As a molecular modeler, brainstorming and creating a library of molecules with slight variations can be both time-consuming and challenging. But what if there were a way to automate this process while experimenting with different fragments? The Replace Fragments functionality in the SMILES Manager extension of SAMSON does exactly this, allowing you to efficiently generate large libraries of molecules by substituting predefined fragments.
Why Replace Fragments?
In molecular modeling and cheminformatics, being able to explore slight variations of a base molecule is critical for tasks such as structure-activity relationship (SAR) studies, property optimization, and screening for molecular candidates. However, modifying a molecule manually to generate new variants can be tedious and repetitive. The Replace Fragments feature offers a way to systematically and quickly test many fragment replacements, thereby accelerating workflows.
Step-by-Step Guide to Using Replace Fragments in SAMSON
Follow these steps to generate your own molecule libraries seamlessly:
- Specify Initial Molecules: Import the molecule you want to use as a starting point. This can be done by opening a relevant file via the File > Open initial molecules menu, manually entering a molecule, or importing an active molecule from the SAMSON Document using Import > SAMSON Selection as initial molecule. For example, let’s start with this molecule:
COc1cc(CNC(=O)CCCC/C=C/C(C)C)ccc1O - Define the Fragment to Replace: Specify the part of the molecule you wish to replace. Input the fragment manually or import it from a selection in the SAMSON Document via Import > SAMSON Selection as replaced fragment. For this example, let’s use:
[*:1]C(=O)NC[*:2] - Provide Replacement Fragments: Feed in the set of fragments to be used as replacements. You can import these from a file via File > Open fragments to replace with… or manually add them. Below is an illustrative list:
1234567891011[*:1]S(=O)(=O)NC[*:2][*:1]C(C(F)(F)(F))NC[*:2][*:1]C1CC1NC[*:2][*:1]C(F)=CC[*:2][*:1]C1COC1NC[*:2][*:1]C1COC1OC[*:2][*:1]C1=NN=C([*:2])N1[*:1]C1=NN=C([*:2])O1[*:1]N1N=NC([*:2])=C1[*:1]N1N=NC([*:2])=N1[*:1]C1=NOC([*:2])=N1
