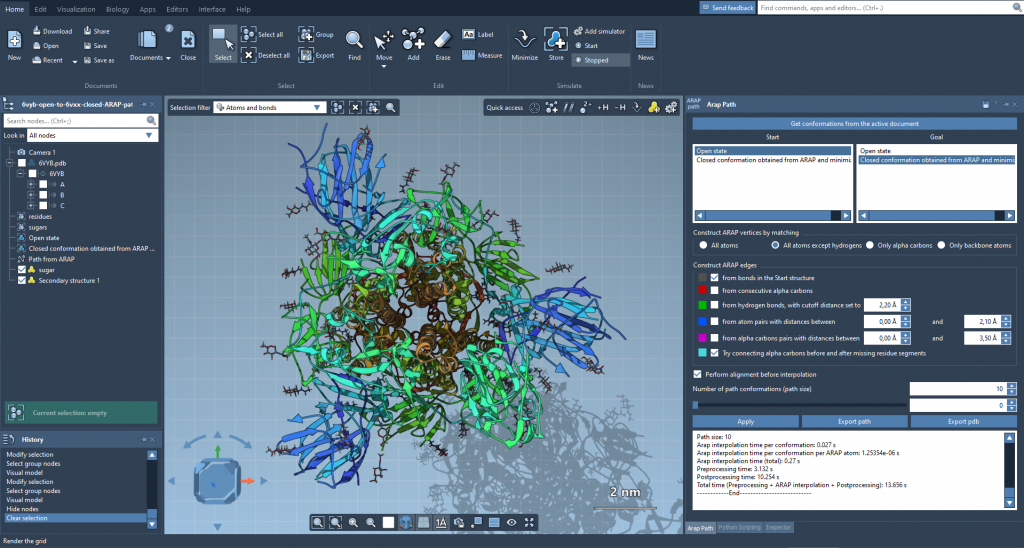
The ARAP Interpolation tool can be applied to problems like generating a path between conformations of the SARS-CoV-2 spike protein, as shown above.
Running the ARAP Interpolation App
Once your conformations are ready:
- Open the ARAP Interpolation app through Home > Apps > Biology > ARAP Path Interpolation.
- Select Start (
1DDT A) and Goal (1MDT A) conformations from the interface. - Choose how atoms will be matched (e.g., “All except hydrogens”) and how edges will be constructed for the ARAP geometric model (e.g., “from bonds in the Start structure”).
- Set the number of intermediate conformations (e.g., 20) to generate a smooth transition path.
When all settings are configured, click Run. Within seconds, the app will compute a physically plausible transition path between the starting and goal conformations.
Analyze and Export Results
After computation, the app provides several useful features:
- Use the slider tool to scrub through and visualize interpolated conformations along the generated path.
- View a color-coded representation of ARAP edges to understand how the interpolation path was constructed.
- Export results into a PDB file for further analysis or save the path as a trajectory object in SAMSON.
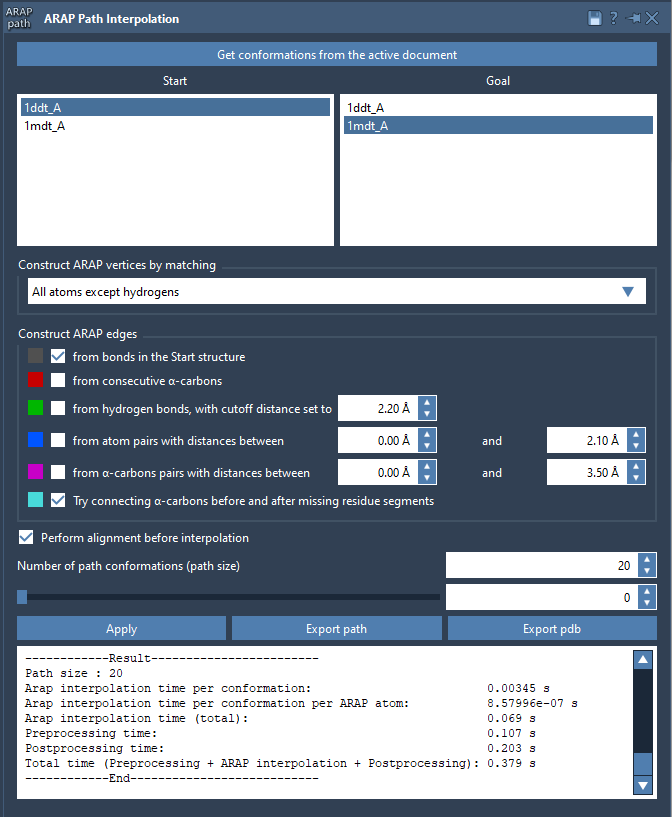
The ARAP app presents results in a clear interface, enabling easier interpretation and analysis.
Get More Out of Your Molecular Simulations
If you’re looking to enhance your computational workflows, ARAP-generated paths can serve as excellent inputs for advanced simulation techniques. Applications range from umbrella sampling in GROMACS Wizard to steered molecular dynamics or dimensionality reduction of structural ensembles.
This tool significantly simplifies transition modeling for protein structures, saving time while providing accurate and biologically meaningful results.
To delve deeper into ARAP Interpolation, visit the official documentation page.
Note: SAMSON and all SAMSON Extensions are free for non-commercial use. You can get SAMSON at SAMSON Connect.





