Preparing ligands for molecular docking can be a tedious task, especially when working with libraries of ligands or when ensuring essential chemical features are present. However, streamlined solutions exist, and the AutoDock Vina Extended SAMSON Extension makes this process much more efficient. In this post, we’ll walk through key features and best practices for ligand preparation with this tool, helping you save time and improve your docking results.
Why Does Ligand Preparation Matter?
The success of molecular docking depends significantly on how well-prepared your ligands are. Problems such as missing hydrogens, incorrect bond configurations, or a lack of adequate minimization can compromise docking accuracy. AutoDock Vina Extended offers tools that address these issues efficiently, ensuring your ligands are ready to deliver meaningful docking simulations.
Step 1: Setting Up a Ligand
Preparing a single ligand in AutoDock Vina Extended is straightforward. Simply check the Single ligand option in the Set ligand interface. From there, select your ligand in the Document view, and click on the associated Set button. Here’s how it looks:
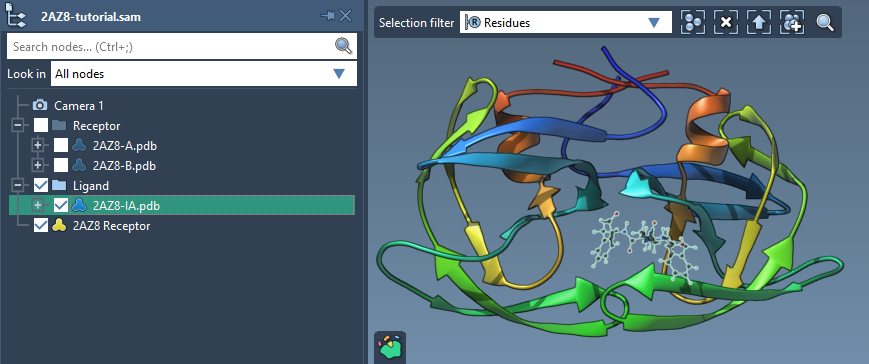
Once a ligand is set, you’ll see visual indicators for rotatable bonds in the Viewport. This lets you control which bonds should remain flexible during docking, optimizing the trade-off between computational efficiency and biological realism.
Step 2: Controlling Rotatable Bonds
Rotatable bonds can be toggled directly in the Viewport by clicking on the green cylinders representing them. Green means the bond is rotatable, and red means it’s locked. Additionally, you can access the Locked bonds settings to specify bond types that should always be rigid. Here’s an example of locking bonds:
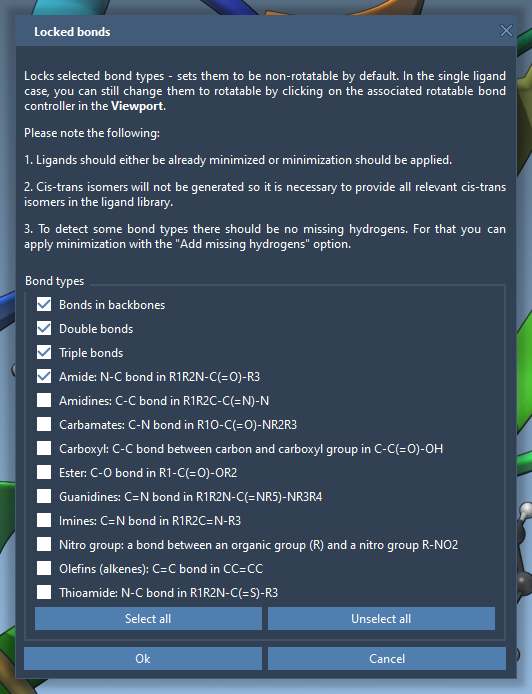
This flexibility allows you to fine-tune bond behavior across complex ligands, enhancing accuracy during docking simulations.
Step 3: Preparing Ligand Libraries
For those working with multiple ligands, AutoDock Vina Extended also supports libraries. Check the Ligand library option and browse to the folder containing your ligands. By default, all bonds in the library are considered rotatable, but you can apply the earlier locking mechanisms for consistency across the library. Here’s the simple interface for setting up a library:
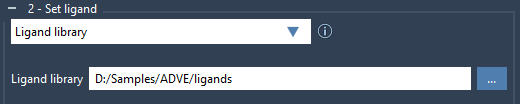
Step 4: Minimizing Ligands
If ligands in your library are not already optimized, AutoDock Vina Extended includes a minimization feature. This ensures ligands have realistic geometries before docking, especially critical for 2D inputs. You just need to check the Minimize option and choose a preset that best matches your needs. Missing hydrogens? No problem—this can be automatically handled during minimization.

The preset efficiently balances time and accuracy, so your ligands are ready for docking without unnecessary delays.
Conclusion
Effective ligand preparation sets the stage for successful docking experiments. By using AutoDock Vina Extended in SAMSON, you can efficiently handle tasks such as minimizing ligands, locking bonds, and processing libraries, ensuring your docking workflows remain smooth and accurate. For further details on how to leverage these features fully, visit the official documentation.
Note: SAMSON and all SAMSON Extensions are free for non-commercial use. You can download SAMSON at samson-connect.net.