- Run Fragment Replacement: Select Run > Replace fragments to initiate the process. SAMSON will systematically generate new molecules by substituting the specified fragment in your initial structure with each of the replacement fragments provided.
- Review and Export Results: The results of the replacements are stored in the Results table within the Replace Fragments tab. From here, you can view them in the active SAMSON document, convert them into 3D structures, or export them into files or grid images. For visual ease, buttons to save the table as a grid or file are conveniently located at the bottom right of the table.
Here’s an example of how the process looks in SAMSON:
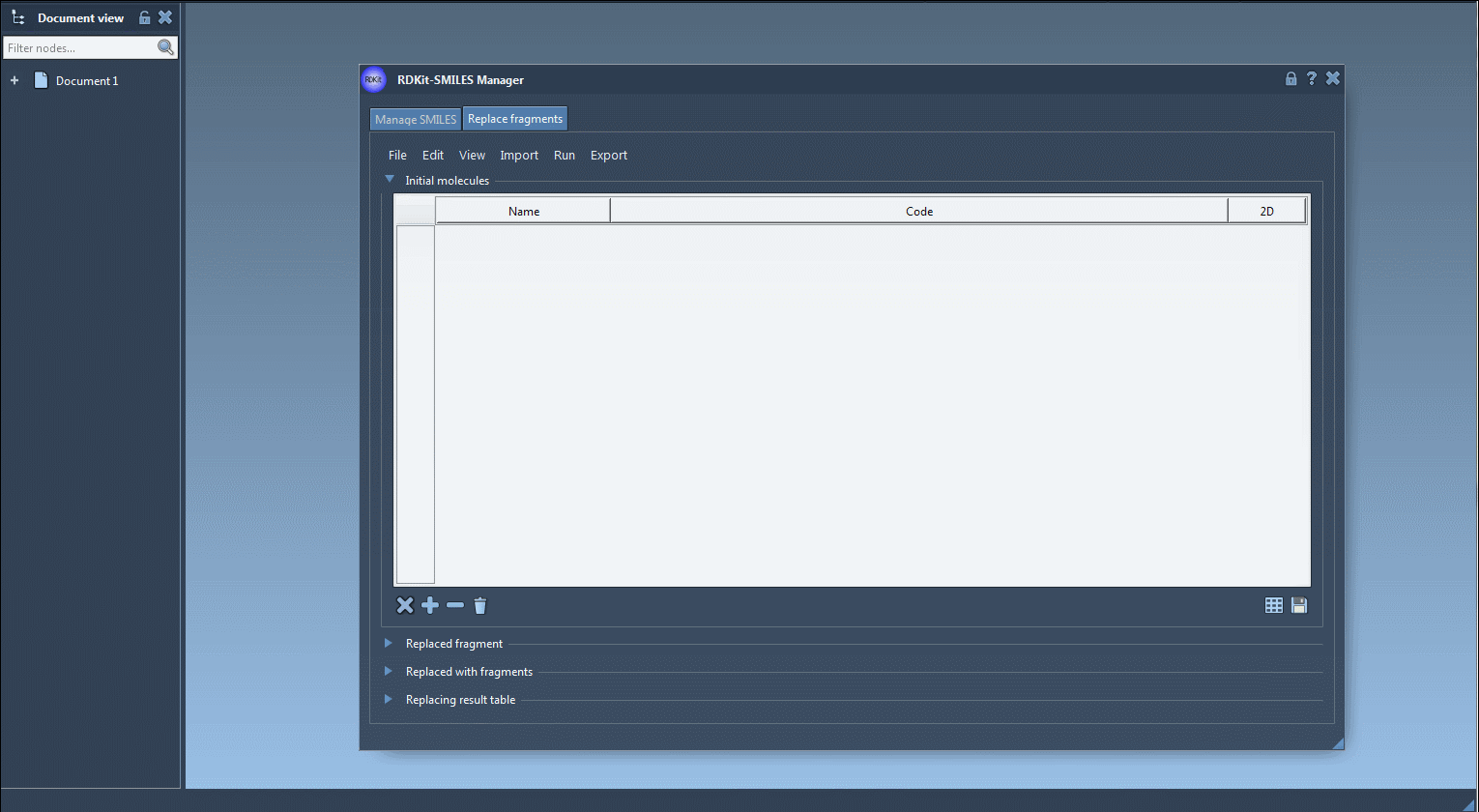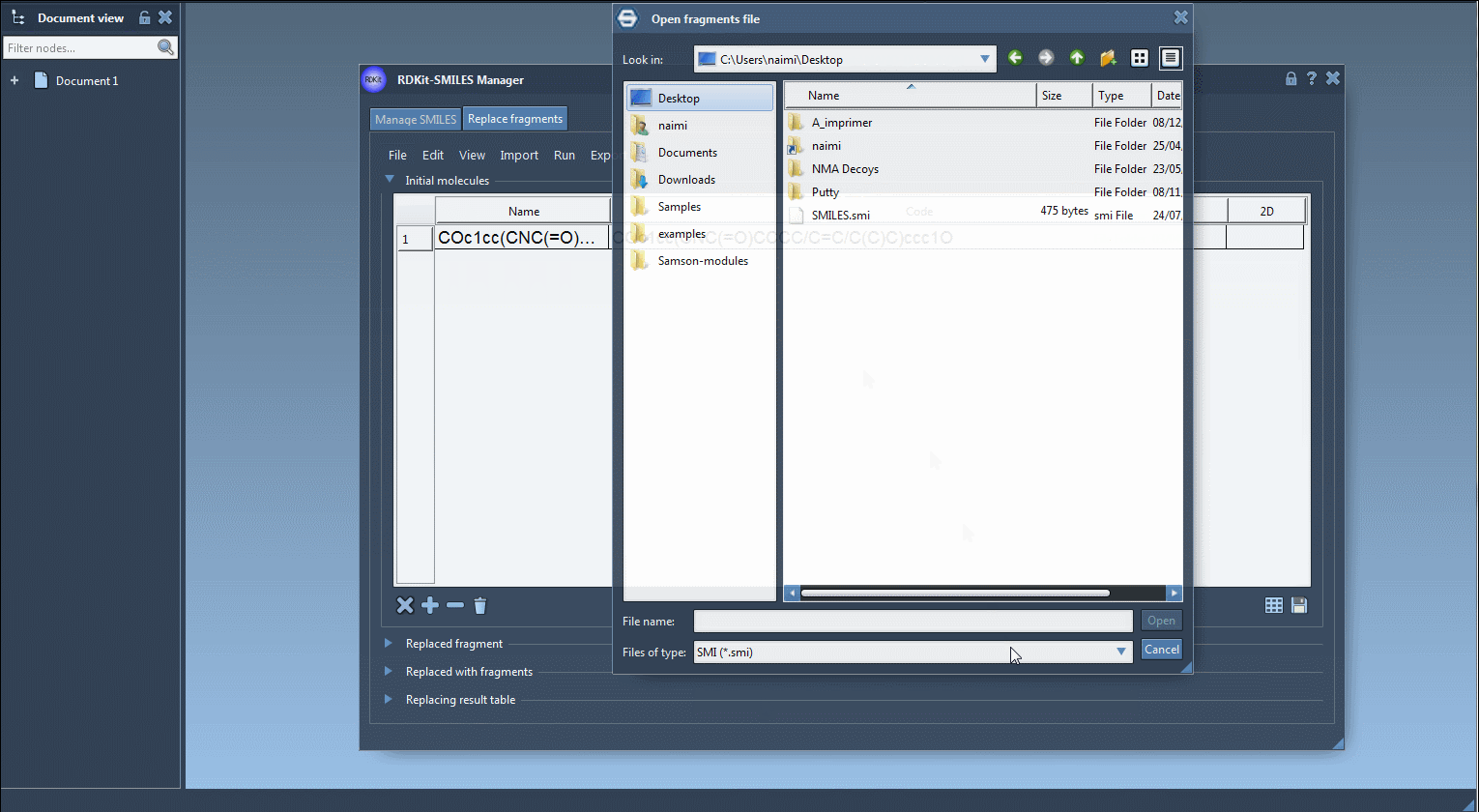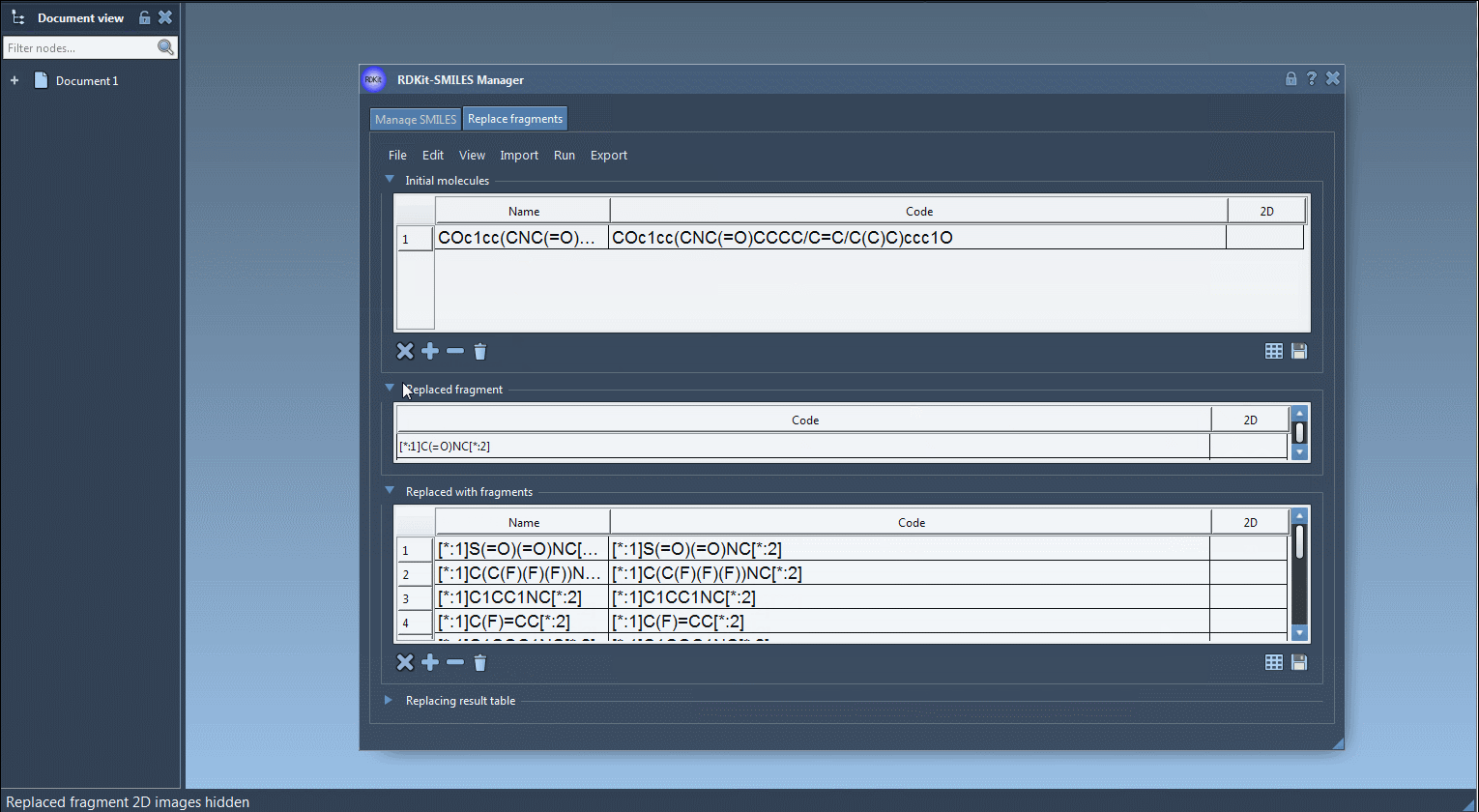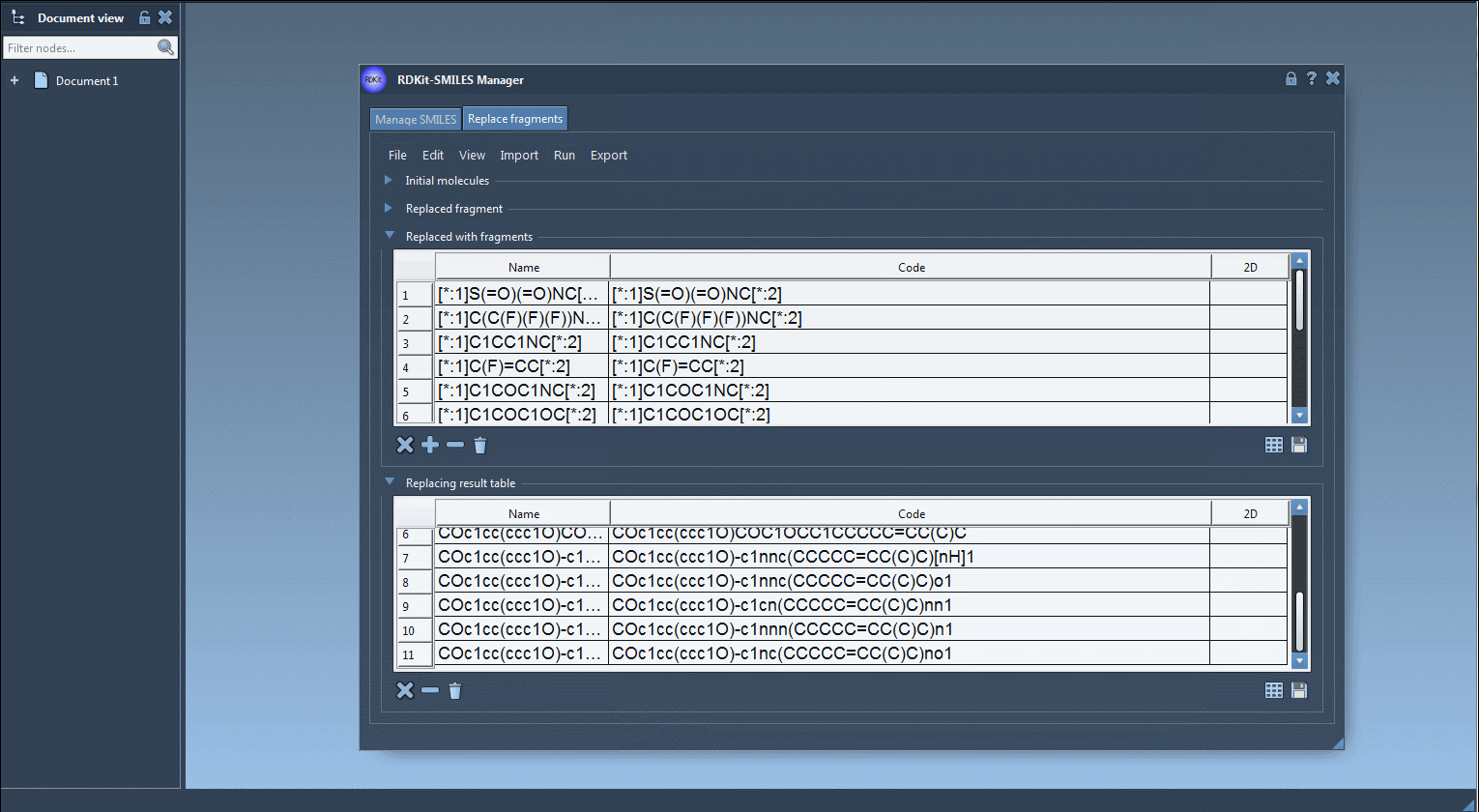
Upon generating your molecule library, you might test various properties, perform structural refinement, or simply save them for further usage. This functionality is especially helpful for quickly creating compound arrays for virtual screening or SAR studies.
Syntax Tips for RDKit Fragments
When specifying fragments in RDKit, use the syntax:
[*:1]C(=O)NC[*:2]
Here, [*:1] and [*:2] denote the two ends of the fragment, ensuring an accurate match during replacements. Proper definition of fragments is essential for obtaining meaningful results. SAMSON’s intuitive interface makes it easy to adjust inputs and regenerate outputs for refinement.
Conclusion
If you’re working on identifying isosteric replacements, crafting diverse molecular libraries, or exploring targeted chemical diversity, the Replace Fragments functionality in SAMSON’s SMILES Manager can save you hours of manual adjustment while maintaining precision and creativity.
Curious to dive deeper? Read the full documentation for a comprehensive guide and other advanced tips.
Note: SAMSON and all SAMSON Extensions are free for non-commercial use. Download SAMSON for free at https://www.samson-connect.net.





