Avoiding Topology Conflicts When Modeling Protein Replicas
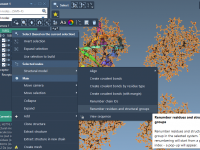
Quickly Finding Conformations with Many Atoms in SAMSON
One common task in molecular modeling—especially when working with large molecular datasets or complex simulations—is identifying which conformations meet certain structural criteria. For example, maybe you’re only interested in conformations that contain a large number of atoms. Manually searching through…
Making Sense of Animation Visibility in Molecular Models
When working on molecular modeling projects involving animations—whether for simulations, presentations, or collaborations with colleagues—clarity and control over what you see and what you don’t are critical. But how do you programmatically control visibility, selection, and naming of animation nodes…
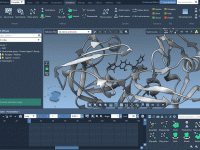
Installing Python Packages in SAMSON Without Leaving the Interface
For many molecular modelers, scripting in Python has become a core part of their workflow. Whether you’re automating analyses, performing simulations, or applying data science tools, access to Python packages is essential. But managing packages—especially across different environments—can be frustrating.…