In molecular modeling, precision is everything. Whether you’re narrowing down specific residues, focusing on atoms of interest, or identifying structural properties, an efficient and intuitive way to search and filter molecular data can save immense effort. This is where SAMSON’s Node Specification Language, or NSL, becomes a vital tool in the modeler’s toolbox. But how exactly does NSL streamline node selection, and how can you leverage it best? Let’s dive in.
Understanding NSL – A Quick Overview
NSL is a powerful command-line language integrated into SAMSON for selecting nodes (atoms, residues, bonds, etc.) and structures from models based on their properties. This is ideal for tasks such as selecting ligands, identifying specific residues, or filtering nodes within proximity to certain coordinates.
Below, we’ll focus on two central use cases: discovering specific nodes using the Find command and filtering nodes in the Document View. With these tools, molecular modelers can significantly enhance their workflows.
Using the Find Command for Precision Node Search
The Find command allows you to input NSL expressions directly to identify targeted nodes. Let’s say you’re dealing with a large protein structure and need to select all alanine residues (ALA) in your workspace. You simply type "ALA" into the Find command interface, press the Tab key for autocompletion, and let context-sensitive suggestions guide your selection.
Not only does this eliminate manual navigation of complex molecular setups, but it also allows queries with logical operations such as and, or, and not. For instance:
C or H: Selects only carbon or hydrogen atoms.n.t residue and not residue.type ALA: Selects all residues except alanine.

Effortlessly combining such expressions allows precise selection of exactly what you’re interested in. And here’s a helpful tip: You can ask the built-in AI Assistant for help! Simply click on the AI button next to the input box, and it will generate an appropriate NSL expression customized to your document’s structure.
Filtering Nodes in the Document View
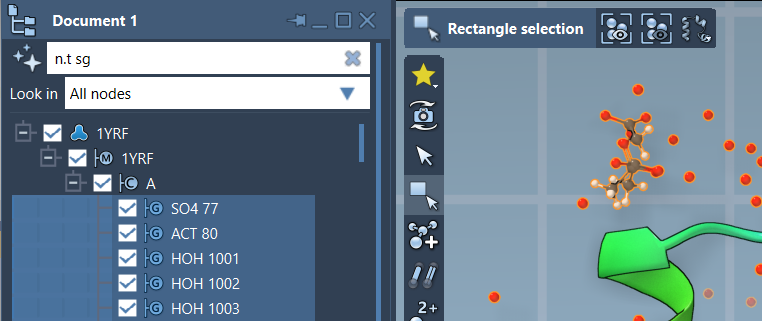
The Document View in SAMSON offers an additional, interactive way to refine your node selection. By entering NSL expressions like n.t sg, which selects structural group nodes, modelers gain the ability to quickly highlight or isolate parts of a molecule and filter directly from the active document.
For example, using n.t backbone (or its short version n.t bb) will immediately highlight all backbone nodes in your model. Structured command inputs also allow proximity-based selection—for example:
"CA" within 5A of S: Matches all nodes named “CA” (alpha carbons) within 5 angstrom of any sulfur atom.residue.secondaryStructure helix: Matches only residues in helical secondary structures.
Just like with the Find command, the AI Assistant is also available to generate highly specific NSL queries based on your dataset in the Document View workspace.
Getting Started with Examples
If all this sounds intriguing, NSL supports intuitive and familiar syntax for various molecular modeling needs. Here are just a few examples to explore:
node.category ligand, receptor: Matches ligands and receptors.residue.id 20:40: Selects residues with IDs in the range 20 to 40.node.type sideChain having S: Matches side chain nodes with at least one sulfur atom."CA" beyond 5A of node.selected: Selects alpha carbons that are beyond 5 angstrom of the current selection.
By mastering these commands and exploring the full potential of NSL, you can achieve superior precision and efficiency in your molecular modeling workflows. To delve deeper, check out the full documentation on Node Specification Language.
SAMSON and all SAMSON Extensions are free for non-commercial use. Download SAMSON today at https://www.samson-connect.net.





